Abstract
Behavior is influenced by genes but can also shape the genetic structure of natural populations. Investigating this link is of great importance because behavioral processes can alter the genetic diversity on which selection acts. Gene flow is one of the main determinants of the genetic structure of a population and dispersal is the behavior that mediates gene flow. Baboons (genus Papio) are among the most intensely studied primate species and serve as a model system to investigate the evolution of social systems using a comparative approach. The general mammalian pattern of male dispersal and female philopatry has thus far been found in baboons, with the exception of hamadryas baboons (Papio hamadryas). As yet, the lack of data on Guinea baboons (Papio papio) creates a taxonomic gap in genus-wide comparative analyses. In our study we investigated the sex-biased dispersal pattern of Guinea baboons in comparison to hamadryas, olive, yellow, and chacma baboons using sequences of the maternally transmitted mitochondrial hypervariable region I. Analyzing whole-range georeferenced samples (N = 777), we found strong evidence for female-biased gene flow in Guinea baboons and confirmed this pattern for hamadryas baboons, as shown by a lack of genetic-geographic structuring. In addition, most genetic variation was found within and not among demes, in sharp contrast to the pattern observed in matrilocal primates including the other baboon taxa. Our results corroborate the notion that the Guinea baboons’ social system shares some important features with that of hamadryas baboons, suggesting similar evolutionary forces have acted to distinguish them from all other baboons.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Clarifying the genetic basis of animal behavior is essential to understand its evolution. Advances in molecular techniques in recent years have enabled researchers to pinpoint an increasing number of genes underlying specific traits, which may eventually help to explain individual behavioral variation in natural populations (Tung et al. 2010). However, behavior and genes are mutually influential, e.g., by triggering or preventing gene expression (Robinson et al. 2008; Tung et al. 2011) or by shaping the genetic structure of natural populations (Altmann et al. 1996; Avise 2004; Bohonak 1999; Di Fiore 2003; Melnick 1987). Investigating the influence of behavior on genetic structure is of great importance because behavioral processes can alter the genetic diversity on which selection acts.
One of the main pathways through which behavior can directly influence genetic diversity and population genetic structure is dispersal. Dispersal, an animal’s movement away from its natal area or group (Pusey and Packer 1987), is an important behavior underlying gene flow. Populations with high gene flow represent a panmictic and genetically more uniform entity, while restricted gene flow leads to several genetically differentiated demes, i.e., local interbreeding populations with distinct gene pools, that may react differently to selection pressures or may eventually diverge into separate species (Avise 2004).
Whereas birds tend to exhibit male philopatry and female-biased dispersal, in mammals male-biased dispersal and female philopatry are the norm, an observation that led Greenwood (1980) to hypothesize that the sex bias in dispersal tightly correlates with the mating system. In group-living species, the composition of the group (social organization, sensu Kappeler and van Schaick 2002) is immediately influenced by the immigration and emigration of individuals. Further, dispersal determines relatedness patterns within a group (Di Fiore 2003) and thus has profound impacts on the social relationships among individuals (social structure), as many social species preferably interact with close kin (Seyfarth and Cheney 2012; Silk 2002).
A sex bias in dispersal translates into a specific pattern of genetic population structure. When dispersal is biased toward one sex, uniparentally inherited genetic markers show incongruent patterns in population structure (Avise 2004). In mammals, the general pattern of female philopatry and male dispersal is reflected in strong geographic structuring of the maternally inherited mitochondrial DNA (mtDNA), but not the paternally inherited Y-chromosomal haplotypes (Avise 2004). Consequently, dispersal is a behavior that connects the social system of a species with its genetic diversity and represents a central factor in population genetics and population dynamics (Broquet and Petit 2009). Moreover, investigating the influence of dispersal patterns on the genetic variation of natural populations may help us to infer the social system of understudied taxa using genetic data (Di Fiore 2003).
The link between the social system and population genetic structure has been investigated in many species, including primates. Papio is among the best studied primate taxa and has widely been used as a model to study the evolution of social systems using a comparative socioecological approach (Barrett 2009; Barton et al. 1996). The wealth of data accumulated on their behavior and their wide distribution throughout Africa promotes them as a useful model to investigate the relationships between social systems and genetic structure.
In southern and eastern African baboons, e.g., yellow baboon (Papio cyncocephalus), chacma baboon (P. ursinus), and Kinda baboon (P. kindae), in which the dispersing sex is male, a strong geographical structuring of mtDNA haplotypes, but of neither Y-chromosomal nor autosomal markers, reflects their matrilineal organization (Burrell 2008; Burrell et al. 2011). Interestingly, the phylogenetically closely related hamadryas baboon (Papio hamadryas) exhibits a different social system in which male philopatry (Sigg et al. 1982; Swedell 2011) leads to a strikingly different genetic structure. For instance, there is no structuring of mitochondrial variation that corresponds to geography (Hammond et al. 2006; Hapke et al. 2001).
The Guinea baboon (Papio papio), on the northwestern fringe of the baboon distribution, has been proposed to share some features with the hamadryas baboon on the northeastern fringe (Jolly 1993, 2009; Jolly and Phillips-Conroy 2006). Like the hamadryas baboon, the Guinea baboon is suspected to be characterized by male philopatry and female dispersal (Jolly 2009). A study using microsatellites indeed found evidence for female-mediated gene flow in a Guinea baboon population in Senegal (Fickenscher et al. 2011), whereas a similar study on a population in Guinea-Bissau did not find signatures of sex-biased dispersal, probably owing to anthropogenic disturbance of the population and group compositions (Ferreira da Silva 2012). If the hypothesis that males are philopatric while females disperse in Guinea baboons is correct, we would expect to find little or no geographic structure in female specific genetic markers (mtDNA) in Guinea baboons. In contrast, if the geographic structure in mtDNA is strong, we would infer that gene flow in Guinea baboons is not female mediated, as in matrilocal primates.
We here investigate the taxon-wide pattern of female gene flow in Guinea, hamadryas, olive, yellow, and chacma baboons using sequences of the maternally transmitted mitochondrial hypervariable region I (HVRI). We infer common patterns by evaluating data over a wide range to overcome the noise induced by different local conditions in single populations. We reconstruct haplotype networks and test for isolation by distance to demonstrate the geographical distribution of genetic variation. We further estimate the hierarchical population structure. We expect to find a high diversity of mitochondrial haplotypes within demes and no significant variation among demes, with shared haplotypes existing between even distantly located demes.
Methods
Sample Collection
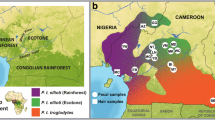
Between 1995 and 2012 we collected fecal samples of free-ranging Guinea and hamadryas baboons covering the whole of their respective ranges. We also collected samples of olive, yellow, and chacma baboons for comparison. Fecal samples were stored either in ethanol or on silica, or according to the two-step method (Nsubuga et al. 2004; Roeder et al. 2004). In addition, we analyzed available tissue samples of hamadryas baboons of known provenance provided by the King Khalid Wildlife Research Center (KKWRC), Saudi Arabia, and published sequences were downloaded from GenBank (Electronic Supplementary Material [ESM] Table SI). In total, our data set included 221 samples of hamadryas baboons (74 and 12 of these samples have previously been published by Hapke et al. [2001] and Winney et al. [2004], respectively) representing 27 different locations, 376 samples of Guinea baboons representing 62 different locations, 112 samples of olive baboons representing 25 different locations (18 of these samples have previously been published by Hapke et al. [2001]), 44 samples of chacma baboons representing 17 different locations, and 24 samples of yellow baboons representing 11 different locations (Fig. 1; overview in Table I; details in ESM Table SI). For each sampling site we recorded GPS coordinates (we used only general site-specific coordinates for our analysis because samples were usually found in a clumped fashion only separated by a few meters). We use sampling location as a proxy for social group, as most samples were collected from unhabituated individuals that in some cases were not observed directly. However, to account for the uncertainty of whether samples actually represent the same social group we use the term “deme” to refer to samples taken from the same location. Because the exact distribution of Guinea baboons in West Africa is unclear, we also collected samples outside of the range indicated in the literature and included them in the analysis if direct observation confirmed that the species was Papio papio. For a subset of the hamadryas and Guinea baboon samples, we tested for repeated sampling of individuals using autosomal microsatellites (Ferreira da Silva 2012; Fickenscher et al. 2011; Hapke et al. 2001). For the remaining samples we did not explicitly test if samples originated from different individuals, as we followed some precautions in the sampling protocol, e.g., only one sampling session per site, a minimum distance between samples of 2 m, that make double sampling negligible (Fickenscher et al. 2011; Hapke et al. 2001).
This project complied with the protocols approved by the German Primate Center, Göttingen, Germany and the animal care regulations and principles of the International Primatological Society for the ethical treatment of nonhuman primates. Permits for research and sample export were obtained from the local authorities and research adhered to the legal requirements of the respective countries in which research was conducted.
Laboratory Analyses
Total genomic DNA was extracted from fecal samples with the QIAamp DNA Stool Mini Kit (Qiagen, Hilden, Germany) and from tissue samples with the DNeasy Blood and Tissue Kit (Qiagen) according to the manufacturer’s protocols with slight modifications (Haus et al. 2013). To avoid contamination we followed established protocols and performed extractions, polymerase chain reaction (PCR), and sequencing in separate laboratory rooms. All steps were monitored for contamination with negative (HPLC water) controls.
We amplified and sequenced a fragment of the HVRI of the mitochondrial genome (D-loop) comprising 341 base pairs (bp) using primers from previous studies (Hapke et al. 2001). PCR amplification was performed on a Sensoquest labcycler in a total volume of 30 μl composed of 1.0 μl of DNA extract (20–40 ng/μl), 19.6 μl of H2O, 3.0 μl of 10× buffer (contains 15 mM MgCl2, Biotherm), 1.0 μl of forward primer (0.33 μM; 5′-CTGGCGTTCTAACTTAAACT-3′) and 1.0 μl of reverse primer (0.33 μM; 5′-GTAGTATTACCCGAGCGG-3’), 0.2 μl of dNTPs (0.16 mM), 4.0 μl of BT (0.6 mg/ml of bovine serum albumin [BSA] + Triton), and 0.2 μl of BioThermTM 5000 Taq polymerase (1U; Genecraft, Germany). PCR conditions comprised a pre-denaturation step at 94°C for 2 min, followed by 35–40 cycles at 94°C for 1 min, 51°C for 1 min, 72°C for 1 min, and a single final extension step at 72°C for 5 min. PCR products were checked on 1% agarose gels, excised, and purified with the Qiagen Gel Extraction Kit (Qiagen). Both strands of each sample were sequenced on an ABI 3130xL sequencer using the BigDye Terminator Cycle Sequencing Kit (Applied Biosystems, Germany). We checked and aligned sequences manually in Bioedit 7.0.5.3 (Hall 1999).
To test for the accuracy of the sequences we amplified random samples and/or sequenced repeatedly. To avoid the amplification of nuclear mitochondrial insertions (numts), we selected primers highly specific to amplify only mitochondrial fragments of Papio (Hapke et al. 2001). We did not observe double peaks in chromatograms or sequence ambiguities when comparing both strands or repeatedly sequenced samples, which would indicate that numts could have flawed our analysis (Bensasson et al. 2001; Thalmann et al. 2004).
Statistical Analyses
We estimated number of segregating sites S, nucleotide diversity π (Nei 1987), number of haplotypes, and haplotype diversity hd (Nei 1987) for each species, both range-wide and separately for each deme, in DnaSP 5.10.01 (Librado and Rozas 2009). Demes with only one sample were excluded from intra-deme diversity calculations. To compare genetic variation within and among demes we performed a hierarchical analysis of molecular variance (AMOVA) (Excoffier et al. 1992) in Arlequin 3.5.1.2 (Excoffier and Lischer 2010) using 10,000 permutations. For this analysis we grouped demes into distinct regions according to their geographic clustering; i.e., the distance to the next closest deme had to be <70 km, as this is a distance that has been shown to affect population structure of nuclear markers in Guinea baboons for two different populations (Ferreira da Silva 2012; Fickenscher et al. 2011) (see Fig. 4; overview in Table I; details in ESM Table SI). Because the grouping may also affect the results of the AMOVA, we also ran the analysis with a weaker clustering, where the smallest distance had to be <150 km. The fixation indices calculated in the AMOVAs, which are measures of genetic differentiation ranging from 0 (no differentiation, high gene flow) to 1 (complete differentiation, no gene flow), were used to evaluate the amount of gene flow within each species at the three respective spatial levels. Using Alleles in Space (AIS) 1.0 (Miller 2005) we furthermore quantitatively analyzed the correlation between genetic and geographic distances with a Mantel test (Mantel 1967) for each species, testing for significance with 10,000 replicates. We split this analysis for hamadryas baboons for the Arabian and the African populations to account for the Red Sea acting as a major barrier to gene flow. To visualize the genetic distances and geographical distribution of haplotypes, we reconstructed a haplotype network using output data generated in Arlequin and visualized using HapStar 0.6 (Teacher and Griffiths 2011) for Guinea and hamadryas baboons, respectively (but not for the other species, where sampling was too sparse).
Results
The 221 hamadryas baboon samples yielded 93 different haplotypes with 84 segregating sites (S), a haplotype diversity (hd) of 0.978, and nucleotide diversity (π) of 0.042. The 376 Guinea baboon samples yielded 104 different haplotypes with S = 90, hd = 0.947, and π = 0.024. The remaining three species (chacma, yellow, and olive baboons) showed very similar hd values, but both π and S were considerably higher than in hamadryas and Guinea baboons (Table I). When comparing the mean intra-deme diversity indices hamadryas baboons showed slightly higher values than Guinea baboons (Table I).
Separate hierarchical AMOVAs for each species revealed highly significant levels of structuring and comparable results for both Guinea and hamadryas baboons and a different pattern for chacma, yellow, and olive baboons (Table II; Fig. 2). In Guinea and hamadryas baboons more than half of the species-wide genetic variation (Guinea baboons: 54%, hamadryas baboons: 54%) was a result of variation within demes (Table II). Only a minor proportion of the genetic variation in these two species was explained by differences between demes (Guinea baboons: 7%, hamadryas baboons: 11%), whereas differentiation between regions contributed slightly more than a third of the genetic variation (Guinea baboons: 39%, hamadryas baboons: 34%) (Table II). In contrast, intra-deme diversity accounted for only 8%, 14%, and 14% of the variation in olive, yellow, and chacma baboons, respectively, while by far the highest percentage of genetic variation in these species was explained by variation among regions (olive baboons: 89%, yellow baboons: 75%, chacma baboons: 79%).
Analysis of molecular variance describing the hierarchical distribution of genetic variation for baboons in comparison to two patrilocal and four matrilocal primates. (1) Excoffier et al. 1992; (2) Goldberg and Ruvolo 1997; (3) Burrell 2008; (4) Rosenblum et al. 1997; (5) Modolo et al. 2005; (6) Nietlisbach et al. 2012; (7) Guschanski et al. 2006.
Changing the clustering from 70 km to 150 km did not greatly affect the overall results and mainly reallocated some of the intraregion variation to among-region variation (Table II). The fixation indices are also considerably smaller in Guinea and hamadryas baboons than in the three matrilocal species, indicating higher mitochondrial gene flow than in olive, yellow, and chacma baboons on all three spatial levels (among regions, among demes within regions, within demes; Table II). We also compared our AMOVA results to published data on both matrilocal and patrilocal primate species. This comparison showed that the distribution of genetic variation in hamadryas and Guinea baboons is very similar to that of humans and chimpanzees (Pan troglodytes) (patrilocal), whereas the distribution of genetic variation in chacma, yellow, and olive baboons is more similar to that in macaques (Macaca spp.), orang-utans (Pongo spp.), and mouse lemurs (Microcebus spp.) (matrilocal) (Fig. 2).
The Mantel test revealed significant correlations of genetic and geographic distance in all five species (Fig. 3). This isolation-by-distance effect was responsible for less than half of the variation in Guinea and hamadryas baboons, with a great deal of scatter around the regression line (Guinea baboons: r = 0.48, P < 0.001; hamadryas baboonsAfrica: r = 0.34, P < 0.001; hamadryas baboonsArabia: r = 0.25, P < 0.001). It was much stronger in the matrilocal baboons: isolation-by-distance explained more than half of the variation in olive, chacma, and yellow baboons and reached a correlation coefficient as high as 0.90 in olive baboons (olive baboons: r = 0.90, P < 0.001; yellow baboons: r = 0.76, P < 0.001; chacma baboons: r = 0.54, P < 0.001).
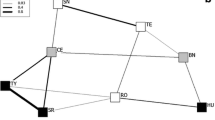
Visualization of the haplotype networks (Fig. 4) showed that there were some haplotype clusters in both Guinea and hamadryas baboons; yet these clusters were not very pronounced. In the Guinea baboons, network clusters only weakly corresponded to the geographic distribution of demes, with many haplotypes being found in several, and even very distant demes (Fig. 4a). Similarly, samples from geographically close demes frequently yielded haplotypes of very distant genetic relationships. The hamadryas baboon network showed a slightly more pronounced geographic clustering, separating most African from Arabian samples, with two distinct Arabian clades (Fig. 4b). Still, haplotypes within regions were very diverse and in some cases very distinct; for instance, some Ethiopian samples clustered closer with Eritrean or Arabian samples than with other samples from Ethiopia. Moreover, several haplotypes in the Arabian clades were shared between demes over a distance of >1000 km (Fig. 4b).
Haplotype network of mtDNA sequences and corresponding distribution of samples of (a) Papio papio and (b) P. hamadryas. One haplotype is represented by one circle and circle size corresponds to haplotype frequency. Branch length is proportional to mutational steps and each dot represents one mutated position. Haplotype color represents the different regions defined for the AMOVA (70 km clustering) depicted on the map.
Discussion
Our results strongly support the hypothesis of female-biased gene flow in Guinea baboons: the female inherited mtDNA marker shows no clear genetic structure that would be consistent with the geographic distribution of our samples. Furthermore, it displays isolation-by-distance, which is consistent with neutral genetic drift driven by dispersal.
Genetic diversity, as inferred from number of haplotypes per species and hd, is comparable between Guinea and hamadryas baboons, both species-wide and at the level of single demes. Species-wide hd is furthermore very similar to that in all other baboon species. However, π is considerably higher in olive, yellow, and chacma baboons compared to hamadryas and Guinea baboons. This probably reflects the more complex evolutionary history of the former three species, which is characterized by multiple events of population isolation and reconnection, leading to deep divergences of haplogroups within these species (Zinner et al. 2009). The very low π in Guinea baboons compared to hamadryas baboons confirms results of a previous study based on a smaller sample size of Guinea baboons from Guinea-Bissau (Ferreira da Silva et al. 2013). The difference in π between Guinea and hamadryas baboons may either be due to a lower effective (female) population size N e or a more recent origin of the species. However, the latter is rather unlikely considering current divergence time estimations that do not suggest a more recent origin of Guinea baboons (Zinner et al. 2013). A smaller effective population size in Guinea baboons could be the result of past demographic changes, e.g., bottlenecks, recent expansion, or a smaller census size, less population substructuring, or a different mating system. Nuclear microsatellite data also suggest that genetic diversity is lower in Guinea baboons than in other baboon species (Ferreira da Silva 2012; Fickenscher et al. 2011). A smaller census size and less substructuring are likely explanations, considering that Guinea baboons have the most restricted distribution of all baboon species (Anandam et al. 2013) and that hamadryas baboons comprise two subpopulations divided by the Red Sea. The similar haplotype diversity between all five species makes us confident that a comparative study of gene flow patterns is feasible and will not be affected by other factors that generally influence the genetic diversity of populations, e.g., differences in female reproductive skew, substructuring of species, differences in demographic history.
MtDNA variation was strikingly similar between Guinea and hamadryas baboons, with the highest proportion of genetic variation being explained by variation within demes. This indicates that female dispersal leads to the accumulation of several mitochondrial haplotypes within a group, a pattern also observed in other female-dispersing species, e.g., chimpanzees (Gagneux et al. 1999; Goldberg and Ruvolo 1997; Morin et al. 1994) and humans (Seielstad et al. 1998). In species with female philopatry, the restriction of female gene flow prohibits the exchange of mitochondrial haplotypes among demes, explaining our results of low genetic variation within demes, but high variation among demes and regions for olive, yellow, and chacma baboons. A study of South African baboons observed similarly low intra-deme variation (Burrell 2008). The higher among-deme and lower among-region variation observed by Burrell (2008) relative to our results for chacma and yellow baboons might be explained by differences in sampling scheme. Our sampling in these species was sparser but included a broader range. Changing the clustering of the AMOVA to larger geographic regions largely eliminates the difference between these two studies.
Burrell (2008) furthermore reports that usually only one haplotype is observed in one specific deme, a pattern that we also observe in yellow and chacma baboons but treated with caution owing to our low intra-deme sampling. In olive baboons we find on average two haplotypes as compared to four and three in demes of hamadryas and Guinea baboons, respectively. Although this difference in intra-deme diversity seems to be rather minor, it is confirmed by the considerably lower intra-deme π in olive relative to hamadryas and Guinea baboons. This suggests that even if several haplotypes are observed within a deme in olive baboons, these are much more closely related than in Guinea and hamadryas baboons.
A comparison of hierarchical distribution of mitogenetic variation between our results and different species with female dispersal and female philopatry, respectively (Modolo et al. 2005; Nietlisbach et al. 2012; Rosenblum et al. 1997) supports our conclusion that Guinea baboons show the typical patterns of a species with female dispersal.
In addition, the less pronounced effect of isolation-by-distance in Guinea and hamadryas baboons is evidence for higher rates of female gene flow in these two species. Although female transfer may be observed on rare occasions in female-philopatric species and has been reported for yellow (Rasmussen, 1981) and olive baboons (Henzi and Barrett, 2003), this apparently has no important impact on the genetic makeup of populations. In hamadryas baboons, the Mantel test revealed the two distinct Arabian clusters that are visible in the haplotype network. These two distinct clusters have already been described in previous studies and are probably a result of the complex colonization history of the Arabian Peninsula by hamadryas baboons (Wildman et al. 2004; Winney et al. 2004).
There was a high degree of shared haplotypes between distant demes in Guinea baboons and, to a lesser extent, in hamadryas baboons. The fact that this pattern is less pronounced in hamadryas baboons could be due to the sampling scheme, which was much patchier in this species. Including more samples from the area between the Ethiopian and the Eritrean clusters may reveal a picture in hamadryas baboons similar to that in Guinea baboons. In both species we observe shared haplotypes over distances of >500 km, a result comparable to that in, e.g., Eastern chimpanzees (Pan troglodytes schweinfurthii; Goldberg and Ruvolo 1997). These shared haplotypes could result from long-distance dispersal, but successive short dispersal events over several generations adding up to larger distances seem to be more likely considering the general biology of baboons. Alternatively, the lack of strong geographic clustering may be explained by shared haplotypes representing ancient diversity and that these ancient lineages are incompletely sorted owing to time constraints. Divergence time estimations and reconstructions of phylogeographic history suggest that Guinea baboons evolved during the same time period as all other baboon species (Zinner et al. 2011, 2013). Consequently, Guinea baboons had as much time as the other species to develop genetic clusters, and this strongly argues against the explanation of incomplete lineage sorting. Further, in female philopatric species one haplotype reaches fixation extremely quickly within demes, causing mitochondrial diversity to disappear rapidly (Hoelzer et al. 1998). This means that polymorphism caused by incomplete lineage sorting would be lost even over short evolutionary timescales, leading to a pattern of mitochondrial variation comparable to what we observed in chacma, yellow, and olive baboons.
Taken together, the results of our study constitute solid evidence for female-biased gene flow in both Guinea and hamadryas baboons, sharply contrasting with the pattern observed in all other baboon species and most mammals. Unfortunately we cannot distinguish between female dispersal in the narrow sense with our mtDNA data set (where single females or small groups of females migrate) and dispersal of social units, e.g., one-male, multifemale groups in hamadryas baboons (Swedell et al. 2011) or parties (Patzelt et al. 2011) in Guinea baboons. This question can be addressed only by long-term observations of individually identified baboons, and at the genetic level by including nuclear markers in future analysis. Whereas direct behavioral observations confirm female dispersal in hamadryas baboons (Swedell et al. 2011), our study is the first indication of a general species-wide pattern of female dispersal in Guinea baboons. These results corroborate the notion that the Guinea baboon’s social system shares some important features with that of hamadryas baboons, suggesting that similar evolutionary forces have acted in their history to distinguish them from all other baboons. Although the details of female dispersal behavior in Guinea baboons remain to be clarified, our study adds to the knowledge of the biology of the genus Papio and improves our understanding of the link between behavior and genetics in primates.
References
Altmann, J., Alberts, S. C., Haines, S. A., Dubach, J., Muruthi, P., Coote, T., Geffen, E., Cheesman, D. J., Mututua, R. S., Saiyalel, S. N., Wayne, R. K., Lacy, R. C., & Bruford, M. W. (1996). Behavior predicts genetic structure in a wild primate group. Proceedings of the National Academy of Sciences of the USA, 93(12), 5797–5801.
Anandam, M. V., Bennett, E. L., Davenport, T. R. B., Davies, N. J., Detwiler, K. M., Engelhardt, A., Eudey, A. A., Gadsby, E. L., Groves, C. P., Healy, A., Karanth, K. P., Molur, S., Nadler, T., Richardson, M. C., Riley, E. P., Roos, C., Rylands, A. B., Sheeran, L. K., Ting, N., Wallis, J., Waters, S. S., Whittaker, D. J., & Zinner, D. (2013). Species accounts of Cercopithecidae. In R. A. Mittermeier, A. B. Rylands, & D. E. Wilson (Eds.), Handbook of the mammals of the world (Primates, Vol. 3, pp. 628–753). Barcelona: Lynx Edicions.
Avise, J. C. (2004). Molecular markers, natural history, and evolution (2nd ed.). Sunderland: Sinauer Associates.
Barrett, L. (2009). A guide to practical babooning: historical, social, and cognitive contingency. Evolutionary Anthropology, 18(3), 91–102.
Barton, R. A., Byrne, R. W., & Whiten, A. (1996). Ecology, feeding competition and social structure in baboons. Behavioral Ecology and Sociobiology, 38, 321–329.
Bensasson, D., Zhang, D.-X., Hartl, D. L., & Hewitt, G. M. (2001). Mitochondrial pseudogenes: evolution’s misplaced witnesses. Trends in Ecology & Evolution, 16(6), 314–321.
Bohonak, A. J. (1999). Dispersal, gene flow, and population structure. The Quarterly Review of Biology, 74(1), 21–45.
Broquet, T., & Petit, E. J. (2009). Molecular estimation of dispersal for ecology and population genetics. Annual Review of Ecology, Evolution, and Systematics, 40(1), 193–216.
Burrell, A. S. (2008). Phylogenetics and population genetics of central African baboons. Department of Anthropology, New York University.
Burrell, A. S., Jolly, C. J., Rogers, J., Phillips-Conroy, J. E., & Disotell, T. R. (2011). Inferring the dispersal behavior of the Kinda baboon (Papio kindae) from multilocus genetic data. American Journal of Physical Anthropology, 144(S52), 99.
Di Fiore, A. (2003). Molecular genetic approaches to the study of primate behavior, social organization, and reproduction. American Journal of Physical Anthropology (Supplement), 37, 62–99.
Excoffier, L., & Lischer, H. E. L. (2010). Arlequin suite ver 3.5: a new series of programs to perform population genetic analyses under Linux and Windows. Molecular Ecology Resources, 10(3), 564–567.
Excoffier, L., Smouse, P. E., & Quattro, J. M. (1992). Analysis of molecular variance inferred from metric distances among DNA haplotypes: application to human mitochondrial DNA restriction data. Genetics, 131(2), 479–91.
Ferreira da Silva, M. J. (2012). Hunting pressure and the population genetic patterns and sex-mediated dispersal in the Guinea Baboon in Guinea-Bissau. Cardiff University.
Ferreira da Silva, M. J., Casanova, C., & Godinho, R. (2013). On the western fringe of baboons distribution: mitochondrial D-loop diversity of Guinea baboons (Papio papio, Desmarest 1820) (Primates: Cercopithecidae) in Coastal Guinea-Bissau, West Africa. Journal of Threatened Taxa, 5, 4441–4450.
Fickenscher, G., Ferreira da Silva, M. J., Patzelt, A., Roos, C., Fischer, J., & Zinner, D. (2011). Genetic evidence for male philopatry in Guinea baboons. Folia Primatologica, 82(6), 365.
Gagneux, P., Wills, C., Gerloff, U., Tautz, D., Morin, P. A., Boesch, C., Fruth, B., Hohmann, G., Ryder, O. A., & Woodruff, D. S. (1999). Mitochondrial sequences show diverse evolutionary histories of African hominoids. Proceedings of the National Academy of Sciences of the USA, 96(9), 5077–5082.
Galat-Luong, A., Galat, G., & Hagell, S. (2006). The social and ecological flexibility of Guinea baboons: implications for Guinea baboon social organization and male strategies. In L. Swedell & S. R. Leigh (Eds.), Reproduction and fitness in baboons: behavioral, ecological, and life history perspectives (pp. 105–121). New York: Springer Science+Business Media.
Goldberg, T. L., & Ruvolo, M. (1997). The geographic apportionment of mitochondrial genetic diversity in east african chimpanzees, Pan troglodytes schweinfurthii. Molecular Biology and Evolution, 14(9), 976–984.
Greenwood, P. J. (1980). Mating systems, philopatry and dispersal in birds and mammals. Animal Behaviour, 28, 1140–1162.
Guschanski, K., Olivieri, G., Funk, S. M., & Radespiel, U. (2006). MtDNA reveals strong genetic differentiation among geographically isolated populations of the golden brown mouse lemur, Microcebus ravelobensis. Conservation Genetics, 8(4), 809–821.
Hall, T. A. (1999). BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symposium Series, 41, 95–98.
Hammond, R. L., Lawson Handley, L. J., Winney, B. J., Bruford, M. W., & Perrin, N. (2006). Genetic evidence for female-biased dispersal and gene flow in a polygynous primate. Proceedings of the Royal Society of London B: Biological Sciences, 273, 479–484.
Hapke, A., Zinner, D., & Zischler, H. (2001). Mitochondrial DNA variation in Eritrean hamadryas baboons (Papio hamadryas hamadryas): life history influences population genetic structure. Behavioral Ecology and Sociobiology, 50, 483–492.
Haus, T., Akom, E., Agwanda, B., Hofreiter, M., Roos, C., & Zinner, D. (2013). Mitochondrial diversity and distribution of African green monkeys (Chlorocebus Gray, 1870). American Journal of Primatology, 75, 350–360.
Henzi, P., & Barrett, L. (2003). Evolutionary ecology, sexual conflict, and behavioral differentiation among baboon populations. Evolutionary Anthropology, 12(5), 217–230.
Hoelzer, G. A., Wallman, J., & Melnick, D. J. (1998). The effects of social structure, geographical structure, and population size on the evolution of mitochondrial DNA: II. Molecular clocks and the lineage sorting period. Journal of Molecular Evolution, 47(1), 21–31.
Jolly, C. J. (1993). Species, subspecies and baboon systematics. In W. H. Kimbel & L. B. Martin (Eds.), Species, species concepts and primate evolution (pp. 67–107). New York: Plenum Press.
Jolly, C. J. (2007). Baboons, mandrills, and mangabeys: afro-papionin socioecology in a phylogenetic perspective. In C. J. Campbell, A. Fuentes, K. C. MacKinnon, M. Panger, & S. K. Bearder (Eds.), Primates in perspective (pp. 240–251). New York: Oxford University Press.
Jolly, C. J. (2009). Fifty years of looking at human evolution: backward, forward, and sideways. Current Anthropology, 50(2), 187–199.
Jolly, C. J., & Phillips-Conroy, J. E. (2006). Testicular size, developmental trajectories, and male life history strategies in four baboon taxa. In L. Swedell & S. R. Leigh (Eds.), Reproduction and fitness in baboons: behavioral, ecological, and life history perspectives (pp. 257–275). New York: Springer Science+Business Media.
Kappeler, P. M., & Van Schaik, C. P. (2002). Evolution of primate social systems. International Journal of Primatology, 23(4), 707–740.
Kingdon, J. (1997). The kingdon field guide to African mammals. London: Academic Press.
Librado, P., & Rozas, J. (2009). DnaSP v5: a software for comprehensive analysis of DNA polymorphism data. Bioinformatics, 25(11), 1451–1452.
Mantel, N. (1967). The detection of disease clustering and a generalized regression approach. Cancer Research, 27(2), 209–220.
Melnick D. (1987). The genetic consequences of primate social organization: a review of macaques, baboons, and vervet monkeys. Genetica, 73(1–2), 117–135.
Miller, M. P. (2005). Alleles in space (AIS): computer software for the joint analysis of interindividual spatial and genetic information. The Journal of Heredity, 96(6), 722–724.
Modolo, L., Salzburger, W., & Martin, R. D. (2005). Phylogeography of Barbary macaques (Macaca sylvanus) and the origin of the Gibraltar colony. Proceedings of the National Academy of Sciences of the USA, 102(20), 7392–7397.
Morin, P. A., Moore, J. J., Chakraborty, R., Jin, L., Goodall, J., & Woodruff, D. S. (1994). Kin selection, social structure, gene flow, and the evolution of chimpanzees. Science, 265(5176), 1193–1201.
Nei, M. (1987). Molecular evolutionary genetics. New York: Columbia University Press.
Nietlisbach, P., Arora, N., Nater, A., Goossens, B., Van Schaik, C. P., & Krützen, M. (2012). Heavily male-biased long-distance dispersal of orang-utans (genus: Pongo), as revealed by Y-chromosomal and mitochondrial genetic markers. Molecular Ecology, 21(13), 3173–3186.
Nsubuga, A. M., Robbins, M. M., Roeder, A. D., Morin, P. A., Boesch, C., & Vigilant, L. (2004). Factors affecting the amount of genomic DNA extracted from ape faeces and the identification of an improved sample storage method. Molecular Ecology, 13(7), 2089–2094.
Patzelt, A., Zinner, D., Fickenscher, G., Diedhiou, S., Camara, B., Stahl, D., & Fischer, J. (2011). Group composition of Guinea baboons (Papio papio) at a water place suggests a fluid social organization. International Journal of Primatology, 32(3), 652–668.
Pusey, A. E., & Packer, C. (1987). Dispersal and Philopatry. In B. B. Smuts, D. L. Cheney, R. M. Seyfarth, R. W. Wrangham, & T. T. Struhsaker (Eds.), Primate societies (pp. 250–266). Chicago: Chicago University Press.
Rasmussen, D. R. (1981). Communities of baboon troops (Papio cynocephalus) in Mikumi National Park, Tanzania: a preliminary report. Folia Primatologica, 36, 232–242.
Robinson, G. E., Fernald, R. D., & Clayton, D. F. (2008). Genes and social behavior. Science, 322(5903), 896–900.
Roeder, A. D., Archer, F. I., Poiner, H. N., & Morin, P. A. (2004). A novel method for collection and preservation of faeces for genetic studies. Molecular Ecology Notes, 4(4), 761–764.
Rosenblum, L. L., Supriatna, J., & Melnick, D. J. (1997). Phylogeographic analysis of pigtail macaque populations (Macaca nemestrina) inferred from mitochondrial DNA. American Journal of Physical Anthropology, 104, 35–45.
Seielstad, M. T., Minch, E., & Cavalli-Sforza, L. L. (1998). Genetic evidence for a higher female migration rate in humans. Nature Genetics, 20(3), 278–280.
Seyfarth, R. M., & Cheney, D. L. (2012). The evolutionary origins of friendship. Annual Review of Psychology, 63, 153–177.
Sigg, H., Stolba, A., Abegglen, J. J., & Dasser, V. (1982). Life history of hamadryas baboons: physical development, infant mortality, reproductive parameters and family relationships. Primates, 23(4), 473–487.
Silk, J. B. (2002). Kin selection in primate groups. International Journal of Primatology, 23(4), 849–875.
Swedell, L. (2011). African Papionins: diversity of social organization and ecological flexibility. In C. J. Campbell, A. Fuentes, K. C. MacKinnon, S. K. Bearder, & R. M. Stumpf (Eds.), Primates in perspective (2nd ed., pp. 241–277). New York: Oxford University Press.
Swedell, L., Saunders, J., Schreier, A., Davis, B., Tesfaye, T., & Pines, M. (2011). Female “dispersal” in hamadryas baboons: transfer among social units in a multilevel society. American Journal of Physical Anthropology, 145(3), 360–370.
Teacher, A. G. F., & Griffiths, D. J. (2011). HapStar: automated haplotype network layout and visualization. Molecular Ecology Resources, 11(1), 151–153.
Thalmann, O., Hebler, J., Poinar, H. N., Pääbo, S., & Vigilant, L. (2004). Unreliable mtDNA data due to nuclear insertions: a cautionary tale from analysis of humans and other great apes. Molecular Ecology, 13(2), 321–335.
Tung, J., Akinyi, M. Y., Mutura, S., Altmann, J., Wray, G. A., & Alberts, S. C. (2011). Allele-specific gene expression in a wild nonhuman primate population. Molecular Ecology, 20(4), 725–739.
Tung, J., Alberts, S. C., & Wray, G. A. (2010). Evolutionary genetics in wild primates: combining genetic approaches with field studies of natural populations. Trends in Genetics, 26(8), 353–362.
Wildman, D. E., Bergman, T. J., al-Aghbari, A., Sterner, K. N., Newman, T. K., Phillips-Conroy, J. E., Jolly, C., & Disotell, T. R. (2004). Mitochondrial evidence for the origin of hamadryas baboons. Molecular Phylogenetics and Evolution, 32(1), 287–296.
Winney, B. J., Hammond, R. L., Macasero, W., Flores, B., Boug, A., Biquand, V., Biquand, S., & Bruford, M. W. (2004). Crossing the Red Sea: Phylogeography of the hamadryas baboon, Papio hamadryas hamadryas. Molecular Ecology, 13(9), 2819–2827.
Zinner, D., Buba, U., & Roos, C. (2011). Pan-African voyagers: the phylogeography of baboons. In V. Sommer & C. Roos (Eds.), Primates of Gashaka: socioecology and conservation in Nigeria’s biodiversity hotspot (pp. 267–307). New York: Springer Science+Business Media.
Zinner, D., Groeneveld, L. F., Keller, C., & Roos, C. (2009). Mitochondrial phylogeography of baboons (Papio spp.)—Indication for introgressive hybridization? BMC Evolutionary Biology, 9, 83.
Zinner, D., Wertheimer, J., Liedigk, R., Groeneveld, L. F., & Roos, C. (2013). Baboon phylogeny as inferred from complete mitochondrial genomes. American Journal of Physical Anthropology, 150, 133–140.
Acknowledgments
We are grateful to James Higham, Lauren Brent, and Amanda Melin for inviting us to contribute to this special issue of International Journal of Primatology. We thank Joanna Setchell, Lauren Brent, and two anonymous reviewers for very helpful comments on earlier versions of the manuscript. We thank Thomas Butynski from the KKWRC for contributing samples from Saudi Arabia; Karim Nasher for samples from Saudi Arabia and Yemen; and Dawit Berhane, Tobias Kopp, Anja Ebenau, Kurt Hammerschmidt, Mamadou Samba Barry, Salian Traore, Amadou Sadio Balde, Jacky Bassene, Cheickh Y. S. Sané, Issakha Ndiaye, and all local field assistants for their assistance in collecting samples. We acknowledge AD (Acção para o Desenvolvimento) and CHIMBO NGOs, the field assistants M. Soares and M. Turé, and the park guides and guards of Cantanhez and Cufada Park for fieldwork logistical support in Guinea-Bissau and The Wild Chimpanzee Foundation for logistical support in Guinea. We also thank Christiane Schwarz for help with laboratory work. We acknowledge the local authorities for their support and permits: the Ministère de l’Environnemnt et de la Protection de la Nature and the Direction des Parcs Nationaux (Senegal); the Opération du Parc National de la Boucle du Baoulé and the Ministère de l’Environnement et de l’Assainissement (Mali); the Office Guinéen de la Diversité Biologique et des Aires Protégées and the Ministère de L’Environnement, des Eaux et Forêts (Guinea); the Ethiopian Wildlife Conservation Organization; IBAP (Institute for Biodiversity and Protected Areas) and DGFF (Direcção Geral de Florestas e Fauna) (Guinea-Bissau); and the Ministère Délégue auprès du Premier Ministre, Chargé de l’Environnement et du Développement Durable (Mauritania). G. H. Kopp was supported by the German Academic Exchange Service (DAAD), The Leakey Foundation, and the German Primate Center. D. Zinner was supported by Grant DFG ZI 548/ 3–1 from the German Science Foundation. M. J. Ferreira da Silva worked under a Portuguese Foundation for Science and Technology (FCT) Post-Doctoral Scholarship (Ref. SFRH/BPD/88496/2012). J. C. Brito was supported by a FCT-contract (Programme Ciência 2007) and fieldwork supported by a grant from National Geographic Society (8412–08) and by FCT (PTDC/BIA-BEC/099934/2008) through the EU programme COMPETE.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(DOCX 67.1 kb)
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution License which permits any use, distribution, and reproduction in any medium, provided the original author(s) and the source are credited.
About this article
Cite this article
Kopp, G.H., Ferreira da Silva, M.J., Fischer, J. et al. The Influence of Social Systems on Patterns of Mitochondrial DNA Variation in Baboons. Int J Primatol 35, 210–225 (2014). https://doi.org/10.1007/s10764-013-9725-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10764-013-9725-5