Abstract
Studying aquatic population dynamics using spatio-temporal monitoring data is associated with a number of challenges and choices. One can let several samples represent the same population over larger areas, or alternatively model the dynamics of each sampling location in continuous space. We analysed the spatio-temporal population dynamics of six phytoplankton taxa in the Baltic Sea applying multivariate state-space models with first-order density dependence. We compared three spatial scales and three models for spatial correlation between predefined subpopulations using information theoretic model selection. We hypothesised that populations close to each other display similar dynamic properties and spatial synchrony decreasing with the distance. We further hypothesize that intermediate-scale grouping of data into subpopulations may parsimoniously represent such dynamics. All taxa showed constant density dependence across space and strong spatial synchrony, consistently requiring a parameter for spatial correlation whenever models included several population states. The most parsimonious spatial structure varied between taxa, most often being one panmictic population or ten intercorrelated population states. Evidently, longer time-series, containing more information, provide more options for modelling detailed spatio-temporal patterns. With a few decade-long plankton time-series data, we encourage determining the appropriate spatial scale on biological grounds rather than model fit.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Time-series data on species’ relative abundances are commonly gathered for monitoring trends and patterns in biological populations and communities, as well as for identifying intrinsic and extrinsic drivers of population dynamics (Turchin, 1995, 2003). Usually, these data consist of counts or density estimates from multiple locations, which show more or less synchronous fluctuations (Ranta et al., 1997, 1999). Spatial synchrony between subpopulations can arise if they are connected by dispersal, or naturally, if sampling points all represent one panmictic population. However, isolated subpopulations also show synchronous dynamics if they have identical density dependence and are subject to the same environmental forcing (Moran, 1953). This phenomenon, known as Moran effect, has later been shown to produce various degrees of synchrony even under more relaxed assumptions, such as somewhat different density-dependent dynamics and non-identical but spatially autocorrelated environment (Ranta et al., 1997, 1999; Engen & Sæther, 2005; Sæther et al., 2007).
In order to analyse large-scale temporal patterns and processes, it is tempting, and not uncommon to let observations from a larger area represent one larger population—an approach which has been applied for trend analysis regionally in many systems and also on plankton in our study area, the Baltic Sea (e.g. Suikkanen et al., 2007). Data from different stations may be treated as replicates of one-state variable, or analysed as a time-series of temporal averages (pooled from several stations). These approaches provide information about sampling noise, which facilitate analyses and summarising of results. On the other hand, they may ignore valuable information about spatial patterns and local dynamic properties, which also contain some implicit information about the contribution of observation error (Dennis et al., 2010). To make well-informed decisions on the analysis of monitoring data, we need to know to which extent population structure can be inferred from the data analysed, and what consequences the choice of scale may have for the parameters estimated (Ward et al., 2010).
State-space models are quantitative dynamic models that explicitly separate between stochastic population fluctuations not captured by the model prediction (process error), and additional stochasticity in the data due to noisy sampling and counting procedures (observation error) (Durbin & Koopman, 2012). These type of models are frequently needed for making inference about density dependence (e.g., Knape, 2008), environmental effects (Lindén & Knape, 2009), trends (Humbert et al., 2009) and in population viability analysis (Tolimieri et al., 2017). It is not uncommon that spatio-temporal population-monitoring data lack replicates for a given location and point in time, which may hamper our ability to estimate observation error variance at the desired scale of observation, and hence, to fit state-space models (Dennis et al., 2010). Observation error is a common issue in marine time-series (Hampton et al., 2013; Scheef et al., 2012), and quality control information from all work stages (from sampling to taxonomical analysis) needs to be considered when plankton data are used for analyses (Zingone et al., 2015).
Our goals are primarily exploratory: we aim to investigate suitable approaches for modelling spatio-temporal population dynamics using practically unreplicated monitoring data. As a model system, we study six common phytoplankton taxa in the Baltic Sea, applying multivariate autoregressive state-space models on three spatial scales, with one-, five-, and ten-population state variables. Our set-up is fairly similar to that of Ward et al. (2010) and Holmes et al. (2012). We hypothesise that (1) the stations will show clear synchrony, presumably due to the Moran effect. (2) Synchronous fluctuations can be modelled on a coarse scale, treating some or all of the stations as representing the same population, without meaningful loss of information. Complex interactions of variables like temperature and nutrients are suggested to cause Moran effects in plankton (Defriez & Reuman, 2017), and the Baltic Sea basins in the study area are all distinct in terms of temperature, nutrient concentrations and salinity (Andersen et al., 2017; Snoeijs-Leijonmalm & Andrén, 2017). We thus hypothesize that our models with five subpopulations corresponding to these basins should be particularly parsimonious. (3) In the cases with several different subpopulations, we expect that synchrony decreases with distance. (4) When applicable, we expect that the subpopulations can be assumed to have the same density dependence parameters due to ecological and genetic similarities. Alternatively, if our prediction does not hold, environmental gradients in, e.g. salinity or basin-specific communities, may result in spatially variable density dependence parameters.
Methods
Study area
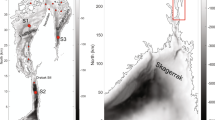
The Baltic Sea is a 442 000 km2 brackish water basin, with only a narrow connection to the North Sea through Kattegat and Skagerrak straits. This leads to varied salinity conditions with highest salinity in the south and decreasing salinities towards north and to the east, with a total salinity range from ca. 2 to 30 practical salinity units (PSUs) for the whole Baltic Sea. The ten chosen monitoring stations are located in the Bothnian Bay (BB), Bothnian Sea (BS), Sea of Åland (AS), Gulf of Finland (GoF) and the Baltic Proper (BP) (Fig. 1). All these sub-basins have distinct differences in mean salinity and depth, and some of them are additionally separated by sills (BS, BB and AS). All stations are pelagic with depths exceeding 80 metres. The salinity within the study area ranges from ca. 3 to 7 PSU.
Monitoring data
We study six phytoplankton taxa, identified either to the genus or species level: Chrysochromulina spp., Hemiselmis spp., Plagioselmis prolonga Butcher 1967, Pseudopedinella tricostata (Roukhiyajnen) Thomsen 1988, Pyramimonas spp. and Teleaulax spp.—with the finest taxonomic resolution resulting in meaningful time-series (nomenclature according to Hällfors, 2004). These are all observed over large areas and considered to be abundant, but little is known about their spatial population structure and dynamics.
The Finnish national monitoring data have been collected since 1979 as part of the Helsinki Commission’s (HELCOM) COMBINE program, according to the methods described in the Manual for Marine Monitoring (HELCOM, 2017). Phytoplankton samples were taken as integrated samples from 0 to 10 metres and preserved with acid Lugol solution. The samples were analysed using an inverted microscope. For each year, the same person analysed and counted the samples for that year for all ten stations. Cells were divided into taxon-specific size classes and then transformed into biovolumes using taxon-specific formulae and size class-specific cell measurements (Olenina et al., 2006). Biovolumes were converted to biomass by assuming that the density was one. The amount of plankton is reported in biomass (wet weight) μg l−1.
Monitoring data, spanning 37 years, was obtained from two different databases, Sumppu (1979–2007) and Hertta (2008–2015, www.ymparisto.fi/oiva). We focus on a seasonal window spanning from the 1 July to the 29 September, as the monitoring has focused on measuring the late summer phytoplankton. The mean Julian day for data gathered during this period is 229 (SD 13.14). All six studied taxa show a reasonably flat phenological pattern of occurrence during the sampled time interval, i.e., variation in sampling date should not introduce severe bias or noise.
Generally, only one sampling event per station and year was carried out during the chosen seasonal window. Exceptions are three years with multiple samples at LL7 and one year with two samples at LL3A. The monitoring stations were chosen based on sampling frequency, disregarding stations that had not been sampled for more than 25% of the full time period (less than 10 years).
Data compilation
As the data come from two different databases, we had to harmonise the data. If the biomass of a taxon was divided into multiple size classes, they were summed to create one taxon-specific biomass for each sample.
We aim to preserve biological relevance by keeping the taxonomic level as accurate as possible, by keeping it as close as possible to the species level, and by choosing distinctive taxa. The taxa chosen for the analysis were required to be present 40% of the time at each of the investigated stations. As the aim is to model already established populations, the earliest possible starting year of each time-series was chosen based on three criteria: (1) non-zero data for the taxon were available for at least eight of the ten stations, (2) if data were missing, or the taxon was not reported in the data that year, there had to be non-zero data the preceding or following year, and (3) the stations with missing data could not be located in the same sub-basin. In three cases, the length of the time-series was influenced by changes in the accuracy of identification through the years. The time-series for P. prolonga, Hemiselmis spp. and Teleaulax spp. are shorter as they were assigned to the order Cryptomonadales prior to 1990 (ESM Table 1).
Zeros in the middle of the time-series were treated as missing values. This was considered the most conservative option as species or taxa might some years have been ascribed to a less accurate taxonomic level (e.g. unknown flagellates), and reported as zero for the taxa studied here. When present, zeros tended to occur simultaneously at all stations in the same year, strongly suggesting inconsistent identification accuracy for some of these fairly common and widespread species. Hence, it is likely that the focal taxa were still present in years with zeros. All time-series were log-transformed prior to analysis.
The statistical model
The applied model consists of an observation and a process component (Eqs. 1 and 3, respectively) with appurtenant variance structures (Eqs. 2 and 4, respectively), and was applied for one taxon at a time. Defined for a focal time step t, the model for a taxon is:
The natural logarithm of estimated biomasses (observed) at all the stations is the multivariate response (vector yt in Eq. 1), which are linked to the underlying hidden states, xt through the matrix Z, the scaling vector a and multinormal observation errors (vector υt in Eqs. 1 and 2). The numbers of underlying hidden states are defined by Z. The observation errors υt are assumed to be independent with equal variances for all population states (the diagonal in variance–covariance matrix R).
The diagonal of matrix B contains the parameters for density dependence, defining how the subpopulation states xt are linked to the situation in previous year xt–1. When the value of B is unity, the model behaves like exponential growth or a random walk, i.e., no density dependence is present. Population growth is considered negatively density dependent when 0 < B < 1 (undercompensatory dynamics), B = 0 (exactly compensatory dynamics), or B < 0 (overcompensatory dynamics), smaller values of B indicating stronger density dependence. On the off-diagonal, all elements of B are zero. The row vector u represents the growth rates of the populations, adjusting the averages. The process errors ωt are multivariate normally distributed with mean zero and variance–covariance matrix Q.
By using different structures for Q we make various assumptions about the correlation between the stations in the unexplained population variation. As a general description of Q, it can be expressed in terms of a common process error variance σ2, the correlation per distance ρ, and a distance matrix D (with zero diagonal), as
We used three options for modelling the correlation between the populations: (1) no correlation (NC), with ρ = 0, which implies zero correlation between the populations; (2) compound symmetry (CS), giving equal correlation between the populations; and (3) an autoregressive covariance structure (AR), where D is the actual distance between the populations, resulting in a correlation declining exponentially with distance between the stations. For the model with one panmictic population, Q is scalar and equal to σ2.
All the data analysis were conducted in the R-environment version 3.4.3 (R Core Team, 2017), using the MARSS (version 3.9) package for fitting multivariate autoregressive Gaussian state-space models (described in Holmes et al., 2012, 2018). To be able to fit all desired model structures, we optimised the process error parameters separately using a combination of MARSS and the optimx function (Nash & Varadhan, 2013) (more detailed description in ESM). Apart from MARSS, other packages that were used include plyr (Wickham, 2016a), reshape2 (Wickham, 2016b), geosphere (Hijmans, 2017) and ggplot2 (Wickham & Chang, 2016).
Competing hypotheses and model selection
By modifying Z, we compared models, where samples from all monitoring stations were considered; those sampled either from one large population, from five subpopulations, or from ten separate subpopulations (Fig. 1). The division of the stations into five populations was done based on salinity and by accounting for sills between sub-basins (Snoeijs-Leijonmalm & Andrén, 2017). However, while alternative criteria of division could have been applied, we stress that the chosen scenario can also be interpreted as an arbitrary intermediate spatial scale, between ten-state variables and a single one. In addition, we also compared models assuming equal or spatially variable density dependence (along the diagonal of B). Considering the different options for Q, Z and B, we investigated in total 13 unique model structures for each taxon (Table 1). Examples and a more detailed account on similar investigations into spatial structure can be found for, e.g. salmon (Hinrichsen & Holmes, 2009), sea lions (Ward et al., 2010) and harbour seals (Holmes et al., 2018).
The structure for a and u were constant for all the models, with station-specific growth rates and a allowing different average biomass levels at the different monitoring stations. Vector a also enables us to define the few sporadic replicates from stations LL7 and LL3A. Inclusion of the replicates in the analysis, effectively prevents the observation error variance (R) from going to zero in the optimization procedure, which is a common problem in state-space modelling (Dennis et al., 2010). The initial values of the state variables (at time step t = 1) were given prior distributions with the mean being the average of the initial time step(s) of the reference time-series and the variance being σ2 (at the diagonal of Q).
The model support for the resulting 13 candidate models for each taxon (see ESM Table 2) was evaluated in terms of parsimony according to their Akaike information criterion corrected for small sample size, AICc (Burnham & Anderson, 2002). To compare models and scenarios between taxa we further calculated Akaike weights and evidence ratios to visualise the results. In the state option comparison (Fig. 2), the panmictic option Akaike weight was multiplied by six to make it comparable to the five- and ten-state options. In comparisons between the spatial correlation scenarios in Q, the panmictic option was excluded (Fig. 5).
The summed Akaike weights for models with different numbers of underlying states. To make the approaches comparable, the Akaike weight of the one-state option was multiplied by six (the number of models with five and ten states) before rescaling. The evidence ratios (ER) for the best vs. the second-best option are displayed in the upper left corner
Additional analyses
To further investigate the plausible role of observation noise in the model selection results, we successively added an increasing amount of simulated noise to the time-series of the taxa, where a multistate population model was favoured by AICc. As the level of observation noise increased, we expected the panmictic model to eventually become the more parsimonious model. This was carried out for the five-state models for P. tricostata, and for the ten-state models for Chrysochromulina spp. both with autoregressive process error structure and one shared density dependence term. The details of this simulation are presented in the ESM.
To assess goodness of fit, we investigated the normality of the residuals in all models within two units of AICc difference (ΔAICc). We applied quantile–quantile plots and the Shapiro-Wilks test for normality.
Results
In general, the fitted models captured the annual and obviously spatially synchronous population changes fairly well. In line with our expectations, all studied taxa showed marked spatial synchrony between monitoring stations, modelled either as correlation between the population states, or as one population state. Accordingly, multistate models with uncorrelated process error consistently performed poorly in terms of AICc (ESM Table 2).
The most parsimonious state option for Hemiselmis spp., P. prolonga, and Pyramimonas spp. was that of one panmictic population (Fig. 2, ESM Table 2). While the panmictic model was not the most parsimonious for Teleaulax spp., it had more weight in comparison to the other options (Fig. 2). For Chrysochromulina spp., the results are suggestive of a ten-state approach and for P. tricostata the level of support was very similar for different numbers of states. Further, the estimated process error correlations for the models within two ΔAICc units for P. tricostata were high (model four, six and twelve: 0.963, 0.976 and 0.921, respectively). The high correlation makes it difficult to judge anything regarding how correlation changes with distance. This is also the case for Pyramimonas spp., for which the best model overall was the panmictic one, and the second-best one was a ten-state model with autoregressive spatial structure with high correlation (0.94). While the five-state model was the best among several competing models within two ΔAICc units for P. tricostata and Teleaulax spp. (ESM Table 2), overall, in contrast to our suggestion, we find no particular support for an intermediate numbers of states (five basin-specific states) being more parsimonious than one or ten states (Fig. 2). By simulation, we could confirm that one-state population models are indeed favoured in noisier systems (ESM Fig. 3). The panmictic population was favoured over the best multistate options with the addition of 2.5–3 times the observation noise of the model estimate.
For three taxa, a single most parsimonious model emerged (Fig. 3), while the other taxa had two, three and five competing models within two units of ΔAICc (Fig. 4). There was no consistency in which single model was the most parsimonious for most taxa. A generally well-performing option was model twelve, with ten-state variables, equal density dependence, and autoregressive spatial correlation in the process error (Q). This model was the best in terms of AICc only for Chrysochromulina spp., but within 1.7 units of AICc from the best model in P. tricostata, Pyramimonas spp. and Teleaulax spp.
Other models that gained support were model four, six and ten (Table 1). The best models for P. tricostata consisted of models six (the best model), twelve (ΔAICc 1.10) and four (ΔAICc 1.40). All mentioned models have equal density dependence. Both model four and six have five-state variables and show spatial correlation in the process error, compound symmetry and autoregressive error structure, respectively. Models ten and twelve are the corresponding pair of models but with ten states. The best models for Teleaulax spp. consisted of models one (best), four, six, ten and twelve (all with ΔAICc ≤ 0.82).
For models with multiple subpopulations, the favoured type of spatial correlation structure in the process error was not consistent between taxa (Fig. 5). The autoregressive structure, where correlation decreases with distance, was clearly superior in Chrysochromulina spp. and Pyramimonas spp. and indicative for P. tricostata, while it was impossible to distinguish between autoregressive and compound symmetry options for the others. Notably, it was never really worse than compound symmetry. For Hemiselmis spp., P. prolonga and Teleaulax spp., the result is expected, as the model with one panmictic population was superior to the models with several states and either spatial correlation structure.
The summed Akaike weights for models with different covariance structure (matrix Q) of the process error: NC no spatial correlation, CS compound symmetry, AR autoregressive. The results are based only on the models with five and ten states. The evidence ratios (ER) for the best versus the second best option are displayed in the upper right corners of the panels
For all taxa, the best models had one shared density dependence parameter on the diagonal of B. This is also the case when looking at the Akaike weights grouped by the number of state variables for most taxa (ESM Fig. 1; ESM Fig. 2). However, the support is not conclusive for Hemiselmis spp. with five populations and P. prolonga with ten populations. The parameter estimates for density dependence ranged between 0.116 and 0.825 in the models within 2 units ΔAICc (Table 2). Three species showed evidence for density dependence, Chrysochromulina spp. and P. tricostata having undercompensatory dynamics and Pyramimonas spp. close to exactly compensatory dynamics. Hemiselmis spp., P. prolonga, and Teleaulax spp. all had point estimates corresponding to weak or moderate undercompensatory density dependence, but accounting for parameter uncertainty, the exponential growth model (diag(B) = 1; no density dependence) cannot be ruled out.
The model residuals did not significantly differ from a normal distribution, with the exception of two models in one species. These were models ten and twelve for Teleaulax spp.
Discussion
An evident result is strong spatial synchrony in population states, which has to be accounted for when modelling this type of data. On the annual scale, the environmental variation is likely to be similar across the study area, leading to a Moran effect. Possible candidate variables causing Moran effects are temperature and salinity. In the Curonian Lagoon, Jaanus et al. (2011) reported that phytoplankton biomass was negatively affected by salinity, while relationships between the dominant phytoplankton groups remained. Another mechanism that can produce synchrony is dispersal. The sea is a dynamic connected environment in constant movement. Earlier it was widely accepted that plankton can be anywhere as long as appropriate resources are available (‘Baas-Becking’s tenet’; de Wit & Bouvier, 2006), in essence having endless dispersal potential. More recently, microorganisms have been shown to exhibit biogeographical patterns, some at least influenced by the environment (Martiny et al., 2006). In practice, the range of dispersal for individual phytoplankton cells is quite small due to their small size, low velocities (Bauerfeind et al., 1986) and short life-span, and thus their horizontal displacement is tied to physical forcing. As the surface currents in the Baltic Sea are relatively weak (Leppäranta & Myrberg, 2009), one individual is unlikely to survive from one monitoring station to the next—with the shortest distance between monitoring stations being 100 km. When Defriez & Reuman (2017) investigated the geography of synchrony in plankton using remote-sensed chlorophyll a data, they also suggested that Moran effect was the more likely mechanism of synchronisation in plankton. We therefore conclude that the observed spatial synchrony must in practice be mediated by the Moran effect rather than by dispersal.
The taxa investigated showed no consistent patterns of population structure. The results suggest that the most appropriate number of states is the one-state option, with four taxa supporting this approach, while Chrysochromulina spp. showed some support for ten states and P. tricostata being inconclusive. Hence, it seems that treating data from nearby monitoring stations as replicates of the same population is often not a bad alternative. This is also supported by the observed very high and uniform correlations between the stations’ process errors in the multiple-state models of these taxa. However, for Chrysochromulina spp., P. tricostata and Pyramimonas spp., higher resolution spatial models performed well, also showing a more moderate correlation in process errors, decreasing by distance. In fact, these three species had longer time-series compared to the rest (Table 1). It is not surprising that species with more data allows for describing more detailed patterns.
Lower biomass of a selected taxon in a sample means that there were fewer cells in a sample, and this may cause lower precision in the estimated biovolume and hence more noise (observation error). We aimed to minimize this by selecting common, numerous taxa with the biovolume of one cell usually ranging only from 14 to 723 µm−3 (Olenina et al., 2006). From the monitoring samples, at least 50 counting units (e.g. cells) of each dominating taxon were counted, and the total count was at least 500 units (HELCOM, 2017), which causes that the counting accuracy for rare taxa is not as good as for common taxa. This is common practice in microscopy based data aiming to monitor plankton communities. For rare, especially the large-sized taxa, the observation error for a biomass result can be much higher than for common small-sized taxa, which usually are more numerous in the monitoring samples of the area (e.g. Suikkanen et al., 2013; Kuosa et al., 2017).
A fundamental question is whether all taxa would be best modelled using a ten-state approach, if the time-series would be long and informative enough. Noisier data and shorter time-series will more likely favour a lower resolution model with fewer states and parameters. As shown in the simulations with added noise in ESM Fig. 3, also the taxa with models favouring multistate options will revert to one panmictic population when 2.5–3 times more observation noise is added. One may ask whether we are misidentifying the best model, with the risk of drawing incorrect conclusions. While this is a problem if one is genuinely identifying population structure, the problem can also be seen merely relating to the scale of investigation, where the study questions and conclusions drawn are tied to that scale. In cases where large-scale population trends and compound-symmetric spatially synchronous patterns dominate (e.g. due to Moran effects), the one-state approach may indeed be detailed enough, or the most parsimonious alternative, even though the population is not in reality panmictic.
Several earlier long-term studies on Baltic Sea phytoplankton and indicator studies utilising phytoplankton-monitoring data have focused on certain sub-basins, where they have grouped the same monitoring data in similar fashion as our intermediate-state option (Suikkanen et al., 2007, 2013; Lehtinen et al., 2016; Kuosa et al., 2017). It is curious that the intermediate option gained low support, since salinity is known to affect the distribution of plankton species (Wasmund et al., 2011), and as there is a notable difference in salinity between the northernmost and southernmost basins. Noticeable, the scale of population dynamics is a different question compared to community composition or species distribution. The five-state option had the most support for the inconclusive taxa P. tricostata and Teleaulax spp., which regardless of model all had high spatial correlation. Chrysophyceae, to which P. tricostata belongs, have been shown to prefer low salinities (Wasmund et al., 2011).
For the taxa that did not have support for the panmictic state option, there was always a spatial structure present either as a spatially autoregressive or a compound symmetry error structure. When looking at the wet weight biomass in the whole Baltic Sea, Olli et al. (2013) observed positive spatial autocorrelation up to 500 km for the whole Baltic Sea and up to 50 km when looking at one basin, the Gulf of Finland. Some plankton species are known to form patches of tens of kilometres (Eppley et al., 1984). Also when looking at genetic and geographical distances in Skeletonema marinoi, Sjöqvist et al. (2015) did not see a change with distance within the Baltic Sea.
An obvious result was the spatially uniform effects of density dependence, applying for both the basin level (five-state variables) and station level (ten-state variables) analyses. Apparently, the ecology of these species and the drivers of population dynamics are similar throughout the study area, despite the gradients in salinity, variable temperatures, different nutrient availabilities and variable winter conditions. This will also be mitigated, and the regional variation will be decreased by including separate growth rates and levels (a and u in Eqs. 1 and 3). The similar ecology and drivers of the population dynamics for the species could be due to genetics, dominating Moran effects, or a combination of these. A in vitro study on Skeletonema marinoi Sarno & Zingone has lent support to the fact that species can have considerably broader salinity spectra then what is covered by our study area (Sjöqvist et al., 2015). Different subpopulations between areas with high salinity differences have been identified using genetic methods for S. marinoi; however, these patterns were at larger spatial scale and the differences were observed between the Baltic Sea and the North Sea (Sjöqvist et al., 2015). This is also in accordance with other studies conducted in the Baltic Sea on macroorganisms, where the genetic variability within the Baltic compared to the North Sea has been considered to be lower (Johannesson & André, 2006). There are also other studies suggesting a cut-of-point at the transition zone between the Baltic and the North Sea (Wennerström et al., 2013). Naturally, the results presented here may not apply for other areas with larger distances, more spatially isolated sub-basins or more spatially variable conditions. Neither can we guarantee that the result applies to all phytoplankton taxa.
On a large scale, density dependence, as described by the autoregressive coefficient, will indeed describe the statistical properties of the time-series. However, it may be different (weaker) at larger scales and perhaps not accurately describing the commonly assumed biological processes behind it, such as intraspecific competition. Then, density dependence is merely technically summarising the population autocorrelation in a system (Turchin, 1995).
It is often notoriously difficult to partition the variance of Q and R. It is not uncommon with a multimodal likelihood surface, where one or both variances show a spurious peak at zero, the realistic peak being too shallow to be detected (Knape, 2008; Auger-Méthé et al., 2016). In an ideal situation, we would have multiple replicates for each time step, which would considerably help partitioning the variances (Dennis et al., 2010). By already including very few replicates effectively, as was done in this study, we prevent the observation error variance from approaching zero. On the other hand, it is not necessarily optimal to use multiple samples from only few years and monitoring stations (Gulf of Finland) as replicates, and to extrapolate this information to the whole system, as the observation error variance may differ between the stations and/or sub-basins.
We are aware that the taxa selected are not representative for the whole phytoplankton community across the Baltic Sea, which includes ca. 2000 species (Hällfors, 2004). Rather our study shows a snapshot of the kinds of patterns that emerge from this kind of data and how the patterns may vary between taxa. In order to investigate our questions on the large spatial scale applied, we simply were restricted to use the species for which sufficient data are available. With a more limited area, it would be possible to include more species. Our study contributes by providing a better understanding of patterns in phytoplankton-monitoring data and helps us to identify suitable modelling approaches, new potential uses and weaknesses.
Conclusions
Our results suggest high spatial synchrony in phytoplankton population fluctuations in the Baltic Sea, implying that spatial correlation must not be ignored in statistical analyses. For many taxa, treating the data as replicates from one panmictic population will not lead to relevant loss of information, as the model with most support either included only one state variable or showed high spatial correlation between multiple populations. This is especially the case when using shorter time-series. We also show that adding noise to the time-series increasingly favour one-state models, suggesting that modelling on a finer spatial scale is more appealing with better quality data. Identification of more detailed patterns, such as ten-state-models with spatial synchrony decreasing by distance, was restricted to the taxa with the longest times series. Naturally, the appropriate scale of investigation may depend on the study question, so approximately equal model parsimony at different spatial scales can be seen as a delighting result. Typically, the models with multiple population states showed uniform patterns of temporal autocorrelation, suggesting that density dependence is most parsimoniously modelled as being uniform across space.
References
Andersen, J. H., J. Carstensen, D. J. Conley, K. Dromph, V. Fleming-Lehtinen, B. G. Gustafsson, A. B. Josefson, A. Norkko, A. Villnäs & C. Murray, 2017. Long-term temporal and spatial trends in eutrophication status of the Baltic Sea. Biological Reviews Cambridge Philosophical Society 92: 135–149.
Auger-Méthé, M., C. Field, C. M. Albertsen, A. E. Derocher, M. A. Lewis, I. D. Jonsen, J. Mills & J. Flemming, 2016. State-space models’ dirty little secrets: even simple linear Gaussian models can have estimation problems. Scientific Reports 6: 26677.
Bauerfeind, E., M. Elbrächter, R. Steiner & J. Throndsen, 1986. Application of laser doppler spectroscopy (LDS) in determining swimming velocities of motile phytoplankton. Marine Biology 93: 323–327.
Burnham, K. P. & D. R. Anderson, 2002. Model selection and multimodel inference: a practical information-theoretic approach, 2nd ed. Springer, New York.
de Wit, R. & T. Bouvier, 2006. ‘Everything is everywhere, but, the environment selects’; what did Baas Becking and Beijerinck really say? Environmental Microbiology 8: 755–758.
Defriez, E. J. & D. C. Reuman, 2017. A global geography of synchrony for marine phytoplankton. Global Ecology and Biogeography 26: 867–877.
Dennis, B., J. M. Ponciano & M. L. Taper, 2010. Replicated sampling increases efficiency in monitoring biological populations. Ecology 91: 610–620.
Durbin, J. & S. J. Koopman, 2012. Time series analysis by state space methods, 2nd ed. Oxford University Press, New York.
Engen, S. & B.-E. Sæther, 2005. Generalizations of the Moran effect explaining spatial synchrony in population fluctuations. The American Naturalist 166: 603–612.
Eppley, R. W., F. M. H. Reid, J. J. Cullen, C. D. Winant & E. Stewart, 1984. Subsurface patch of a dinoflagellate (Ceratium tripos) off Southern California: Patch length, growth rate, associated vertically migrating species. Marine Biology 80: 207–214.
Hällfors, G., 2004. Checklist of Baltic Sea Phytoplankton species (including some heterotrophic protist groups). Baltic Sea Environment Proceedings 95. Helsinki Commission, Baltic Marine Environment Protection Commission
Hampton, S. E., E. E. Holmes, L. P. Scheef, M. D. Scheuerell, S. L. Katz, D. E. Pendleton & E. J. Ward, 2013. Quantifying effects of abiotic and biotic drivers on community dynamics with multivariate autoregressive (MAR) models. Ecology 94: 2663–2669.
HELCOM, 2017. Monitoring of phytoplankton species composition, abundance and biomass. In: Manual for Marine Monitoring in the HELCOM COMBINE Programme of HELCOM (Last updated: July 2017. Internet page visited on 21.3.2018): http://www.helcom.fi/Documents/Action%20areas/Monitoring%20and%20assessment/Manuals%20and%20Guidelines/Guidelines%20for%20monitoring%20phytoplankton%20species%20composition,%20abundance%20and%20biomass.pdf
Hijmans, R. J., 2017. geosphere: Spherical Trigonometry. R package version 1.5-7.
Hinrichsen, R. A. & E. E. Holmes, 2009. Using multivariate state-space models to study spatial structure and dynamics. In Cantrell, C., C. Cosner & S. Ruan (eds.), Spatial Ecology. CRC/Chapman Hall, Boca Raton, Florida: 145–166.
Holmes, E. E., E. J. Ward & K. Wills, 2012. MARSS: multivariate autoregressive state-space models for analyzing time-series data. R Journal 4: 11–19.
Holmes, E. E., E. J. Ward, & M. D. Scheuerell, 2018. Analysis of multivariate time-series using the MARSS package. Version 3.10.8 https://cran.r-project.org/web/packages/MARSS/vignettes/UserGuide.pdf
Humbert, J.-Y., L. S. Mills, J. S. Horne & B. Dennis, 2009. A better way to estimate population trends. Oikos 118: 1940–1946.
Jaanus, A., A. Andersson, I. Olenina, K. Toming & K. Kaljurand, 2011. Changes in phytoplankton communities along a north–south gradient in the Baltic Sea between 1990 and 2008. Boreal Environment Research 16: 191–208.
Johannesson, K. & C. André, 2006. Life on the margin: genetic isolation and diversity loss in a peripheral marine ecosystem, the Baltic Sea. Molecular Ecology 15: 2013–2029.
Knape, J., 2008. Estimability of density dependence in models of time-series data. Ecology 89: 2994–3000.
Kuosa, H., V. Fleming-Lehtinen, S. Lehtinen, M. Lehtiniemi, H. Nygård, M. Raateoja, J. Raitaniemi, J. Tuimala, L. Uusitalo & S. Suikkanen, 2017. A retrospective view of the development of the Gulf of Bothnia ecosystem. Journal of Marine Systems 167: 78–92.
Lehtinen, S., S. Suikkanen, H. Hällfors, P. Kauppila, M. Lehtiniemi, J. Tuimala, L. Uusitalo & H. Kuosa, 2016. Approach for supporting food web assessments with multi-decadal phytoplankton community analyses –case Baltic Sea. Frontiers in Marine Science 3: 220.
Leppäranta, M. & K. Myrberg, 2009. Physical oceanography of the Baltic Sea. Springer, Berlin.
Lindén, A. & J. Knape, 2009. Estimating environmental effects on population dynamics: consequences of observation error. Oikos 118: 675–680.
Martiny, J. B., B. J. M. Bohannan, J. H. Brown, R. K. Colwell, J. A. Fuhrman, J. L. Green, M. C. Horner-Devine, M. Kane, J. Adams Krumins, C. R. Kuske, P. J. Morin, S. Naeem, L. Øvreås, A.-L. Reysenbach, V. H. Smith & J. T. Staley, 2006. Microbial biogeography: putting microorganisms on the map. Nature Reviews Microbiology 4: 102–112.
Moran, P. A. P., 1953. The statistical analysis of the Canadian lynx cycle. II. Synchronization and meteorology. Australian Journal of Zoology 1: 291–298.
Nash, J. & R. Varadhan, 2013. A Replacement and Extension of the optim() Function. R Package Version 2013(8): 7.
Olenina, I., S. Hajdu, L. Edler, A. Andersson, N. Wasmund, S. Busch, J. Göbel, S. Gromisz, S. Huseby, M. Huttunen, A. Jaanus, P. Kokkonen, I. Ledaine & E. Niemkiewicz, 2006. Biovolumes and size-classes of phytoplankton in the Baltic Sea. Baltic Sea Environment Proceedings 106. Helsinki Commission, Baltic Marine Environment Protection Commission.
Olli, K., O. Trikk, R. Klais, R. Ptacnik, R. Andersen, S. Lehtinen & T. Tamminen, 2013. Harmonizing large data sets reveals novel patterns in the Baltic Sea phytoplankton community structure. Marine Ecology Progress Series 473: 53–66.
R Core Team, 2017. R: A Language and Environment for Statistical Computing. R Foundation for Statistical Computing, Vienna Austria. https://www.R-project.org
Ranta, E., V. Kaitala, J. Lindström & E. Helle, 1997. The Moran effect and synchrony in population dynamics. Oikos 78: 136–142.
Ranta, E., V. Kaitala & J. Lindström, 1999. Spatially autocorrelated disturbances and patterns in population synchrony. Proceedings of the Royal Society B Biological Sciences 266: 1851–1856.
Scheef, L. P., S. E. Hampton & R. Izmest’eva, 2012. Inferring plankton community structure from marine and freshwater long-term data using multivariate autoregressive models. Limnology and Oceanography: Methods 11: 475–484.
Snoeijs-Leijonmalm, P. & E. Andrén, 2017. Why is the Baltic Sea so special to live in? In Snoeijs-Leijonmalm, P., H. Schubert & T. Radziejewska (eds.), Biological Oceanography of the Baltic Sea. Springer, Dordrecht: 23–80.
Sjöqvist, C., A. Godhe, P. R. Jonsson, L. Sundqvist & A. Kremp, 2015. Local adaptation and oceanographic connectivity patterns explain genetic differentiation of a marine diatom across the North Sea-Baltic Sea salinity gradient. Molecular Ecology 24: 2871–2885.
Suikkanen, S., M. Laamanen & M. Huttunen, 2007. Long-term changes in summer phytoplankton communities of the open northern Baltic Sea. Estuarine, Coastal and Shelf Science 71: 580–592.
Suikkanen, S., S. Pulina, J. Enström-Öst, M. Lehtiniemi, S. Lehtinen & A. Brutemark, 2013. Climate change and eutrophication induced shifts in northern summer plankton communities. PLoS ONE 8: e66475.
Sæther, B.-E., S. Engen, V. Grøtan, W. Fiedler, E. Matthysen, M. E. Visser, J. Wright, W. P. Møller, F. Adriaensen, H. Van Balen, D. Balmer, M. C. Mainwaring, R. H. McCleery, M. Pampus & W. Winkel, 2007. The extended Moran effect and large-scale synchronous fluctuations in the size of great tit and blue tit populations. Journal of Animal Ecology 76: 315–325.
Tolimieri, N., E. E. Holmes, G. D. Williams, R. Pacunski & D. Lowry, 2017. Population assessment using multivariate time-series analysis: a case study of rockfishes in Puget Sound. Ecology and Evolution 7: 2846–2860.
Turchin, P., 1995. Population regulation: old arguments and a new synthesis. In Capuccino, N. & P. W. Price (eds.), Population Dynamics: New Approaches and Synthesis, 1st ed. Academic Press, Cambridge: 19–39.
Turchin, P., 2003. Complex Population Dynamics: a Theoretical/Empirical Synthesis., Vol. 35. Princeton University Press, Princeton.
Ward, E. J., H. Chirakkal, M. González-Suárez, D. Aurioles-Gamboa, E. E. Holmes & L. Gerber, 2010. Inferring spatial structure from time-series data: using multivariate state-space models to detect metapopulation structure of California sea lions in the Gulf of California, Mexico. Journal of Applied Ecology 47: 47–56.
Wasmund, N., J. Tuimala, S. Suikkanen, L. Vandepitte & A. Kraberg, 2011. Long-term trends in phytoplankton composition in the western and central Baltic Sea. Journal of Marine Systems 87: 145–159.
Wickham, H., 2016a. Tools for splitting, applying and combining data. R package version 1.8.4.
Wickham, H., 2016b. Flexibly reshape data: a reboot of the reshape package. R package version 1.4.2.
Wickham, H & W. Chang, 2016. Create elegant data visualisations using the grammar of graphics. R package version 2.2.1
Wennerström, L., L. Laikre, N. Ryman, F. M. Utter, N. I. Ab Ghani, C. André, J. DeFaveri, D. Johansson, L. Kautsky, J. Merilä, N. Mikhailova, R. Pereyra, A. Sandström, A. G. F. Teacher, R. Wenne, A. Vasemägi, M. Zbawicka, K. Johannesson & C. R. Primmer, 2013. Genetic biodiversity in the Baltic Sea: species-specific patterns challenge management. Biodiversity and Conservation 22: 3045–3065.
Zingone, A., P. J. Harrison, A. Kraberg, S. Lehtinen, A. McQuatters-Gollop, T. O’Brien, J. Sun & H. H. Jakobsen, 2015. Increasing the quality, comparability and accessibility of phytoplankton species composition time-series data. Estuarine, Coastal and Shelf Science 162: 151–160.
Acknowledgements
Open access funding provided by Åbo Akademi University (ABO). We thank Jonna Engström-Öst and the anonymous reviewer for comments that improved the manuscript. Louise Lindroos was funded by Onni Talaan Säätiö.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Handling editor: Judit Padisák
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Forsblom, L., Lehtinen, S. & Lindén, A. Spatio-temporal population dynamics of six phytoplankton taxa. Hydrobiologia 828, 301–314 (2019). https://doi.org/10.1007/s10750-018-3826-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10750-018-3826-2