Abstract
The brown trout (Salmo trutta L.) is widely distributed all around Europe but its natural diversity is threatened by massive stocking with Atlantic domestic strains. Describing the remaining natural genetic diversity and the proportion of domestic hatchery strains in rivers is a prerequisite for smart conservation. The high genetic diversity of brown trout populations around the Tyrrhenian Sea is well known. Use of twelve microsatellites has allowed description of the natural genetic structure of populations and detection of the consequences of stocking. Mitochondrial DNA control region sequences and the LDH-C1* gene enabled placement of each population into one of the six mitochondrial and two allozymic known evolutionary lineages. The Corsican populations showed low intra-population genetic diversity but an exceptionally high level of inter-population differentiation. More southern Tyrrhenian regions exhibited opposite pattern of diversity, partly due to the Atlantic domestic introgression. Globally, the natural structure outlines two north–south clines: high inter-population differentiation and predominance of the Adriatic lineage in the north, but lower inter-population differentiation and the presence of the natural Atlantic lineage in the south. In addition, the Tyrrhenian region is the contact zone between the widespread Adriatic lineage and a local natural Atlantic lineage probably coming from North Africa through the Strait of Gibraltar.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The brown trout is widespread all around Europe, western Asia, and North Africa at medium and high elevation (Behnke, 1972, 1986). It is one of the most extensively managed freshwater fish species worldwide because of its high economical value, mainly for sport fishing. As a consequence, brown trout is regularly stocked in most part of its distribution from multiple hatcheries that breed various strains in many countries. This species has also been introduced and acclimatized in numerous countries around the world where sometimes it is considered as an invader (De Moor & Bruton, 1988; Olsson et al., 2006; Miró & Ventura, 2013).
The brown trout is a complex taxon, first investigated through morphology, then with molecular markers. The term of “Salmo trutta complex” (Patarnello et al., 1994; Bernatchez & Osinov, 1995; Giuffra et al., 1996; Bernatchez, 2001) is used to indicate that the complex is composed of numerous differentiated geographic forms (without consensus on the taxonomy) that can easily hybridize with each other (Largiadèr & Scholl, 1996; Meldgaard et al., 2007). Its ecological diversity is also remarkable with sedentary populations in the cold upstream of rivers, and anadromous ones that migrate to seas or lakes. Based on phenotypes, many species have been described. Kottelat & Freyhof (2007) reviewed the published nomenclature and reached the number of 28 nominal species. This has been augmented by at least fifteen new species described during the last 12 years (Delling & Doadrio, 2005; Turan et al., 2009, 2010; Delling, 2010; Turan et al., 2011, 2012, 2014a, 2014b; Doadrio et al., 2015).
Phylogenetic studies based on mitochondrial DNA (mtDNA) provided a first clear and testable way of classification, dividing S. trutta into five main evolutionary lineages, considered as geographic variants but not as species. Thus, Atlantic (AT), Mediterranean (ME), Marble (MA), Adriatic (AD), and Danubian (DA) lineages have been described (Bernatchez et al., 1992) and confirmed in many publications. Other lineages, i.e., monophyletic regional variants at the base of large phylogenetic lineages, such as Duero (DU, Suárez et al., 2001), Tigris (TI, Bardakci et al., 2006), the Balkan cluster (Snoj et al., 2009), and Dades (Snoj et al., 2011), have also been proposed.
The Western Mediterranean basin is considered to extend from the Siculo–Tunisian to the Gibraltar Straits. Its eastern part consisted of more or less isolated zones constituting the Tyrrhenian and Ligurian seas. The Adriatic and Ionian seas belong to the Eastern Mediterranean basin and are at the contact with the Western basin. Several publications analyzed the genetic diversity of trout in limited parts of its distribution around the Tyrrhenian Sea using allozymes, RFLP, and sequences of a portion of the mitochondrial control region (mtDNA CR) and microsatellites (Nonnis Marzano et al., 2003; Lucentini et al., 2006; Gratton et al., 2014; Fabiani et al., 2018), but the genetic diversity of brown trout in the Tyrrhenian Sea region has not yet been examined in detail.
According to the large literature based on morphology and molecular genetics proposing taxonomic organization of the trout populations inhabiting the Tyrrhenian periphery, a rather confusing picture has emerged. Nera River (Roma region) was considered to be an indigenous pool, home of the Mediterranean S. trutta lineage (Lucentini et al., 2006). Corsican rivers were proposed to host S. macrostigma (Roule, 1933; Guyomard, 1989; Bernatchez et al., 1992) or S. trutta (Berrebi, 2015), while Sardinian rivers were inhabited by S. macrostigma (Boulenger, 1901; Mola, 1928; Pomini, 1940; Gandolfi et al., 1991; Patarnello et al., 1994; Massidda, 1995; Sabatini et al., 2006; Orrù et al. 2010) or S. trutta (Sabatini et al., 2011) or S. cettii (Zaccara et al., 2015). South Sicilian trout are of Atlantic S. trutta lineage (Schöffmann et al., 2007) or S. cettii (Kottelat, 1997; Kottelat & Freyhof, 2007; Schöffmann et al., 2007; Duchi, 2011, 2018; Bianco, 2014; Sabatini et al., 2018).
In the present study, the whole area around the Tyrrhenian Sea was sampled in order to describe the overall structure of trout populations. For this, microsatellite genotypes (12 loci), mtDNA CR sequences, and LDH-C1* genotypes were analyzed. In addition, the impact of stocking was measured. In light of this new knowledge, the principles of conservation are discussed.
Materials and methods
Sampling distribution
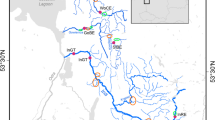
The present survey concentrates sampling to the freshwater basins distributed all around the Tyrrhenian Sea with some Ionian Sea tributaries (Fig. 1). In order to introduce comparative samples, a river entering the Ligurian Sea at the French–Italian boundary and three commercial Atlantic domestic samples from northern France and central Italy have been included (Table 1). The French hatchery is representative of the international commercial Atlantic strain and the Italian ones of local domestic strains, both derived from genitors from several countries (Bohling et al., 2016).
Geographic position of 29 samples (the last one, number 30, positioned far in the north, is not shown). The station numbers refer to the first column of Table 1
Electrofishing was conducted between 2004 and 2014. A total of 365 river trout (sub-adults and adults) were anaesthetized and a small fin clip taken (preserved in 96% ethanol) before each fish was returned to its river. In the three hatcheries sampled, the method was similar but the fish were caught with dip nets.
Some of the samples contained few fish (Table 1): lower Tiber in Lazio region (7–9 specimens), Calabria (3–7), and Sicily (6–11), except for the Anapo River. These samples were very difficult to constitute because of the low trout density. In Sicily and Calabria, the species is protected and special permits are needed with a given maximum number of trout to be caught. Fortunately, most often, this concerned nearby small rivers which could be concatenated for population parameter calculations.
DNA extraction
DNA was extracted using the Chelex/proteinase K-based method described by Walsh et al. (1991) and Estoup et al. (1996) and then improved by Yue & Orban (2005). A small piece of fin was incubated overnight at 56°C in 195 µl of 5% Chelex 100 Resin (Biorad) solution containing 50 mM of Tris-HCL (pH 7) and 500 µg/ml of proteinase K. Samples were then incubated at 95°C for 10 min before centrifugation at 3500 g for 5 min. Supernatants were recovered and frozen at − 20°C until required for use.
Mitochondrial marker
The mtDNA CR was amplified by PCR using the PST and FST primers (Cortey & García-Marín, 2002). Each 50 μL reaction included 0.4 μM of each primer (Eurofins MWG Operon), dNTP (2 mM each), 2 mM of MgCl2, 10 μL of 5 × PCR buffer, 1 U of Taq polymerase (GoTaq® Promega), and about 50 ng of genomic DNA. The PCR conditions included initial denaturation (95°C, 5 min), followed by 30 strand denaturation (94°C, 1 min), primer annealing (52°C, 1 min), and DNA extension (72°C, 1 min) cycles, and then by a final extension (72°C, 5 min). All PCR amplifications were performed in Eppendorf Mastercycler thermocyclers. The amplified DNA fragments were run on a 0.8% agarose gel to verify the amplification efficiency. The amplified products were purified and sequenced in both directions to confirm the polymorphic sites in an ABIPRISM 3130/xl/sequencer (Applied Biosystems).
The mtDNA CR sequences were aligned using the computer program Clustal X (Thompson et al., 1997) implemented in MEGA version 6 (Tamura et al., 2013). In order to assign Tyrrhenian trout to a lineage, data were aligned and compared with reference S. trutta CR sequences from GenBank (AT, DU, ME, MA, AD and DA) (see Table S1 for haplotype details). The genealogical relation of haplotypes was depicted using a 95% statistical parsimony network constructed using TCS 1.3 (Clement et al., 2000).
LDH-C1* genotypes
Some of the sampled specimens were genotyped at the LDH-C1* gene coding the LDH enzyme (Table 1). This marker, largely used in the past through allozyme electrophoreses (Hamilton et al., 1989; Almodóvar et al., 2006; Berrebi, 2015), was analyzed by the PCR–RFLP method of McMeel et al. (2001). This popular marker separates the ancestral allele *100 found in Atlantic salmon and southern brown trout lineages and Atlantic domestic strains carrying the derived *90 allele (Hamilton et al., 1989; García-Marín et al., 1999; Berrebi, 2015). Native Mediterranean trout populations belong to the mitochondrial trout lineages ME, AD, MA, and AT. This last lineage is represented by the so-called southern Atlantic clade, sensu Cortey et al. (2004) and also exhibits the LDH-C1*100 allele (Aurelle & Berrebi, 2002). Therefore, this marker is used, here, mainly to distinguish between the AT haplotypes belonging to the southern Atlantic clade and the AT haplotypes of northern origin (marked with LDH-C1*90) that invaded the Mediterranean region subsequent to stocking activities. According to Hamilton et al. (1989), all Mediterranean trout populations are characterized by the allele *100, except the marble trout (allele LDH-C1*120). On the contrary, most north Atlantic populations carry the allele *90 with several exceptions (García-Marín et al., 1999). This marker can be used to distinguish natural trout, including natural Atlantic populations in Sicily carrying the *100 allele (Schöffmann et al., 2007) from introduced hatchery Atlantic trout.
Microsatellite loci
The primers of the twelve microsatellite markers used in this study were obtained from the literature. Repeated sequences are all dinucleotide except Ssa197 which is a tetranucleotide microsatellite (O’Reilly et al., 1996). Details about each locus and multiplexes are given in Table 2.
For each marker, one of the 5′ ends of the two primers was end-labeled with a fluorescent dye, either 6-FAM, HEX, or NED. Polymerase chain reactions (PCR) were performed using the Qiagen multiplex PCR kit in a final volume of 10 µl, containing 3 µl of genomic DNA diluted at 10 ng/µl, 5 µl of Qiagen PCR Master Mix, 1 µl of Qiagen Q-solution, and 1 µl of primer mix at 2 µM each (Eurofins MWG Operon). Amplifications were carried out in a GeneAmp PCR System 2700 thermal cycler (Applied Biosystems), according to the supplier’s instructions (Qiagen multiplex PCR kit): initial denaturation step (95°C, 15 min), followed by 35 cycles of denaturation (94°C, 30 s), annealing (55°C for the three multiplexes, 90 s), and extension (72°C, 60 s), with a final extension step (60°C, 30 min). Amplified PCR fragments were then diluted and separated on an ABIPRISM 3130/xl/sequencer (Applied Biosystems) with GeneScan 500 Rox dye size standards. Allele sizes were determined using the GeneMapper v4.1 software system (Applied Biosystems, Life Technologies).
A genotype matrix was then constructed and used as a basis for all the following statistical analyses mainly carried out using GENETIX 4.05 (Belkhir et al., 2004).
Microsatellites statistics
For estimation of genetic polymorphism, expected heterozygosity (He), unbiased expected heterozygosity (Hnb: Nei, 1978), and observed heterozygosity (Ho) were calculated for each sample or river system. The mean number of alleles by locus (A) is another way to estimate the diversity in a population. These calculations were limited to samples of ten or more specimens. For this reason, nearby locations characterized by very low fish number were grouped in order to increase the sample size.
Inter-sample or inter-river system differentiations (Fst) and the intra-river panmixia (Fis) were estimated (θ and f estimators of Weir & Cockerham, 1984, respectively). The significance of the Fst and Fis values was tested by random permutation procedures: 5000 individual permutations between samples for Fst and 5000 allele permutations within samples for Fis, processed using GENETIX.
The sequential Bonferroni correction was applied for multiple tests (Rice, 1989). Micro-Checker software (van Oosterhout et al., 2004) was run in order to detect null alleles, drop-out, or stuttering perturbations.
A general picture of the trout genetic diversity was first obtained through multidimensional analyses. Here Factorial Correspondence Analyses (FCA: Benzécri, 1973), allowing the overall structure of the sampling to be explored, were carried out as implemented in GENETIX. Three focuses were chosen: all the samples, the Sicilian two lineages, and the Atlantic diversity of the sampling (Fig. 3a–c). The clusters (or clouds) observed in the diagrams correspond to nuclear genetic homogeneous lineages. The mathematical method is clearly detailed in She et al. (1987).
In order to detect differentiated subgroups, hierarchical STRUCTURE assignment analyses were performed for the whole sample set (Vähä et al., 2007; Marić et al., 2017). The program STRUCTURE 2.3.2 (Pritchard et al., 2000) runs Markov chain Monte Carlo simulations to partition individuals into K clusters (here, K was run between 1 and 24). Basic assignment criteria are the minimization of Hardy–Weinberg and of linkage disequilibria (Pritchard et al., 2000). Different run lengths were used at each step (from 20,000 to 100,000 burn-in and 100,000 to 500,000 total lengths, repeated seven times for each K) depending on convergence (proportion of identical structure among the seven repeated runs at each step). The admixture ancestry model and correlated allele frequency options were chosen. Sampling location was not used as prior information.
The ΔK method (Evanno et al., 2005) was applied to estimate the most probable K at each step (see Table S2). The first step determines the first hierarchical level, the whole sample set being split into K subgroups. Then each subgroup was analyzed separately, allowing for more precise clustering of individuals without eliminating admixed individuals. This hierarchical method was applied until no further substructure was observed. A comparison with the simple assignment test with the best K (here K = 19) is provided in Figs. S5 and S6.
Finally, the parentage software COLONY 2.0.6.4. (Jones & Wang, 2010) was applied especially on Corsican and Sardinian samples. We have chosen the Pair-Likelihood-Score (PLS)/Full-Likelihood (FL) combined (= FPLS) algorithm in order to establish only full-sibs listing. The objective is to understand the family structure of these very low polymorphic and very differentiated/isolated populations.
Results
Mitochondrial sequences
A total of 144 mtDNA CR sequences were obtained in this survey corresponding to sixteen different haplotypes, seven of which are undescribed so far (AD-Tyrrh1 to 6 and AT-Tyrrh1: Tables 3 and S1 for distribution and accession numbers). The alignment length was 987 bp providing 34 parsimony informative sites. Haplotype classification allows the relationships between Tyrrhenian populations and the five main known lineages widespread around Europe plus the DU junior lineage to be understood. The network (Fig. 2) represents the haplotype organization within the Tyrrhenian region, with GenBank published haplotypes used as references. Tyrrhenian haplotypes all belong here to AD or AT lineages exclusively.
Network presenting the relationships between Tyrrhenian control region sequences (colored circles) and published ones (white circles). This network allows classification of the Tyrrhenian haplotypes within the five main known lineages (plus DU). The circle sizes are proportional to the haplotype occurrence except the white ones
The new haplotype AD-Tyrrh1 is the most common, widespread in all regions except Sicily, and in a central position in the AD haplogroup (Fig. 2). AD-Tyrrh5 and 6 are exclusive to Nera River (i.e., upper Tiber). With nine different haplotypes detected, the AD haplogroup was the main non-domestic mtDNA haplogroup observed in the Tyrrhenian region (56%, Table 3). Note that the sequence of the AD-Tyrrh4 haplotype, newly discovered in the present study, contains an 82 bp repeat towards the 3′-end of the CR. As the elongation model of this repeat is generally thought to be the result of intra-molecular processes (Buroker et al., 1990), only the first copy was kept in the analysis. The native AD lineage was present in 96% of the Corsican Tyrrhenian trout, 100% of the upper Tiber River (Nera), 67% of the lower Tiber River (Simbrivio and Aniene), 50% of the Sardinian trout, 46% of Calabrian trout, and 0% of Sicilian ones, representing a north–south cline partly linked to stocking activities. In the present survey, six AD haplotypes are new, suggesting their endemism (Table S1).
Corsica was mainly characterized by the new haplotype AD-Tyrrh1, except for the Acqua d’Acelli population that had only the ADcs15 haplotype. AD-Tyrrh1 accounted for 88% of the Corsican AD haplotypes, 31% of the Sardinian ones, 8% in the Tiber basin in central Italy, 50% of the Calabrian diversity (but with a sample size of 6) and was not present in Sicily. Here again, a cline affected the distribution of this variant, exhibiting a decrease from north to south. The other AD haplotypes were distributed without clear structure in Sardinia, the Tiber River, and Calabria (Table 3).
Among the AT lineage, the AT-s6 haplotype should represent a natural migration of Atlantic trout into the Mediterranean (Schöffmann et al., 2007). This haplotype of 380 bp is included in the 985-bp haplotype ATSic detected here. This haplotype was the only one present in the Anapo River (Sicily). “Clone JE1,″ the other Sicilian AT haplotype, was the only one found in the Manghisi River, a tributary of the River Cassibile. On the contrary, haplotypes 1–4 (Table 3) are well known to characterize north Atlantic populations and the domestic commercial Atlantic hatchery strain (Cortey & García-Marín, 2002; Cortey et al., 2004). These haplotypes, synonymous with ATcs1 to 4 (Cortey et al., 2004), are well represented in both Italian and French hatcheries, and in rivers where they had been introduced by stocking. They were observed in Corsica (globally 4%), Sardinia (50%), Calabria (54%), and in the lower Tiber basin (33%). The new and rare haplotype AT-Tyrrh1 was found only in the Cantiano hatchery (sample 15) but not in any river. In Sicily, only the Atlantic haplotype was observed. Their natural/domestic origin is discussed below.
LDH-C1* genotypes
Genotyping this gene is especially useful to distinguish between natural and domestic origins of the AT haplotypes of Sicily. Some of the samples were tested, especially those with AT mtDNA haplotypes (Table 1). Hatchery samples were nearly all characterized by the LDH-C1*90 allele except for one *90/100 heterozygote genotype among the 20 Cantiano hatchery trouts (sample 15). This hatchery *90 allele was also found in 8/8 in the Allaro River and 7/10 in the Diga Giulia River, both in Calabria, and just 1/10 in the Anapo River in Sicily. The domestic allele LDH-C1*90 was not found in Corsica (but only four fishes analyzed) nor in the Manghisi River in Sicily (five trouts).
Microsatellite genotypes
For samples numbering over ten specimens (20 of the 30 considered), the Micro-Checker software made no drop-out detection, two tests among 240 had suspect stuttering, and 17 among 240 suggested the occurrence of null alleles. The 17 possible null allele detections were spread among 8 of the 12 loci, with 0–3 cases each, showing that there was no systematic presence of null alleles in a given marker, probably not disturbing the calculations.
The first representation of overall microsatellite genetic diversity is given by multidimensional analyses. Figure 3a first isolates Corsican populations at the negative part of the first axis, with very low intra-population and very high inter-population diversity, together with the absence of Atlantic influence. The southern samples of Sardinian trout (numbers 12–14) clustered in one group. Note that because this is a 2D representation of a 3D simplification of the 273D hyperspace (274 × 324 matrix), Sardinian autochthonous trout (green envelope) are clearly separated from E Maghjine Corsican trout (red circles) along axis 3 despite the apparent overlapping in the diagram (see Fig. S4). The same is true between Lataga (gray circles) and Acqua d’Acelli (dark green circles) fishes (Fig. S4).
Sampling diversity analyzed through Factorial Correspondences Analysis. Circles correspond to France and Corsica, diamonds to Sardinia, triangles to continental Italy, squares to Sicily, and crosses to domestic samples. The different colors distinguish different populations of the same region. a General picture gathering all the samples. C. Italy = Central Italy (the Marche and Lazio); ellipses: red = Corsica, green = part of Sardinia, blue = part of C. Italy, black = domestic Atlantic, dotted orange = overlapping samples developed in Fig. 3b. b In order to focus on the samples included in the doted orange envelope of Fig. 3a, a new FCA was performed with these samples only (except the Roya River ME sample), giving new insight into the Anapo River differentiation. c Focus on the samples included in the dotted orange envelope of Fig. 3a, after withdrawal of the Roya River ME and the Anapo River AT samples. Black ellipse = true domestic Atlantic trout (samples 10, 11, 15, 16, 18, 23, and 30). Red ellipse = Calabrian sample containing both wild and domestic trout (samples 20–26). Lower Tiber = populations close to Rome. The upper Tiber gathers the two domestic strains from the hatcheries Cantiano and Visso (samples 15 and 16). South Sicily samples are Manghisi and San Marco (28 and 29)
In contrast, several trout populations seemed to be introgressed or hybridized with the commercial international Atlantic strain represented here by the Italian and French hatcheries (Cantiano, Visso and Isère hatcheries, samples 15, 16 and 30). They are positioned at the very end of the positive part of axis one. In order to understand this part of the diagram, Atlantic-like samples were reanalyzed alone. The second analysis (Fig. 3b) showed clear separation of the Anapo population (Sicily, sample 27) from the remaining populations. Finally, the third analysis (Fig. 3c) that considered the Atlantic-like samples without the Anapo one, gathered at the left the domestic Atlantic strains and the Ermolinus and Sadali samples of central-eastern Sardinia (numbers 10 and 11 in Table 1) and those from Allaro and Precariti in Calabria (23 and 26), probably of domestic origin. The remaining samples collected around two clusters: first, numbers 18, 19, 28, and 29 (respectively from the Tiber River in Lazio province and the southern samples in Sicily), and second, numbers 20–25 corresponding to most samples from Calabria (except for 23, the Allaro River, probably of full domestic origin).
Summarizing, the multidimensional analyses highlighted several types of populations:
-
insular/endemic differentiated populations: all Corsican (samples 2–9) and southern Sardinian samples (12–14); the Nera population in continental Italy (17) (Fig. 3a);
-
the Anapo River population (27) clearly isolated from the remaining Atlantic populations (Fig. 3b);
-
southern populations, close to the domestic cluster: downstream Tiber River samples (numbers 18 and 19); two Sicilian samples (28 and 29); most Calabrian samples (numbers 20–24, except for sample 23 from the Allaro River that clustered with domestic trout);
-
domestic Atlantic samples: Italian and French hatchery strains (numbers 15, 16 and 30); central-eastern Sardinian samples (10 and 11); the Simbrivio River, a Tiber River tributary (18); the Allaro River in Calabria (23) (Fig. 3c).
The hierarchical STRUCTURE analysis gave very similar results (Fig. 4, Table S2). In the first hierarchical step, trouts were separated into two groups. Group A mostly consisted of natural samples from Corsica and Sardinia, while group B was mainly composed of samples containing the AT lineage. In group B (subgroup B2) some are domestic hatchery samples of AT lineage (Cantiano, Visso and Isère). Others are an assemblage of domestic AT individuals in different proportions with native ME (Roya) and AD lineage (U Furcone, Ermolinus, Sadali, lower Calabria, and lower Tiber), which is evident from the second to the fifth step, depending on the sample (subgroups B2 and B3). Finally, several are native samples of AT (Anapo and south Sicily) and AD (Ancinale and Nera) lineages (subgroup B1). However, in southern Sicilian rivers, there is a small proportion of domestic AT genotypes visible in the second step of the hierarchical assignment. This heterogeneity (especially in group B) explains that the hierarchical analysis reached up to 22 clusters. Some clusters showed substructures (lower Tiber, lower Calabria, Isère hatchery, Cantiano hatchery, Ermolinus, U Furcone, and Marroccu) which are marks of migrations or introductions. The Nera sample is probably not sub-structured since the program cuts each individual into two lineages but do not partition them.
Population structure as inferred by hierarchical STRUCTURE analysis of microsatellite data. White lines separate the sampling sites. Cluster names correspond to river names or to regions. In order to link each cluster to the elementary sample(s) involved, the numbers from the first column in Table 1 are indicated at the left of the figure—first histogram. The most probable K for the analyzed samples shown in the arrows is based on the ΔK method; no further structures were detected in subsequent rounds (after the sixth step) and within the excluded clusters (K = 1). Arrows delineate the progress of the hierarchical approach, where subsets of the data were subsequently analyzed. Corresponding mtDNA (control region haplotypes) and LDH-C1* alleles are indicated at the right of the figure
Unlike the hierarchical Structure analysis and ΔK method (Evanno et al., 2005) which estimated 22 groups, method based on maximizing the mean estimated ln probability of data (ln P(D); Pritchard et al., 2000) show that the most probable K = 19 (Figs. S5 and S6). This last analysis method is less informative than hierarchical Structure, and, unlike hierarchical method, some of populations are not detected as distinct (Pozzi, Val d’Esse and Marroccu), which is very important for delimitation of MU.
Parentage analyses performed with COLONY software detected one to three families in each autochthonous Corsican or Sardinian population. However, in north-east Sardinian populations, the Ermolinus and Sadali populations are composed of 7–9 families of generally 1 or 2 individuals (except two families of 3 and 5 trouts, see Table S3).
Comparing different marker systems
Comparing data provided by the three categories of markers allowed the general genetic structure of trout around the Tyrrhenian Sea to be described. The LDH-C1* marker is monomorphic for the considered natural populations and was used as a diagnostic marker between natural and domestic trout.
The Roya trout population belongs to the ME lineage, according to mtDNA CR sequences (unpublished data) constituting the ME lineage reference for microsatellite diversity and assignment (Table 4, Fig. 4). In the sequence data constituting the present study, this ME lineage was not observed all around the Tyrrhenian Sea.
Corsican and Sardinian natural populations were mostly characterized by the AD-Tyrrh1 haplotype (with a few AD-Tyrrh2 and AD-Tyrrh4). Among this mitochondrial homogeneity, there were two exceptions: the Corsican population of Acqua d’Acelli had only the ADsc15 haplotype and the Sardinian population Is Abius was characterized by the A_2 Adriatic haplotype. The microsatellite markers divided Corsican and Sardinian natural populations into nine clusters in Fig. 3a and at the third step of the hierarchical assignment analysis (Fig. 4).
Along the Tyrrhenian-Ionian Italian Peninsula, the upper Tiber River trout (Nera River, sample 17) appear as a separate cluster in the third step of the hierarchical analysis, which is congruent with the mtDNA haplotype composition. Here, two private haplotypes were observed (AD-Tyrr5 and 6) that probably have a trans-Apennine origin due to their genetic similarity with haplotype ADcs11 (see Fig. 2), this latter being very common in the Adriatic region (Sušnik et al., 2007; Berrebi et al., 2013).
On the contrary, the lower Tiber River trout (samples 18 and 19) showed wide and distinct haplotype diversity of more western origin (AD-Tyrrh1, 2 and 4).
Further south, the Calabrian samples were separated into two lineages at the second step of the hierarchical analysis (B1 and B2): the Ancinale River system (samples 20–22) and lower Calabria (samples 23–26). The Ancinale cluster was characterized by the common haplotype AD-Tyrrh1 and the lower Calabrian by two private haplotypes: AD-Tyrrh3 and ADcs1.
Sicilian trout populations exhibited two AT haplotypes (ATSic and “clone JE1”). The clone JE1 haplotype, fixed in the Manghisi River population is of native origin on the basis of the associated fixation of the LDH-C1*(100) allele (Table 3). Microsatellite Sicilian clusters separated lately (at the third step) the Manghisi and San Marco populations (south Sicily—samples 28 and 29) from the Anapo population (27) in correlation with the two haplotypes (Table 3). With only one exception, all analyzed Sicilian trouts showed the LDH-C1*100 wild genotype.
All around the Tyrrhenian Sea, AT haplotypes AT-Tyrrh1 and “haplotypes 1–4” marked the domestic lineage assigned to the B2 cluster (Fig. 4) based on microsatellites and recognizable with the LDH-C1*90 allele, while clusters B1 and B3 dominantly represented composite natural lineages.
Population parameters
Population parameters add some biological information. Heterozygosity, as measured with various H parameters and with the mean number of alleles A (Table 4), was very low in Corsica (Hnb = 0.17), but very high in Sardinia (0.58). Parameter A, the mean number of alleles by locus, was also very distinctive (respectively 1.9 and 4.7). Hatchery populations are considered to be highly polymorphic (Berrebi et al., 2000; Bohling et al., 2016) and displayed large values: 0.67 < Hnb < 0.78 and 5.7 < A < 8.3.
Inter-sample differentiation can be determined using the estimations of Fst (Table 5). Most sample pairs were highly significantly differentiated except for a few cases which became moderately significant after Bonferroni correction. Considering only feral trout, in Corsica the mean inter-sample Fst was 0.74, which is a very high value among neighboring rivers (0.47 > Fst > 0.90). The mean Fst was lower in Sicily (0.34) and among the Tiber tributaries (0.29). In Sardinia, the Ermolinus and Sadali trout populations were deeply introgressed by domestic forms. Only Marroccu, Is Abius and Camboni populations are native, with very low differentiation (0.03 < Fst < 0.08). In Calabria, the small sample sizes limited the estimations between groupings to an opposition between the Ancinale and Alaro/Assi/Stilaro/Precariti basins (Fst was 0.17).
Discussion
Combining nuclear and mitochondrial markers provided a rather clear structure of the Tyrrhenian (and Ionian) trout populations. Very high inter-population diversity was observed in Corsica. A double north–south cline was detected: (i) high inter-population differentiation in the north (mainly Corsica) and lower in the south (ii) predominance of the Adriatic lineage in the north, which decreased towards the south due to stocking in Sardinia, the Tiber basin, and Calabria and because of the natural Atlantic lineage settlement in Sicily.
Origin of Tyrrhenian trout diversity
Corsican trout can be considered as the best conserved natural stock around the Tyrrhenian Sea, since all samples (except number 2) were in the A cluster, which characterizes mostly AD individuals (Fig. 4). As a possible explanation, the riverine ecosystems have been relatively well preserved in the island and the upstream part of the watersheds is generally free of pollution. Hydrogeological characteristics are also favorable to salmonids. According to Gauthier & Berrebi (2007), the ancestral lineage (AD) has been isolated upstream by impassable waterfalls since the last glacial maximum. Below these waterfalls, postglacial Mediterranean invader trout (ME, not sampled here) hybridized with resident AD populations. The numerous upstream isolated ancestral populations are free of ME invader or of domestic introduction (Fig. 4), confirming the Berrebi (2015) survey of 38 samples, some of which were re-sampled for this study (stations 2, 6 and 7). Traces of stocking (AT) are rare there (imperceptible with nuclear markers and 4% according to the mitochondrial sequences). This favorable situation explains the preserved high differentiation between populations (0.47 > Fst > 0.90; mean value 0.74, the highest value among all the samples). Moreover, the small size of streams together with possible bottlenecks could explain the very low intra-population diversity (0.08 < Hnb < 0.26, the lowest values among all the samples) compared to native grouped Tiber populations (0.51 < Hnb < 0.75), native Sardinian trout (0.45 < Hnb < 0.51), and even the grouped Calabrian (0.62 < Hnb < 0.78) and Sicilian populations (0.49 < Hnb < 0.59). Table 3 shows that these populations exhibit only one haplotype each (except for sample 9 from the Lataga River), mostly the newly described AD-Tyrrh1.
In order to better describe these island isolated small populations, COLONY software (Jones & Wang 2010) was used to research family structure in Corsican populations (not shown). In Corsica, each population shows only one to three families (Table S3). In fact, the polymorphism is so low (0.08 < Hnb < 0.26) that we can deduce that these small populations, totally isolated, have suffered several drastic recent bottlenecks. We can also suppose that one or two pairs of parents have recently re-funded each population making naturally each sample 1–3 families. The other interpretation is that such low polymorphism does not allow families detection. In Sardinia, the landscape is totally different: (i) in north-east populations (Flumendosa basin), mostly composed of Atlantic domestic lineage, the Ermolinus and Sadali populations are composed of 7–9 families of generally 1 or 2 individuals. These highly polymorphic populations (here 0.74 < Hnb < 0.79) are composed of various origins. (ii) In the south, the same local lineage has been observed in three samples of the same watershed (Cixerri basin) with possible exchanges. The medium level of polymorphism (0.41 < Hnb < 0.51) corresponds to rather large populations and possibly exchanges between them. The whole Cixerri sampling forms 12 families of full-sibs frequently composed of individuals of the three sampled locations (Table S3).
If we compare these values with populations analyzed in the literature with similar markers (9–11 microsatellite loci), the first difficulty is the diversity of cases mainly in terms of population size. However, with the filter of case similarity, we observed very high diversity in northern populations (generally of AT mtDNA lineage) as in Switzerland (Stelkens et al., 2012: 0.73 < He < 0.81) or in Romania (Popa et al., 2016: 0.79 < He < 0.82), but lower values in Catalonia/Spain, as around the Tyrrhenian Sea (Araguas et al., 2017: 0.36<He < 0.66).
The Sardinian populations seem to be far more degraded than the Corsican ones. Currently trout populations can be found in a few basins in Sardinia, where they are confined to areas of medium elevation. In this island, known pure natural populations are limited to one southern basin located within the Regional Natural Reserve (Foresta di Monte Arcosu): the Cixerri basins (Sabatini et al., 2006, 2011; Zaccara et al., 2015; Sabatini et al., 2018). Cixerri basin samples (numbers 12–14) were grouped at the second step of hierarchical assignment (Fig. 4). Flumendosa basin (samples 10 and 11), located east-central of the island, is inhabited by hybrid trout with domestic dominance, assigned to the B2 cluster gathering the domestic Atlantic lineage (Fig. 4), with a majority (76%) of hatchery haplotypes (Table 3).
Tiber River trout are probably influenced by the numerous upstream hatcheries settled in the Marche (samples 15 and 16) and Umbria regions. However, this influence seems limited in the wild according to the mitochondrial marker (no AT haplotype upstream in the Nera River and 33% downstream). Nuclear markers clearly isolate a first natural group, the upstream Nera River sample alone, visible outside the central part of the multidimensional diagram (Fig. 3a) in the B1 lineage (Fig. 4) and with haplotypes AD-Tyrrh5 and 6, not observed elsewhere (Table 3). According to microsatellites (Fig. 3c and the B2 cluster in Fig. 4.), another lower Tiber group is genetically closer to the domestic samples (haplotypes AD-Tyrrh 1, 2, and 4, all three not endemic to the Tiber River).
Calabria is a dry area cut at the south by small parallel coastal rivers flowing into the Ionian Sea. Domestic Atlantic strains were introduced into this region, and totally replaced the Allaro population (Table 3, Fig. 3c and B2 cluster in Fig. 4). The Diga Giulia population is hybridized and shows 70% of domestic LDH alleles (*90) and 30% of Atlantic haplotypes (Table 3). Natural Calabrian trout are limited to Ancinale and partly in the Diga Giulia Rivers with haplotypes AD-Tyrrh1 and 3, and ADcs1.
According to Schöffmann et al. (2007) and the present study, Sicily is inhabited by trout belonging only to the AT mitochondrial lineage. Schöffmann et al. (2007) described haplotype AT-s6 in a sample of 26 specimens from three rivers including the River Anapo. The AT-s6 haplotype is a short synonym (380 bp reported by Bernatchez, 2001) of three published longer sequences: the ATSic haplotype (endemic to Sicily, Snoj et al., 2011), and the ATM1 and ATM6 haplotypes, known in Morocco (Snoj et al., 2011). These natural AT haplotypes are phylogenetically far from “haplotypes 1–4” marking domestic AT trout (Snoj et al., 2011). Using one enzymatic marker, Schöffmann et al. (2007) indicated that the LDH-C1*100/100 genotype is dominant in Sicily (24/26). This last genotype is absent from north Atlantic domestic strains (Berrebi et al., 2000; Cortey et al., 2004). It was thus deduced that the Sicilian lineage should be a natural immigration of the south Atlantic lineage, characterized by the LDH-C1*100 allele. Schöffmann et al. (2007) also indicated that the AT-s6 haplotype sequence is close to those found in Atlantic basins in the Iberian Peninsula (Weiss et al., 2000; Suárez et al., 2001) and southern France (Aurelle & Berrebi, 2001). The other haplotype found in Sicily in the Manghisi River (tributary of the River Cassibile), “clone JE1,” was first described in the Jerte River (tributary of the River Tajo) in Spain (Suárez et al., 2001). This haplotype is phylogenetically close to the ATSic natural haplotype but also to several domestic haplotypes like “haplotype 3” (Fig. 2). Therefore, the presence of the sole allele LDH-C1*100 in a sub-sample from the Manghisi River (Table 3) demonstrates the natural origin of the haplotype clone JE1. Taking these new results into account confirm the delimitations of two Atlantic sublineages: the natural one mainly in Sicily and the domestic ones mainly in hatchery samples but dispatched over the whole zone. This is justified by (i) the two exclusive groups of haplotypes, haplotypes 1 to 4 vs ATsic and clone JE1 for domestic and natural AT and (ii) by Fig. 4, where the Atlantic lineage is in black at step 1 and in green and black (B1 and B2) at step 2 for natural and domestic Atlantic sublineages.
Recently, Splendiani et al. (2016) detected the ATsc33 haplotype in sub-fossil remains of brown trout from a Tyrrhenian locality of northern Calabria dating about 13,000-8,000 years BP. This haplotype is considered to belong to the southern Atlantic haplogroup together with ATSic and clone JE1 haplotypes, and was already detected in Sicily (Fruciano et al., 2014), Spain (Suárez et al., 2001; Cortey et al., 2009), and Morocco (Snoj et al., 2011). This observation is coherent with the recent expansion of this haplogroup related to the last glacial maximum as proposed by Snoj et al. (2011).
In the samples from Calabria analyzed in the present study, no native Atlantic haplotypes were found. However, due to the very low size of the Calabrian samples analyzed here (13 sequenced specimens), we cannot make any assumptions about a possible natural presence of the Atlantic lineage in current brown trout populations of the southern Italian peninsula.
Conservation purposes
The brown trout is generally accepted as a diversified and complex assemblage of salmonids of the genus Salmo. It has been investigated for a long time by several methods, but without a clear description of its taxonomy. According to Sanz (2018), who reviewed the entire S. trutta complex data, the origin of this difficulty is due to the multiple episodes of colonization and secondary contacts. This high phylogenetic complexity with large genetic differences within lineages is accompanied by morphological and life-history diversity that has led to the description of dozens of nominal species (Kottelat & Freyhof, 2007).
The increase in the number of new trout species, sometimes elevating known subspecies to the species rank (Isaac et al., 2004; Zachos et al., 2013), sometimes accepting ancient morphological species as true species (Ninua et al., 2018), is a strategy to be considered as a threatened taxon in the IUCN Red List criteria. This should be a help for conservation (Garnett & Christidis, 2017) but such a strategy can lead to mismatches with described molecular lineages (Phillimore & Owens, 2006). Considering the complex evolutionary history of the brown trout, Sanz (2018) recommended defining conservation units based on genetic assessments at the population level.
Thirty years ago, molecular investigations proposed a first global description of the S. trutta complex. The most popular description of genetic structure is based on the mtDNA CR sequences, showing five main clusters closely linked to geographic distribution (Bernatchez et al., 1992; Bernatchez, 2001). This organization has been confirmed with rDNA ITS markers (Presa et al., 2002).
Numerous publications used the CR marker, increasing the number of sequences or the sequence length (Weiss et al., 2000; Cortey & García-Marín, 2002) until sequencing the 1250 bp that covers the whole CR in Giuffra et al. (1994) or 1013 bp in Cortey et al. (2004). This marker is perhaps not the best for phylogenetic reconstruction, but it is the most used and so the most practical for general phylogenies with numerous GenBank sequences involved.
Besides the S. trutta complex, few taxa have been demonstrated as being outside the species complex according to mtDNA phylogenies. S. ohridanus and S. obtusirostris are rare Salmo taxa considered as distinct species (Snoj et al., 2002; Sušnik et al., 2006; Snoj et al., 2009) as is S. salar. At the same time, other morphological species have been returned to the S. trutta complex (S. platycephalus: Sušnik et al., 2004; S. dentex: Snoj et al., 2010).
Most molecular publications apply an exclusivity criterion which is the reciprocal monophyly, the basis of the phylogenetic species concept (Hebert et al., 2003; Sites & Marshall, 2004), so that taxonomy is as close as possible to the phylogeny. However, discrepancies between morphology and genetics (especially for recent species) should be explained by the expected discrepancy between gene history (Avise 2000) and/or gene phylogeny and species phylogeny (Knowles & Carstens, 2007).
According to Moritz (1994) and to the arguments given here, conservation units should be management units (MUs), i.e., conservation units based on molecular assessments at the population level, without phylogenetic prerequisites, considering population assemblages forming units that deserve distinct management for conservation.
In the Tyrrhenian region, several attempts at nomenclature have been made. The proposition of Kottelat & Freyhof (2007) to retain the name S. cettii for the whole Tyrrhenian region ignores the local high diversity of trout populations and numerous genetic subgroups observed with two molecular markers (Figs. 2, 3 and 4). Trout in the Tyrrhenian Sea region were rarely investigated with molecular methods and never as a whole in order to distinguish the different taxa and conservation units, although these data are necessary for intelligent management and prioritization of stock preservation. The present survey demonstrated that several categories of molecular markers are necessary for this. Crossing the well visible clusters in Figs. 2 and 3a and b and the distinct lineages obtained by assignment (Fig. 4), the following significantly differentiated MUs can be proposed:
The eight Corsican samples (U Furcone, A Tassineta, Aqua d’Acelli, E Maghjine, Speloncellu, Pozzi, Val d’Ese, and Lataga Rivers) and the three Sardinian samples of the Cixerri basin divided into two MUs (Marroccu and Is Abius-Camboni) constitute ten MUs, genetically clearly different, mostly grouped in cluster A of Fig. 4. They constitute the island part of the Tyrrhenian trout diversity. The detection of these MUs allows much more isolated and differentiated populations to be described in Corsica. These island MUs are almost not subject to domestic admixture.
Secondly, continental Italy revealed only two MUs positioned in cluster B1 of Fig. 4, i.e., dominated by natural trout: (i) the Nera River (sample 17, central Italy) and (ii) the Ancinale River (sample 22, Calabria), which seems to be the only local population dominated by a natural lineage.
Third, the Sicilian trouts, confirmed in their natural AT lineage, are nevertheless separated into two MUs: (i) the well-known Anapo River (Schöffmann et al., 2007) characterized by haplotype ATSic and (ii) the pure wild populations of Manghisi and San Marco with the “clone JE1” haplotype, although the San Marco River specimens were not haplotyped.
Conclusions
Analyzed as a whole for the Tyrrhenian Sea region, trout have revealed their geographical genetic organization.
The northern islands host precious MUs (not seen elsewhere) with special mention of Corsica, where the relative good health of very small populations suggests that dozens of MUs need protection. Sardinia is similar except that very few trout populations remain for investigation with molecular markers: two MUs could be defined in the Cixerri basin. Sicily is similar to Sardinia since only the south of the island is populated by small populations, but at least two MUs were detected. We do not know if the limited trout range in Sicily is natural or due to human activity.
Populations from continental Italy appear less structured, probably as a consequence of both fewer sampling efforts and a strong influence of genetic introgression with the Atlantic genome of domestic origin. Here, therefore, on one hand, two MUs are still recognizable (in central and south Tyrrhenian Italy). On the other hand, caution should be taken for estimation of the number of MUs detected. In future, more effort should be made to include samples from other parts of Tyrrhenian Italy where the historical presence of native brown trout genetic diversity has been highlighted (e.g., Splendiani et al., 2017; Fabiani et al., 2018).
The maintenance of these natural lineages, some without domestic introgression, allows optimism regarding conservation of Tyrrhenian trout, which is indispensable for their role in freshwater biodiversity. In view of the great genetic diversity that characterizes Mediterranean trout and the risks of undesirable genetic homogenization by domestic forms, an intelligent conservation strategy should never be carried out without preliminary genetic description of the populations involved in conservation actions. Unfortunately, the number of conservation programs carried out without any kind of control is becoming more and more frequent.
References
Almodóvar, A., G. G. Nicola, B. Elvira & J. L. García-Marín, 2006. Introgression variability among Iberian brown trout evolutionary significant units: the influence of local management and environmental features. Freshwater Biology 51: 1175–1187.
Angers, B., L. Bernatchez, A. Angers & L. Desgroseillers, 1995. Specific microsatellite loci for brook charr (Salvelinus fontinalis Mitchill) reveal strong population subdivision on a microgeographic scale. Journal of Fish Biology 48: 177–185.
Araguas, R. M., M. Vera, E. Aparicio, N. Sanz, R. Fernandez-Cebrian, C. Marchante & J. L. García-Marín, 2017. Current status of the brown trout (Salmo trutta) populations within eastern Pyrenees genetic refuges. Ecology of Freshwater Fish 26: 120–132.
Aurelle, D. & P. Berrebi, 2001. Genetic structure of brown trout (Salmo trutta, L.) populations from south-western France: data from mitochondrial control region variability. Molecular Ecology 10: 1551–1561.
Aurelle, D. & P. Berrebi, 2002. Natural and artificial secondary contact in brown trout (Salmo trutta, L.) in the French western Pyrenees assessed by allozymes and microsatellites. Heredity 89: 171–183.
Avise, J. C., 2000. Phylogeography—the history and formation of species. Harvard University Press, Cambridge.
Bardakci, F., N. Degerli, O. Ozdemir & H. H. Basibuyuk, 2006. Phylogeography of the Turkish brown trout Salmo trutta L.: mitochondrial DNA PCR-RFLP variation. Journal of Fish Biology 68: 36–55.
Behnke, R. J., 1972. The systematics of salmonid fishes of recently glaciated lakes. Journal of the Fisheries Research Board of Canada 29: 639–671.
Behnke, R. J., 1986. Brown trout. Trout 27: 42–47.
Belkhir, K., P. Borsa, J. Goudet & F. Bonhomme, 2004. GENETIX 4.05: logiciel sous Windows pour la génétique des populations. Laboratoire Génome et Population, CNRS-UPR, Université de Montpellier II, Montpellier, France.
Benzécri, J.-P., 1973. L’analyse des données. Dunod, Paris.
Bernatchez, L., 2001. The evolutionary history of brown trout (Salmo trutta L.) inferred from phylogeographic, nested clade, and mismatch analyses of mitochondrial DNA variation. Evolution 55: 351–379.
Bernatchez, L. & A. Osinov, 1995. Genetic diversity of trout (genus Salmo) from its most eastern native range based on mitochondrial DNA and nuclear gene variation. Molecular Ecology 4: 285–297.
Bernatchez, L., R. Guyomard & F. Bonhomme, 1992. DNA sequence variation of the mitochondrial control region among geographically and morphologically remote European brown trout Salmo trutta populations. Molecular Ecology 1: 161–173.
Berrebi, P., 2015. Three brown trout Salmo trutta lineages in Corsica described through allozyme variation. Journal of Fish Biology 86: 60–73.
Berrebi, P., C. Poteaux, M. Fissier & G. Cattaneo-Berrebi, 2000. Stocking impact and allozyme diversity in brown trout from Mediterranean southern France. Journal of Fish Biology 56: 949–960.
Berrebi, P., C. Tougard, S. Dubois, Z. Shao, I. Koutseri, S. Petkovski & A. J. Crivelli, 2013. Genetic diversity and conservation of the Prespa trout in the Balkans. International Journal of Molecular Sciences 14: 23454–23470.
Bianco, P. G., 2014. An update on the status of native and exotic freshwater fishes of Italy. Journal of Applied Ichthyology 30: 62–77.
Bohling, J., P. Haffray & P. Berrebi, 2016. Genetic diversity and population structure of domestic brown trout (Salmo trutta) in France. Aquaculture 462: 1–9.
Boulenger, G. A., 1901. On the occurrence of Salmo macrostigma in Sardinia. Annals and Magazine of Natural History 8(43): 14.
Buroker, N. E., J. R. Brown, T. A. Gilbert, P. J. O’Hara, A. T. Beckenbach, W. K. Thomas & M. J. Smith, 1990. Length heteroplasmy of sturgeon mitochondrial DNA, an illegitimate elongation model. Genetics 124: 157–163.
Charles, K., R. Guyomard, B. Hoyheim, D. Ombredane & J.-L. Baglinière, 2005. Lack of genetic differentiation between anadromous and resident sympatric brown trout (Salmo trutta) in a Normandy population. Aquatic Living Resources 18: 65–69.
Clement, M., D. Posada & K. A. Crandall, 2000. TCS: a computer program to estimate gene genealogies. Molecular Ecology 9: 1657–1660.
Cortey, M. & J. L. García-Marín, 2002. Evidence for phylogeographically informative sequence variation in the mitochondrial control region of Atlantic brown trout. Journal of Fish Biology 60: 1058–1063.
Cortey, M., C. Pla & J. L. García-Marín, 2004. Historical biogeography of Mediterranean trout. Molecular Phylogenetics and Evolution 33: 831–844.
Cortey, M., M. Vera, C. Pla & J. L. García-Marín, 2009. Northern and Southern expansions of Atlantic brown trout (Salmo trutta) populations during the Pleistocene. Biological Journal of the Linnean Society 97: 904–917.
De Moor, I. J. & M. N. Bruton, 1988. Atlas of alien and translocated indigenous aquatic animals in southern Africa. National Scientific Programmes Unit: CSIR, SANSP Report 144: 317.
Delling, B., 2010. Diversity of western and southern Balkan trouts, with the description of a new species from the Louros River, Greece (Teleostei: Salmonidae). Ichthyological Exploration of Freshwaters 21: 331–344.
Delling, B. & I. Doadrio, 2005. Systematics of the trouts endemic to Moroccan lakes, with description of a new species (Teleostei: Salmonidae). Ichthyological Exploration of Freshwaters 16: 49–64.
Doadrio, I., S. Perea & A. Yahyaoui, 2015. Two new species of Atlantic trout (Actinopterygii, Salmonidae) from Morocco. Graellsia 71(2): e031.
Duchi, A., 2011. Fecundity, egg and alevin size in the River Irminio population of the threatened Salmo cettii Rafinesque-Schmaltz, 1810 (Sicily, Italy). Journal of Applied Ichthyology 27: 868–872.
Duchi, A., 2018. Flank spot number and its significance for systematics, taxonomy and conservation of the near-threatened Mediterranean trout Salmo cettii: evidence from a genetically pure population. Journal of Fish Biology 92: 254–260.
Estoup, A., C. R. Largiader, E. Perrot & D. Chourrout, 1996. Rapid one-tube DNA extraction for reliable PCR detection of fish polymorphic markers and transgenes. Molecular Marine Biology and Biotechnology 5: 295–298.
Evanno, G., S. Regnaut & J. Goudet, 2005. Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Molecular Ecology 14: 2611–2620.
Fabiani, A., P. Gratton, I. A. Zappes, M. Seminara, A. D’Orsi, V. Sbordoni & G. Allegrucci, 2018. Investigating the genetic structure of trout from the Garden of Ninfa (central Italy): suggestions for conservation and management. Fisheries Management and Ecology 25: 1–11.
Fruciano, C., A. M. Pappalardo, C. Tigano & V. Ferrito, 2014. Phylogeographical relationships of Sicilian brown trout and the effects of genetic introgression on morphospace occupation. Biological Journal of Linnean Society 112: 387–398.
Gandolfi, G., S. Zerunian, P. Torricelli & A. Marconato, 1991. I Pesci delle acque interne italiane. Ministero dell’Ambiente, Instituto Poligrafico e Zecca dello Stato, Rome. pp. 616.
García-Marín, J.-L., F. M. Utter & C. Pla, 1999. Postglacial colonization of brown trout in Europe based on distribution of allozyme variants. Heredity 82: 46–56.
Garnett, S. T. & L. Christidis, 2017. Taxonomy anarchy hampers conservation—the classification of complex organisms is in chaos. Nature 546: 25–27.
Gauthier, A. & P. Berrebi, 2007. La colonisation de l’île par différentes souches de truite. Guide de gestion de la truite macrostigma. pp 4–10.
Giuffra, E., L. Bernatchez & R. Guyomard, 1994. Mitochondrial control region and protein coding genes sequence variation among phenotypic forms of brown trout Salmo trutta from northern Italy. Molecular Ecology 3: 161–171.
Giuffra, E., R. Guyomard & G. Forneris, 1996. Phylogenetic relationships and introgression patterns between incipient parapatric species of Italian brown trout (Salmo trutta L. complex). Molecular Ecology 5: 207–220.
Gratton, P., G. Allegrucci, V. Sbordoni & A. Gandolfi, 2014. The evolutionary jigsaw puzzle of the surviving trout (Salmo trutta L. complex) diversity in the Italian region. A multilocus Bayesian approach. Molecular Phylogenetics and Evolution 79: 292–304.
Guyomard, R., 1989. Diversité génétique de la truite commune. Bulletin Français de Pêche et de Pisciculture 314: 118–135.
Hamilton, K. E., A. Ferguson, J. B. Taggart, T. Tomasson, A. Walker & E. Fahy, 1989. Post-glacial colonisation of brown trout, Salmo trutta L.: Ldh-5 as a phylogeographic marker locus. Journal of Fish Biology 35: 651–664.
Hebert, P. D. N., A. Cywinska, S. L. Ball & J. R. DeWaard, 2003. Biological identifications through DNA barcodes. Proceedings of the Royal Society Series B, Biological Sciences 270: 313–321.
Holm, L. E. & C. Bendixen, 2000. Oncorhynchus mykiss clone TAA72-13, sequence tagged site [available on internet at http://www.ncbi.nlm.nih.gov/genbank/]. Accession number AF239038.
Isaac, N. J. B., J. Mallet & G. M. Mace, 2004. Taxonomic inflation: its influence on macroecology and conservation. Trends in Ecology and Evolution 19: 464–469.
Jones, O. R. & J. I. Wang, 2010. COLONY: a program for parentage and sibship inference from multilocus genotype data. Molecular Ecology Resources 10: 551–555.
Knowles, L. L. & B. C. Carstens, 2007. Delimiting species without monophyletic gene trees. Systematic Biology 56(6): 887–895.
Kottelat, M., 1997. European freshwater fishes. An heuristic checklist of the freshwater fishes of Europe (exclusive of former USSR), with an introduction for non-systematists and comments on nomenclature and conservation. Biologia, Bratislava 52 (supplement 5): 1–271.
Kottelat, M. & J. Freyhof, 2007. Handbook of European Freshwater Fishes. Publications Kottelat, Cornol, CH, xiv + 646 pp.
Largiadèr, C. R. & A. Scholl, 1996. Genetic introgression between native and introduced brown trout (Salmo trutta L.) populations in the Rhone River Basin. Molecular Ecology 5: 417–426.
Lucentini, L., A. Palomba, L. Gigliarelli, H. Lancioni, P. Viali & F. Panara, 2006. Genetic characterization of a putative indigenous brown trout (Salmo trutta fario) population in a secondary stream of the Nera River Basin (Central Italy) assessed by means of three molecular markers. Italian Journal of Zoology 73: 263–273.
Marić, S., S. Sušnik Bajec, J. Schöffmann, V. Kostov & A. Snoj, 2017. Phylogeography of stream-dwelling trout in the Republic of Macedonia and a molecular genetic basis for revision of the taxonomy proposed by S. Karaman. Hydrobiologia 785: 249–260.
Massidda, P., 1995. Salmo (trutta) macrostigma in Sardegna. Biologia Ambientale 5: 40–43.
McMeel, O. M., E. M. Hoey & A. Ferguson, 2001. Partial nucleotide sequences, and routine typing by polymerase chain reaction-restriction fragment length polymorphism, of the brown trout (Salmo trutta) lactate dehydrogenase, LDH-C1*90 and *100 alleles. Molecular Ecology 10: 29–34.
Meldgaard, T., A. J. Crivelli, D. Jesensek, G. Poizat, J.-F. Rubin & P. Berrebi, 2007. Hybridization mechanisms between the endangered marble trout (Salmo marmoratus) and the brown trout (Salmo trutta) as revealed by in-stream experiments. Biological Conservation 136: 602–611.
Miró, A. & M. Ventura, 2013. Historical use, fishing management and lake characteristics explain the presence of non-native trout in Pyrenean lakes: Implications for conservation. Biological Conservation 167: 17–24.
Mola, P., 1928. Facies della fauna e della flora delle acque interne. Contributo alla idrobiologia della Sardegna. Internationale Revue der gesamten Hydrobiologie und Hydrographie 20: 117–173.
Moritz, C., 1994. Defining evolutionarily significant units for conservation. Trends in Ecology and Evolution 9: 373–375.
Nei, M., 1978. Estimation of average heterozygosity and genetic distance from a small number of individuals. Genetics 89: 583–590.
Ninua, L., D. Tarkhnishvili & E. Gvazava, 2018. Phylogeography and taxonomic status of trout and salmon from the Ponto-Caspian drainages, with inferences on European brown trout evolution and taxonomy. Ecology and Evolution. https://doi.org/10.1002/ece3.3884.
Nonnis Marzano, F., N. Corradi, R. Papa, J. Tagliavini & G. Gandolfi, 2003. Molecular evidence for introgression and loss of genetic variability in Salmo (trutta) macrostigma as a result of massive restocking of Apennine populations (Northern and Central Italy). Environmental Biology of Fishes 68: 349–356.
Olsson, K., P. Stenroth, P. Nyström, N. Holmqvist, A. R. McIntosh & M. J. Winterbourn, 2006. Does natural acidity mediate interactions between introduced brown trout, native fish, crayfish and other invertebrates in West Coast New Zealand streams? Biological Conservation 130: 255–267.
O’Reilly, P. T., L. C. Hamilton, S. K. McConnell & J. M. Wright, 1996. Rapid analysis of genetic variation in Atlantic salmon (Salmo salar) by PCR multiplexing of dinucleotide and tetranucleotide microsatellites. Canadian Journal of Fisheries and Aquatic Sciences 53: 2292–2298.
Orrù, F., A. M. Deiana & A. Cau, 2010. Introduction and distribution of alien freshwater fishes on the island of Sardinia (Italy): an assessment on the basis of existing data sources. Journal of Applied Ichthyology 26(2): 46–52.
Patarnello, T., L. Bargelloni, F. Caldara & L. Colombo, 1994. Cytochrome b and 16S rRNA sequence variation in the Salmo trutta (Salmonidae, Teleostei) species complex. Molecular Phylogenetics and Evolution 3: 69–74.
Phillimore, A. B. & I. P. F. Owens, 2006. Are subspecies useful in evolutionary and conservation biology? Proceedings of the Royal Society of London, Series B, Biological Sciences 273: 1049–1053.
Pomini, F. P., 1940. Ricerche sul Salmo macrostigma Dum. Bollettino di Pesca, Piscicoltura e Idrobiologia 16(3): 12–63.
Popa, G.-O., A. Curtean-Bănăduc, D. Bănăduc, I. E. Florescu, A. Burcea, A. Dudu, S. E. Georgescu & M. Costache, 2016. Molecular markers reveal reduced genetic diversity in Romanian populations of brown trout, Salmo trutta L., 1758 (Salmonidae). Acta Zoologica Bulgarica 68: 399–406.
Presa, P. & R. Guyomard, 1996. Conservation of microsatellites in three species of salmonids. Journal of Fish Biology 49: 1326–1329.
Presa, P., F. Krieg, A. Estoup & R. Guyomard, 1994. Diversité et gestion génétique de la truite commune: apport de l’étude du polymorphisme des locus protéiques et microsatellites. Genetics Selection Evolution 26(suppl. 1): 183–202.
Presa, P., B. G. Pardo, P. Martinez & L. Bernatchez, 2002. Phylogeographic congruence between mtDNA and rDNA ITS markers in brown trout. Molecular Biology and Evolution 19(12): 2161–2175.
Pritchard, J. K., M. Stephens & P. Donnelly, 2000. Inference of population structure using multilocus genotype data. Genetics 155: 945–959.
Rexroad III, C. E., R. L. Coleman, W. K. Hershberger & J. Killefer, 2002. Thirty-eight polymorphic microsatellite markers for mapping in rainbow trout. Journal of Animal Science 80: 541–542.
Rice, W. R., 1989. Analyzing tables of statistical tests. Evolution 43: 223–225.
Roule, L., 1933. Le peuplement des cours d’eau de la Corse en poissons. Bulletin Français de Pisciculture 63: 61–62.
Sabatini, A., F. Orrù, R. Cannas, P. Serra & P. Cau, 2006. Conservation and management of Salmo (trutta) macrostigma in Sardinian freshwaters: first results of genetic characterization. Quaderni ETP 34: 335–340.
Sabatini, A., R. Cannas, M. C. Follesa, F. Palmas, A. Manunza, G. Matta, A. A. Pendugiu, P. Serra & A. Cau, 2011. Genetic characterization and artificial reproduction attempt of endemic Sardinian trout Salmo trutta L., 1758 (Osteichthyes, Salmonidae): Experiences in captivity. Italian Journal of Zoology 78(1): 20–26.
Sabatini, A., C. Podda, G. Frau, M. V. Cani, A. Musu, M. Serra & F. Palmas, 2018. Restoration of native Mediterranean trout Salmo cetti Rafinesque, 1810 (Actinopterygii, Salmonidae) populations using an electric barrier as mitigation tool. The European Zoological Journal. https://doi.org/10.1080/24750263.2018.1435664.
Sanz, N., 2018. Phylogeographic history of brown trout: a review. In Lobon-Cervia, J. & N. Sanz (eds), Brown trout: biology, ecology and management. Wiley, Hoboken, NJ: 17–63.
Schöffmann, J., S. Sušnik & A. Snoj, 2007. Phylogenetic origin of Salmo trutta L 1758 from Sicily, based on mitochondrial and nuclear DNA analyses. Hydrobiologia 575: 51–55.
Scribner, K. T., J. R. Gust & R. L. Fields, 1996. Isolation and characterization of novel salmon microsatellite loci: cross-species amplification and population genetic applications. Canadian Journal of Fisheries and Aquatic Sciences 53: 833–841.
She, J. X., M. Autem, G. Kotoulas, N. Pasteur & F. Bonhomme, 1987. Multivariate analysis of genetic exchanges between Solea aegyptiaca and Solea senegalensis (Teleosts, Soleidae). Biological Journal of the Linnean Society 32: 357–371.
Sites, J. W. & J. C. Marshall, 2004. Empirical criteria for delimiting species. Annual Review of Ecology, Evolution, and Systematics 35: 199–227.
Slettan, A., I. Olsaker & Ø. Lie, 1995. Atlantic salmon, Salmo salar, microsatellites at the SSOSL25, SSOSL85, SSOSL311, SSOSL417 loci. Animal Genetics 26: 277–285.
Slettan, A., I. Olsaker & Ø. Lie, 1996. Polymorphic Atlantic salmon, Salmo salar, microsatellites at the SSOSL438, SSOSL439 and SSOSL444 loci. Animal Genetics 27: 57–64.
Snoj, A., E. Melkic, S. Sušnik, S. Muhamedagic & P. Dovc, 2002. DNA phylogeny supports revised classification of Salmothymus obtusirostris. Biological Journal of the Linnean Society 77: 397–411.
Snoj, A., S. Marić, P. Berrebi, A. J. Crivelli, S. Shumka & S. Sušnik, 2009. Genetic architecture of trout from Albania as revealed by mtDNA control region variation. Genetics Selection Evolution 41: 22.
Snoj, A., B. Glamuzina, A. Razpet, J. Zablocki, I. Bogut, E. Lerceteau-Kohler, N. Pojskic & S. Sušnik, 2010. Resolving taxonomic uncertainties using molecular systematics: Salmo dentex and the Balkan trout community. Hydrobiologia 651: 199–212.
Snoj, A., S. Marić, S. Sušnik Bajec, P. Berrebi, S. Janjani & J. Schöffmann, 2011. Phylogeographic structure and demographic patterns of brown trout in North-West Africa. Molecular Phylogenetics and Evolution 61: 203–211.
Splendiani, A., T. Fioravanti, M. Giovannotti, A. Negri, P. Ruggeri, L. Olivieri, P. Nisi Cerioni, M. Lorenzoni & V. Caputo Barucchi, 2016. The effects of paleoclimatic events on Mediterranean trout: preliminary evidences from ancient DNA. PLoS ONE 11(6): e0157975.
Splendiani, A., T. Fioravanti, M. Giovannotti, L. Olivieri, P. Ruggeri, P. Nisi Cerioni, S. Vanni, F. Enrichetti & V. Caputo Barucchi, 2017. Museum samples could help to reconstruct the original distribution of Salmo trutta complex in Italy. Journal of Fish Biology 90: 2443–2451.
Stelkens, R. B., G. Jaffuel, M. Escher & C. Wedekind, 2012. Genetic and phenotypic population divergence on a microgeographic scale in brown trout. Molecular Ecology 21: 2896–2915.
Suárez, J., J. M. Bautista, A. Almodóvar & A. Machordom, 2001. Evolution of the mitochondrial control region in Paleartic brown trout (Salmo trutta) populations: the biogeographical role of the Iberian Peninsula. Heredity 87: 198–206.
Sušnik, S., J. Schöffmann & A. Snoj, 2004. Phylogenetic position of Salmo (Platysalmo) platycephalus Behnke 1968 from south-central Turkey, evidenced by genetic data. Journal of Fish Biology 64: 947–960.
Sušnik, S., I. Knizhin, A. Snoj & S. Weiss, 2006. Genetic and morphological characterization of a Lake Ohrid endemic, Salmo (Acantholingua) ohridanus with a comparison to sympatric Salmo trutta. Journal of Fish Biology 68: 2–23.
Sušnik, S., A. Snoj, I. F. Wilson, D. Mrdak & S. Weiss, 2007. Historical demography of brown trout (Salmo trutta) in the Adriatic drainage including the putative S. letnica endemic to Lake Ohrid. Molecular Phylogenetics and Evolution 44: 63–76.
Tamura, K., G. Stecher, D. Peterson, A. Filipski & S. Kumar, 2013. MEGA6: molecular evolutionary genetics analysis version 6.0. Molecular Biology and Evolution 30: 2725–2729.
Thompson, J. D., T. J. Gibson, F. Plewniak, F. Jeanmougin & D. G. Higgins, 1997. The CLUSTAL_X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Research 25: 4876–4882.
Turan, D., M. Kottelat & S. Engin, 2009. Two new species of trouts, resident and migratory, sympatric in streams of northern Anatolia (Salmoniformes: Salmonidae). Ichthyological Exploration of Freshwaters 20: 333–364.
Turan, D., M. Kottelat & S. Engin, 2010. Two new species of trouts, resident and migratory, sympatric in streams of northern Anatolia (Salmoniformes: Salmonidae). Ichthyological Exploration of Freshwaters 20: 289–384.
Turan, D., M. Kottelat & Y. Bektas, 2011. Salmo tigridis, a new species of trout from the Tigris River, Turkey (Teleostei: Salmonidae). Zootaxa 2993: 23–33.
Turan, D., M. Kottelat & S. Engin, 2012. The trouts of the Mediterranean drainages of southern Anatolia, Turkey, with description of three new species (Teleostei: Salmonidae). Ichthyological Exploration of Freshwaters 23: 219–236.
Turan, D., E. Doğan, C. Kaya & M. Kanyılmaz, 2014a. Salmo kottelati, a new species of trout from Alakır Stream, draining to the Mediterranean in southern Anatolia, Turkey (Teleostei, Salmonidae). ZooKeys 462: 135–151.
Turan, D., M. Kottelat & S. Engin, 2014b. Two new species of trouts from the Euphrates drainage, Turkey (Teleostei: Salmonidae). Ichthyological Exploration of Freshwaters 24: 275–287.
Vähä, J. P., J. Erkinaro, E. Niemelä & C. R. Primmer, 2007. Life-history and habitat features influence the within-river genetic structure of Atlantic salmon. Molecular Ecology 16: 2638–2654.
van Oosterhout, C., W. F. Hutchinson, D. P. Wills & P. Shipley, 2004. MICRO-CHECKER: software for identifying and correcting genotyping errors in microsatellite data. Molecular Ecology Notes 4: 535–538.
Walsh, P. S., D. A. Metzger & R. Higushi, 1991. Chelex 100 as a medium for simple extraction of DNA for PCR-based typing from forensic material. Biotechniques 10(4): 506–513.
Weir, B. S. & C. C. Cockerham, 1984. Estimating F-statistics for the analysis of population structure. Evolution 38: 1358–1370.
Weiss, S., A. Antunes, C. Schlötterer & P. Alexandrino, 2000. Mitochondrial haplotype diversity among Portuguese trout Salmo trutta L. populations: relevance of the post-Pleistocene recolonization of northern Europe. Molecular Ecology 9: 691–698.
Yue, G. H. & L. Orban, 2005. A simple and affordable method for high-throughput DNA extraction from animal tissues for polymerase chain reaction. Electrophoresis 26: 3081–3083.
Zaccara, S., S. Trasforini, C. M. Antognazza, C. Puzzi, J. R. Britton & G. Crosa, 2015. Morphological and genetic characterization of Sardinian trout Salmo cettii Rafinesque, 1810 and their conservation implications. Hydrobiologia 760: 205–223.
Zachos, F. E., M. Apollonio, E. V. Barmann, M. Festa-Bianchet, U. Gohlich, J. C. Habel, E. Haring, L. Kruckenhauser, S. Lovari, A. D. McDevitt, C. Pertoldi, G. E. Rossner, M. R. Sanchez-Villagra, M. Scandura & F. Suchentrunk, 2013. Species inflation and taxonomic artefacts—a critical comment on recent trends in mammalian classification. Mammalian Biology 78: 1–6.
Acknowledgements
The authors thank Bernard Lasserre, Sophie Dubois, Yuki Minegishi, and Zhaojun Shao for laboratory technical help, Pietro Armenia, Giuseppe Pisani, and Andrea Belluscio for field help, and David Schikorski (of the private company Labofarm-Genindexe—France) for genotyping most of the considered trouts. Thank you to Martin Laporte and Séverine Roques who participated to statistical solving. Some of the samples were constituted by the French anglers’ Federations of Alpes Maritimes and Corsica and the Genesalm project (CIPA). They are warmly thanked. Sicilian samples were collected with permission from the Regional Province of Syracuse and the professional assistance of Pietro Armena. In Calabria and Lazio (Rome), corresponding managers (Giuseppe Pisani in Calabria and Andrea Belluscio in Lazio) linked us with professional fishermen for sampling. We thank them for their irreplaceable assistance. The authors would also like to acknowledge OSU-OREME and Juliette Fabre for their help in database construction and mapping. Saša Marić was supported by the Ministry of Education, Science and Technological Development of the Republic of Serbia (Grant No. 173045).
Author information
Authors and Affiliations
Corresponding author
Additional information
Handling editor: Cécile Fauvelot
Electronic supplementary material
Below is the link to the electronic supplementary material.
10750_2018_3734_MOESM1_ESM.docx
Table S1: Accession numbers of all haplotypes positioned in Fig. 2 (* = new), completed with their geographic distribution according to twenty main publications. Supplementary material 1 (DOCX 34 kb)
10750_2018_3734_MOESM2_ESM.docx
Table S2: Hierarchical steps in estimating K (the number of genetic clusters) from STRUCTURE runs using the ΔK method. L(K) - posterior probability of K; stdev - standard deviation of L(K) from seven independent runs; ΔK - an ad hoc quantity, predictor of the real number of clusters (Evanno et al., 2005), best ΔK are highlighted. Supplementary material 2 (DOCX 33 kb)
10750_2018_3734_MOESM4_ESM.docx
Fig. S4: New projection of the FCA analysis presented in Fig. 3a according to axes 1 and 3. In this new perspective, Sardinian samples and E Maghine Corsican one, together with the Corsican samples Lataga and Aqua d’Acelli, which seemed similar in Fig. 3a, are in fact clearly different (red arrows) while Corsican samples Pozzi and Val d’Ese confirm their similarity. Supplementary material 4 (DOCX 150 kb)
10750_2018_3734_MOESM5_ESM.jpg
Fig. S5: Estimation of the number of genetic clusters (K) for the first level, from STRUCTURE runs using the ΔK method. Supplementary material 5 (JPEG 132 kb)
10750_2018_3734_MOESM6_ESM.jpg
Fig. S6. Estimated population structure as inferred by STRUCTURE analysis of microsatellite marker DNA data. White lines separate sampling sites, the most probable K = 19 is based on maximizing the mean estimated ln probability of data (Pritchard et al., 2000). Names and codes of sampling sites/clusters are reported in Tables 1 and 4 and Fig. 4. Supplementary material 6 (JPEG 660 kb)
Rights and permissions
About this article
Cite this article
Berrebi, P., Caputo Barucchi, V., Splendiani, A. et al. Brown trout (Salmo trutta L.) high genetic diversity around the Tyrrhenian Sea as revealed by nuclear and mitochondrial markers. Hydrobiologia 826, 209–231 (2019). https://doi.org/10.1007/s10750-018-3734-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10750-018-3734-5