Abstract
Anthocyanins occur in potato tuber skin and flesh, sprouts, leaves, stems and flowers. The goal of this study was to identify genomic regions and candidate gene alleles key for accumulation of anthocyanins in potato corolla in various quantities. QTL analyses were performed in two mapping populations segregating for flower colour intensity and candidate genes were identified on the basis of function and location (chalcone isomerase, chi; chalcone synthase, chs) or location (RNA-dependent RNA polymerase 1, RDR1). We detected three and four QTL affecting the violet flower colour intensity using the two mapping populations, respectively. In both populations a locus F, necessary for violet flower colour, segregated and we used different approaches to differentiate the qualitative effect of this locus and to detect the genetic factors affecting the quantitative flower colour intensity. The strongest QTL and the only one common for the two mapping populations was located on chromosome V. The role of all three candidate genes, chi, chs and RDR1, in control of flower colour intensity is supported to different extents by the performed genetic analyses. The most important QTL on chromosome V is most likely in the same position as the QTL for anthocyanin tuber flesh coloration described previously, which indicates that the natural variation in some biosynthetic and/or regulatory genes may influence anthocyanin levels in multiple tissues.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In the 16th century, potato (Solanum tuberosum L.) was brought to Europe and grown as an ornamental curiosity or a medicinal plant. It was valued for white or violet flowers, until the nutritional value of the tubers was appreciated and popularized. Now, the potato is the third most important food crop worldwide with the global production that reached 385 million tons in 2014 (FAOSTAT). Numerous cultivars of potato suitable for diversified purposes have been bred with focus on the agronomic and tuber quality traits. However, some variation of flower colour is still present in the cultivated potato gene pool. Among cultivars in The European Cultivated Potato Database (www.europotato.org), the cultivars with white flowers dominate (1477 cultivars), but ones with red violet (887), blue violet (148) and light blue (38) flowers are also present. Potato can also have pigmented tuber skin and flesh, sprouts, leaves and stems. Anthocyanins in potato tuber skin and flesh are not only serving as a genetic model but, just like in other anthocyanin-rich crops and vegetables, enhance the market value of the plant by increasing its visual appeal to the consumer, reducing softening, shrivelling and fungal rots (Passeri et al. 2016). What is also important, plant products rich in anthocyanins are health-promoting components of the human diet. The health benefits associated with anthocyanins intake include: prevention of the degenerative diseases and cancer, reduction of retinal damage, decreasing inflammatory markers expression, reduction of risk of type-2 diabetes and reduction of weight gain (Passeri et al. 2016).
The flower colour has been studied in potato as one of the model traits for genetics. Potato flowers are not white, when the dominant allele F is present at locus on potato chromosome X and when there is at least one dominant allele in loci P or R that confers general ability to synthesise anthocyanin pigments (van Eck et al. 1993, 1994). The F locus was also mapped in tetraploid potato and located on DM1-3 physical map to superscaffold PGSC0003DMB000000106 (Hackett et al. 2014). Loci P and R affect the anthocyanin presence and composition in all parts of the potato plant. The pigmentation of potato petals, but also tuber skin and flesh vary from blue violet based on petunidin (delphinidin derivative) and conferred by locus P on chromosome XI to pelargonidin-based red violet encoded in locus R (also known as D) on chromosome II and the presence of diversified alleles of these loci in different combinations results in various shades of violet (Salaman 1910; Dodds and Long 1955; van Eck et al. 1993, 1994; Lewis et al. 1998).
De Jong et al. (2004) has demonstrated that map position of the locus P corresponds to the location of gene coding flavonoid 3′,5′-hydroxylase (F3′,5′H) in tomato. The gene has been then cloned from potato and its co-segregation with violet tuber colour was described in F1 diploid potato family. Furthermore, the red-skinned tubers of cultivar Desiree have been changed to violet after transforming the cultivar with the “violet” allele of the F3′,5′H, thus confirming that locus P encodes this enzyme (Jung et al. 2005). In tomato a gene encoding flavonoid 3′,5′-hydroxylase was shown to accept flavones, flavanones, dihydroflavonols and flavonols as substrates as well as to respond transcriptionally to nitrogen deprivation (Olsen et al. 2010).
Dihydroflavonol 4-reductase (DFR) has been associated with the ability for red anthocyanin synthesis in potato (De Jong et al. 2003), which was later supported by mapping in tomato of the gene encoding the enzyme to the position overlapping with locus R on chromosome II (De Jong et al. 2004). Further evidence that the locus R encodes for DFR was obtained by transformation. Introduction of “red” allele of DFR to cultivar Prince Hairy with white tuber and pale violet flowers, has changed the flower colour to red violet and, when an appropriate allele of tissue-specific regulatory gene developer (D) was introduced by crossing, the tuber skin colour changed to red violet, too (Zhang et al. 2009a).
Both F3′,5′H and DFR have been manipulated in order to change the colour of potato tubers and, for example, the petals of roses and carnations to violet but to change the quantity of the anthocyanins present in plant tissue, a variety of transcription factors have been targeted by genetic modifications and classical plant breeding. However, it may not be sufficient to enhance the synthesis of the anthocyanins for increased contents of these pigments since their final yield is affected also by their degradation in vacuoles (reviewed by Passeri et al. 2016).
In potato, an R2R3 MYB transcription factor, designated Stan2 (homolog of petunia’s ANTHOCYANIN2, AN2), was shown to be encoded in locus D (also known as I) (Salaman 1910; De Jong 1987) on chromosome X controlling the presence of anthocyanins in tuber skin. An allele of Stan2 cosegregating with red tuber skin colour was used for transformation of red-skinned cultivar Desiree as well as white-skinned Bintje and diploid potato clones resulting in enhanced pigment accumulation not only in tuber skin but also in young foliage, flower petals and tuber flesh (Jung et al. 2009). Allelic diversity and interacting proteins of StAN1 were investigated by D’Amelia et al. (2014) leading to confirmation of its impact on foliage pigmentation. Tuber skin colour of cultivar Desiree deepened also after overexpressing of StAN11, a WD40-repeat gene that regulates expression of DFR (Li et al. 2014). Other study has shown that transcription factors StbHLH1 and StJAF13 are essential for regulation of anthocyanins biosynthesis in potato, while StAN1, StMYBA1 and StMYB113 are expressed also in the absence of pigmentation (Liu et al. 2016). However, not only transcription factors were exploited to enhance potato anthocyanin accumulation and intensity of colour. A potato UDP-glucose: flavonoid-3-O-glucosyltransferase (3GT), when overexpressed in cultivar Desiree, also caused increase in anthocyanin content and more intensive colour of the tuber skin (Wei et al. 2012). This enzyme acting downstream of F3′,5′H and DFR, catalyses the transfer of glucosyl moiety to anthocyanidins thus improving their stability and water solubility. A different method to increase anthocyanin content has been based on the overexpressing key enzymes involved in their synthesis also proved to be successful. Single-gene overexpression of genes encoding chalcone synthase (chs) and chalcone isomerase (chi), enzymes that mediate the first and second steps in the flavonoid biosynthetic pathway, resulted in a significant increase of phenolic acids and anthocyanins in potato tubers (Lukaszewicz et al. 2004).
From the above review of the attempts to modify the quantity of anthocyanins in potato plants it is difficult to conclude which genes are most important for anthocyanin accumulation because any overexpression of genes involved in regulation of the process, or directly in the various stages of synthesis, indeed was successful. A genetic approach to tackle this problem has been rarely used. A quantitative trait loci (QTL) analysis of pigmented tuber flesh in potato identified three regions affecting the trait and located on chromosome V, VIII and IX (Zhang et al. 2009b). Location of chi overlapped with QTL on chromosome V, while a bHLH transcription factor similar to petunia an1 (Stan1) co-localised with QTL on chromosome IX. Each of QTL on chromosomes V and IX explained up to 8.1% of variance observed in phenotypes of the mapping population and contribution of Stan1 allele to tuber flesh pigmentation was demonstrated using various other tetraploid potato genotypes (Zhang et al. 2009b). The goal of this study was to explore further the natural diversity of potato regarding the intensity of the anthocyanin pigmentation and to use flower colour as a model to study genomic regions and candidate gene alleles key for accumulation of these pigments in various quantities. QTL analyses were performed in two diploid mapping populations segregating for flower colour intensity and candidate genes were identified on the basis of function and location (chi, chs) or location (RNA-dependent RNA polymerase 1, RDR1). The first reason for including RDR1 as a candidate gene was its location in the peak of QTL in our preliminary analysis. However, the RDR function also might be related to anthocyanin pigmentation, since another RDR in A. thaliana has been associated with regulation of the anthocyanin biosynthesis (Willmann et al. 2011). In case of one population, which was an F1 progeny of a wild parent and cultivar dihaploid, we analysed the sequences of candidate genes fragments descending from both parents. The flower colour treated as a model trait enabled identification of QTL and candidate gene alleles affecting pigmentation intensity. We postulate that the same QTL when present in potato genotypes with dominant alleles in loci D (I) (Salaman 1910; De Jong 1987) or Pf (de Jong 1987) that govern pigmentation of tuber skin and flesh, may cause enhancement of colour also in tuber skin and flesh, respectively.
Materials and methods
Plant material
The studied material consisted of unselected F1 populations of diploid potato derived from two crosses (Table 1). Population 12–3 (N = 159) was a progeny of hybrid clones DG 00-683 (white flowers) and DG 08-28/13 (dark violet flowers) and it has been previously used to map loci controlling tuber starch and leaf sucrose contents (Śliwka et al. 2016). The number of individuals in current study was reduced from the published one because of removal of the ones with missing flower colour data. Alleles in both parents might have originated not only from S. tuberosum but also from other Solanum species: S. chacoense (28 and 11% in the genomes of DG 00-683 and DG 08-28/13, respectively), S. gourlayi, S. microdontum, S. verrucosum and S. yungasense present in their pedigree. There is 9% contribution of S. phureja to the DG 08-28/13 genome. Using population 12–3 and 1597 Diversity Array Technology (DArT) and CAPS markers, a genetic map of both parental genomes has been constructed with total length of 1117 cM (Śliwka et al. 2016).
Population 05–18 (dH Balbina × 99–10/36; N = 114) has served previously for mapping a late blight resistance gene Rpi-rzc1 (Śliwka et al. 2012). The late blight resistant parent, clone 99–10/36 (pale violet flowers), has been selected from the accession VIR 7370 (VIR 8664) from the N.I. Vavilov Institute for Plant Genetic Resources (VIR), Russia, previously identified as Solanum ruiz-ceballosii Cárd. (Zoteyeva et al. 2012). Specimens made from the clone (Śliwka 99–10/36, sheets BM001211939 and BM001211940) were re-identified as Solanum brevicaule Bitter (sensu Spooner et al. 2016) following examination of material at the Natural History Museum (London) herbarium. The other parent (dH Balbina) is a dihaploid of Polish potato cultivar Balbina and has white flowers. A genetic map of population 05–18 is available and consists of 1603 DArT markers and 48 sequence-specific PCR markers with the total map length of 1204.8 cM (Śliwka et al. 2012).
More seeds of the cross dH Balbina × 99–10/36 have been sown to obtain an extended population 12–4 for fine mapping of the Rpi-rzc1 gene. In that work, only locus on potato chromosome X containing the Rpi-rzc1 gene was enriched with sequence-specific markers (Brylińska et al. 2015). In this study, 315 full-sibling individuals with flower colour data available (from populations 05–18 to 12–4) were treated jointly as population 12–4. The candidate gene markers were scored in this population but DArT genotyping data was not available for QTL mapping (Table 1).
Flower colour assessment
Flower colour was evaluated on 1–4 scale, where 1 = white corollas, 2 = pale violet, 3 = violet and 4 = dark violet. It is a relative scale of increasing flower colour intensity, where score 4 is the most intensive violet observed in each mapping population. However, in population 12–3 the darkest violet (4) was lighter than the darkest violet in population 12–4. The assessment was done in vegetation seasons 2012–2014 using field-grown plants (population 12–4) or plants grown in a net tunnel (population 12–3). Assessments of population 05–18 were repeated in 2006, 2007 and 2008 in the field (Śliwka et al. 2012). Flower colour categories 2–4 were merged in one group and flower colour has been previously mapped as a qualitative trait to locus F on potato chromosome X both in populations 05–18 (Śliwka et al. 2012) and 12–3 (Śliwka et al. 2016).
Sequence-specific DNA markers
PCR markers At1g14790 (RDR1 fragment; Śliwka et al. 2012), chi and chs (Zhang et al. 2009b) were amplified as described earlier. The reaction mixture contained in 20 μl: 2 μl of 10 × PCR buffer, the four deoxynucleotides (0.1 mM), MgCl2 (1.5 mM), primers (0.2 μM), Taq polymerase (0.05 U/μl) and 10–30 ng of template DNA. The thermal conditions of PCR reaction were following: 94 °C—180 s; 40 cycles of: 94 °C—30 s, 55 °C—45 s, 72 °C—90 s; 72 °C—420 s. The polymorphism of all three markers was revealed directly as PCR products of different sizes in population 12–3, while in population 12–4 it was revealed after digestion of the chi and chs markers with Tru1I and EcoRI restriction endonucleases, respectively. The chi and chs PCR products obtained for dH Balbina and 99–10/36 were purified with a GenElute PCR Clean-Up Kit (Sigma-Aldrich, St. Louis, MO, USA) according to the manufacturer’s protocol. The PCR products were cloned into blunt pCRScript Amp SK cloning vector (Promega, Madison, Wisconsin, USA). After ligation, E. coli Top10 chemocompetent cells were transformed and spread on lysogeny broth (LB) plates containing 100 µg/ml ampicillin and X-gal (40 µg/ml) for blue-white selection. Bacteria from white colonies were used as a source of plasmid template in colony PCR for positive clones screening. Universal M13F and M13R primers were used to bidirectionally sequence transformants containing cloned insert of interest. Sequencing reactions were done using BigDye Terminator v3.1 kit (Life Technologies Polska Ltd., Warsaw, Poland) and sequencing products for six or seven clones for each of markers were resolved on ABI3730XL genetic analyser. DNA cloning and sequencing were performed by an external company (Laboratory of DNA Sequencing and Oligonucleotide Synthesis, oligo.pl, Institute of Biochemistry and Biophysics Polish Academy of Sciences). Quality check and trimming of the DNA sequences were done with ChromasLite 2 software and the consensus sequences were produced using MultAlin online software (Corpet 1988).
Data analyses and QTL mapping
Two approaches for QTL mapping were applied. In populations 05–18 and 12–3 where DArT linkage maps were available (Table 1), restricted Multi-QTL Mapping (rMQM) was performed in MapQTL® 6 software using F locus (1 = violet flowers, 0 = white flowers) as a covariate/experimental design cofactor (Van Ooijen 2009). QTL with an LOD score > 3 were considered to be significant. Candidate gene markers were incorporated to the existing genetic maps using JoinMap® 4.1 (Van Oijen 2006) with setting described earlier (Śliwka et al. 2012, 2016). rMQM was then repeated using the enriched maps. When the linkage map was not available, i.e. in the extended population 12–4, associations between candidate gene markers and flower colour intensity as well as determination coefficients (R 2) were estimated by the Student’s T test and analysis of variance, respectively, excluding the individuals with white flowers from the analyses. The same was performed in the population 12–3, to enable comparisons. The fit of segregation to the expected ratio was checked by the χ 2 test. All statistical analyses were done using Statistica 10 software (Statsoft Inc. 2011).
Results
In the population 12–4 (population 05–18 included), ratio of individuals with white flowers to the ones with violet flowers was 151: 164 which was a proportion not deviated significantly from the 1:1 (χ 2 = 0.29; p = 0.59). Similarly, in the population 12–3, 88 individuals had white and 71 individuals had violet flowers, (1:1: χ 2 = 1.35; p = 0.25). Such ratios were in agreement with a hypothesis on segregation of a single gene with Ff × ff genotypes of parental forms, where allele F confers anthocyanin pigmentation in petals (Śliwka et al. 2012, 2016). Application of rMQM in populations with available genetic maps (05–18 and 12–3) and using F locus (1 = violet flowers, 0 = white flowers) as a covariate allowed detection of QTL affecting flower colour intensity otherwise masked by the F locus. The strongest QTL was preliminary located on chromosome V in both populations (data not shown), therefore three candidate gene markers were chosen on the basis of function and location (chi, chs) or location (RDR1 fragment: At1g14790). The markers were tested for polymorphism, scored and incorporated to both genetic maps. The three markers were mapped to corresponding positions on chromosome V of both 05–18 and 12–3 maps, as shown in Table 2. rMQM procedure was then repeated using the enriched maps and the results are also shown in Table 2.
In population 05–18, a QTL with the strongest effect on flower colour intensity was found on chromosome V, spanning 11.4–33.6 cM with a 1 cM gap at 21.8–22.8 cM (Fig. 1a). A DArT marker pPt-471439 at the QTL peak (LOD = 4.77; 27.7 cM) explained 6.0% of the variance observed in the trait and was inherited from the S. ruiz-ceballosii parent 99–10/36 with violet flowers. Candidate gene markers chi and chs were located outside the QTL, while marker At1g14790 had a significant effect on the trait (LOD = 4.43; R 2 = 5.6%). Three more QTL for flower colour intensity were detected on genetic map 05–18 on chromosomes VI, VII and XII. From 4.4 to 5.1% of variance in phenotype could be ascribed to these QTL and they covered narrow regions up to 2.7 cM (Table 2).
In population 12–3, three QTL affecting flower colour were detected by rMQM (Table 2). The most important one was located on chromosome V (0–27.1 cM) in a region in majority overlapping with the most significant QTL detected using population 05–18 (Fig. 1b). Within the QTL on chromosome V, a DArT marker pPt-456666 (descending from both parents) and candidate gene marker At1g14790 (descending from the white flowering DG 00-683) had highest effects on flower colour intensity (LOD = 4.71; R 2 = 1.9%). The chi marker, also descending from DG 00-683, was significantly associated with the trait (LOD = 4.71; R 2 = 1.9%) and the chs was not. The remaining two QTL for flower colour intensity were located on chromosomes VI (44.5–45.6 cM) and VIII (19.3–19.5 cM) of the population 12–3 and explained up to 1.7 and 1.6% of the variance, respectively. The position of the QTL on chromosome VI was different from the position of the QTL detected in population 05–18 on the same chromosome (Table 2).
To characterize QTL for flower colour intensity on chromosome V, candidate gene markers were scored in larger population 12–4.The same analyses were performed in the population 12–3. To ignore the effect of locus F, individuals with different flower colours were analysed separately in four categories (Figs. 2, 3) or individuals with white flowers were excluded from analysis (Table 3). Under assumption that if candidate gene markers were linked to the flower colour intensity, the proportion of their alleles among individuals of violet flower categories would be deviated from 1:1, we tested marker segregation data for deviation from that ratio by a χ 2 test (Figs. 2, 3).
Distribution of marker alleles (dark grey: chi.a, At1g14790.a and chs.a, light grey: chi.b, At1g14790.b and chs.b) descending from clone 99–10/36 in the population dH Balbina × 99–10/36. The population (N = 315) divided into four groups according to the flower colour: a white, b pale violet, c violet, d dark violet. Numbers of individuals with particular alleles are shown as data label on the chart. Significant deviation from 1:1 ratio of alleles in each group is calculated by a χ 2 test and marked by asterisks: *− p < 0.05, ***− p < 0.001
Distribution of marker alleles (dark grey: chi.e, At1g14790.e and chs.g, light grey: chi.f, At1g14790.f and chs.h) in the population 12–3. The population (N = 159) is divided into four groups according to the flower colour: a white, b pale violet, c violet, d dark violet. Numbers of individuals with particular alleles are shown as data labels on the chart. Significant deviation from 1:1 ratio of alleles in each group is calculated by a χ 2 test and marked by asterisks: *− p < 0.05, **− p < 0.01, ***− p < 0.001
In population 12–4, the three candidate gene markers segregated 1:1 within individuals with violet flowers of medium intensity (Fig. 2c) and within individuals with white flowers, with the exception of marker chs (Fig. 2a). In the group of individuals with pale violet flowers (Fig. 2b), there was a significant excess of alleles chi.a, At1g14790.a and chs.b, while among the individuals with dark violet flowers alleles chi.b and At1g14790.b were more frequent than expected (Fig. 2d).
In the smaller population 12–3, alleles of candidate gene marker of different origins segregated, therefore named with names different that in population 12–4 (Fig. 3). The alleles chi.f and At1g14790.f were here significantly more frequent among the progeny with pale violet flowers (Fig. 3b) and, unexpectedly also in the group with white flowers (Fig. 3a). All eight individuals with dark violet flowers possessed allele chs.h (Fig. 3d), while among the individuals with medium violet flowers (Fig. 3c) no significant deviations from 1:1 in allele proportions were detected. However, in the population 12–3 the limited number of progeny in particular flower colour categories probably was the reason why the differences although present, were not statistically significant.
The Student’s T test and analysis of variance, performed on the individuals with violet flowers only, indicated that all three candidate gene markers were significantly associated with the violet flower colour intensity in both population 12–4 and 12–3 (Table 3). In the bigger dataset (population 12–4, N = 164) the percentages of variance explained ascribed to the markers were higher than in the smaller one (population 12–3, N = 71). In both populations, the strongest association was between colour intensity and the marker At1g14790: 33.1% of variance explained in population 12–4 and 18.0% in population 12–3 could be ascribed to this marker. The presence of the marker allele At1g14790.b was associated with the 0.9 score increase of the flower colour intensity in the population 12–4, while the presence of the allele At1g14790.e was associated with 0.6 score increase in population 12–3 (Table 3). For the other two markers the percentages of the variance explained ranged from 6% for the marker chs in population 12–3 to 11.3% for the marker chi in the same population. The effects of particular chi and chs alleles on the colour intensity scores were 0.4 or 0.5 (Table 3).
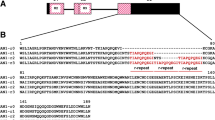
For 99–10/36 parent 7 clones of marker chi and 6 clones of marker chs, while for dH Balbina 6 clones of marker chi and 7 clones of marker chs, were sequenced with the forward and reverse primers. From 99 to 10/36 parent consensus alleles named a and b for both markers were reconstructed using MultAlin software and for dH Balbina c allele of chi as well as c and d alleles of chs were retrieved. Chi fragments differed in length: 806 bp (allele chi.c), 807 bp (allele chi.a) and 811 bp (allele chi.b). Fragments of chs had following sizes: 1222 bp (alleles chs.a, chs.b, chs.c) and 1220 bp (allele chs.d). The GenBank accession numbers of the obtained sequences and their comparison to the sequence of the reference genome DM1-3 v.4.03 are shown in Supplementary Fig. 1. A Tru1I restriction site characteristic for the allele chi.a was located at 535 bp and an EcoRI restriction site characteristic for the allele chs.a was at 502 bp (Supplementary Fig. 1). BLAST query of the chi fragments to reference potato genome DM1-3 v.4.03 matched to an unannotated fragment of chr05:1401031..1401838, while a query against general Nucleotide Collection (NCBI) retrieved fragments of chalcone-flavonone isomerases, e.g. NM_001320711.1 from Solanum lycopersicum in 97% identical to the allele chi.a. Chs fragments matched to the reference potato genome DM1-3 v.4.03 chr05:48887142..48888370 encoding chalcone synthase 2 (PGSC0003DMG400019110). There were two introns in the chi fragments and one in the chs fragment (Supplementary Fig. 1). Comparison of the chi alleles to the reference genome DM1-3 sequence showed one non-synonymous substitution in allele chi.b, two in allele chi.a and four in allele chi.c. The same comparison of chs showed three non-synonymous substitutions in allele chs.c, four in allele chs.d and six in alleles chs.a and chs.b. Alleles chs.a and chs.b differed from each other by two non-synonymous substitutions (Supplementary Fig. 1).
Discussion
We detected three and four QTL affecting the intensity of anthocyanin pigmentation in potato corolla using the mapping populations 05–18 (12–4) and 12–3, respectively. In both populations a locus F, necessary for violet flower colour, segregated and we used three different approaches to differentiate the qualitative effect of this locus and to detect the genetic factors affecting the quantitative flower colour intensity: 1. rMQM and using locus F as a covariate (Fig. 1, Table 2), 2. analysis of markers segregations separately in different flower colour categories (Figs. 2, 3) and 3. T-Student test and ANOVA performed on individuals with violet flowers only. In the first approach, rMQM, genetic maps of whole parental genomes were analyzed and the strongest QTL for flower colour intensity was located on chromosome V. It was the only QTL common for the two mapping populations. To characterize that region better, the candidate gene markers, chosen on the basis of function and location (chi, chs) or location (At1g14790), were used in the second and third approach. Markers chi and chs were taken from the earlier study in which marker chi was mapped within QTL for tuber flesh anthocyanin pigmentation in potato (Zhang et al. 2009b).
Depending on the approach the strength of the QTL on chromosome V and the significance of the three candidate gene markers were estimated differently. According to T-Student test and ANOVA, all three markers (chi, chs and At1g14790) were significantly associated with flower colour intensity in both populations and 18.0 and 33.1% of the variance explained could be ascribed to the marker At1g14790 in population 12–3 and 12–4, respectively. In the rMQM, these effects were weaker (1.9% in population 12–3 and 5.6% in population 05–18), marker chs was not significant for the flower colour intensity in any population and marker chi was only significant in population 12-3. Analyses of markers segregations resulted in an intermediate picture with various combination of significant and insignificant marker-flower colour associations in particular flower colour categories (Figs. 2, 3). In the population 05–18/12–4, number of individuals included in rMQM was smaller (114) than in other analyses (315), which may be a factor affecting the results of analysis. However, in population 12–3 the number of individuals was the same in all analyses that still produced different outcomes demonstrating that the number of individuals was not the only source of differences. One possible explanation of the significant effects of the chi and chs markers is that they are just linked to the gene(s) underlying the trait. However, a second possible explanation would be that the specific combination of chs, chi and RDR1 gene alleles result in particular phenotypes (pale violet, violet and dark violet flower colour) and therefore all three genes contribute to the QTL (as shown in Figs. 2, 3).
The QTL for flower colour intensity located in our study on chromosome V was most likely in the same position as the QTL for anthocyanin tuber flesh coloration described by Zhang et al. (2009b), which is an important finding indicating that the same genetic factors may be involved in anthocyanin accumulation in different parts of the plant. Depending on the presence of loci: F (van Eck et al. 1993, 1994), D (I) (Salaman 1910; De Jong 1987) or Pf (de Jong 1987), the QTL on chromosome V may cause enhancement of violet colour in corolla, tuber skin and flesh, respectively, which may be useful information for breeding potatoes with pigmented tubers. The eggplant chromosomes V and X harbor QTL for anthocyanin distribution in tissues and organs such as fruit skin, stem, leaf, calyx, corolla, venation or fruit peduncle (Barchi et al. 2012; Ge et al. 2013; Toppino et al. 2016). Although direct comparisons with eggplant QTL maps are hampered by the differences in genome organization between this plant and potato (Wu and Tanksley 2010), this may be an indication of conserved character of QTL on chromosome V affecting anthocyanin content in Solanaceae. The eggplant chromosome V, when compared directly to potato chromosome V (Wu and Tanksley 2010) or indirectly via the tomato one (Fukuoka et al. 2012; Rinaldi et al. 2016), has an upper part orthologous to chromosome V (inversed in tomato) and lower part orthologous to chromosome XII. The synteny has been shown between eggplant QTL for anthocyanin pigmentation on chromosome X and the tomato region of chromosome V containing chs (Barchi et al. 2012).
The role of chi and chs diversity in intensity of flower colour in potato has not been elucidated, but we showed that these genes may potentially contribute to the QTL effect. By sequencing the marker fragments from the parents of the mapping population 05–18 (12–4), we showed that they are indeed derived from the genes encoding chalcone isomerase and chalcone synthase. Chi fragments matched to the unannotated fragment of potato genome and the presence of functional chi at chr05:1400665..1403145 should be validated experimentally although it is supported by genetic mapping in our study and by Zhang et al. (2009b). The particular fragmentary alleles of both chi and chs showed polymorphisms between clone 99–10/36 and dH Balbina, that would be manifested also on the protein level and could affect the protein activity. In an earlier study, both chi and chs, when overexpressed, caused a significant increase of phenolic acids and anthocyanins in potato tubers (Lukaszewicz et al. 2004).
The third candidate gene marker, At1g14790 (C2_ At1g14790) originated from Conserved Ortholog Set II (SGN, Solanaceae Genomics Network) and has been designed on the basis of Arabidopsis thaliana RNA-dependent RNA polymerase 1 (AtRDR1, accession NM_101348.4). Two genes encoding RDR1, StRDR1a (PGSC0003DMG400031261) and StRDR1b (PGSC0003DMG400031261), have been identified in the potato reference genome DM 1–3 on chromosome V (Hunter et al. 2016). The location of those genes on reference physical map did not correspond precisely to our genetic map data (Table 2). Hunter et al. (2016) demonstrated that StRDR1 expression is regulated by salicylic acid but failed to show evidence for RDR1 involvement in defence against viruses in potato, although such role of RDR1 has been reported many times in thale cress and tobacco. The authors speculate that RDRs 1 in potato may have lost the antiviral role, perhaps taken over by RDRs 6, and play a different one (Hunter et al. 2016). Plant RDRs are grouped in seven families (Hunter et al. 2016) and their functions include those processes where dsRNA and siRNAs are involved: development, reproduction, biotic and abiotic stress responses. The RDR6 in A. thaliana has been associated with siRNA-producing locus TAS4, which produces ta-siRNAs that target transcripts of MYB transcription factors regulating enzymes in the anthocyanin biosynthesis (Willmann et al. 2011). Therefore, we cannot exclude that the association of the marker At1g14790 with potato flower colour intensity in our study was not only genetic but also functional, indicating StRDR1 involvement in anthocyanin synthesis regulation. This hypothesis can be supported by the fact that anthocyanins accumulation is responsive to salicylic acid treatments as it was shown in sweet cherry (Giménez et al. 2017) and grapes (Champa et al. 2015).
References
Barchi L, Lanteri S, Portis E, Valè G, Volante A, Pulcini L, Ciriaci T, Acciarri N, Barbierato V, Toppino L, Rotino GL (2012) A RAD tag derived marker based eggplant linkage map and the location of QTLs determining anthocyanin pigmentation. PLoS ONE 7(8):e43740
Brylińska M, Tomczyńska I, Jakuczun H, Wasilewicz-Flis I, Witek K, Jones JD, Śliwka J (2015) Fine mapping of the Rpi-rzc1 gene conferring broad-spectrum resistance to potato late blight. Eur J Plant Pathol 143:193–198
Champa WA, Gill MI, Mahajan BV, Arora NK (2015) Preharvest salicylic acid treatments to improve quality and postharvest life of table grapes (Vitis vinifera L.) cv. Flame Seedless. J Food Sci Technol 52(6):3607–3616
Corpet F (1988) Multiple sequence alignment with hierarchical clustering. Nucl Acids Res 16(22):10881–10890
D’Amelia V, Aversano R, Batelli G, Caruso I, Castellano Moreno M, Castro-Sanz AB, Chiaiese P, Fasano C, Palomba F, Carputo D (2014) High AN1 variability and interaction with basic helix-loop-helix co-factors related to anthocyanin biosynthesis in potato leaves. Plant J 80:527–540
De Jong H (1987) Inheritance of pigmented tuber flesh in cultivated diploid potatoes. Am Potato J 64:337–343
De Jong WS, De Jong DM, De Jong H, Kalazich J, Bodis M (2003) An allele of dihydroflavonol 4-reductase associated with the ability to produce red anthocyanin pigments in potato (Solanum tuberosum L.). Theor Appl Genet 107(8):1375–1383
De Jong WS, Eannetta NT, De Jong DM, Bodis M (2004) Candidate gene analysis of anthocyanin pigmentation loci in the Solanaceae. Theor Appl Genet 108(3):423–432
Dodds KS, Long DH (1955) The inheritance of colour in diploid potatoes. I. Types of anthocyanidins and their genetic loci. J Genet 53:136–149
Fukuoka H, Miyatake K, Nunome T, Negoro S, Shirasawa K, Isobe S, Asamizu E, Yamaguchi H, Ohyama A (2012) Development of gene-based markers and construction of an integrated linkage map in eggplant by using Solanum orthologous (SOL) gene sets. Theor Appl Genet 125(1):47–56
Ge HY, Liu Y, Zhang J, Han HQ, Li HZ, Shao WT, Chen HY (2013) Simple sequence repeat-based association analysis of fruit traits in eggplant (Solanum melongena). Genet Mol Res 12(4):5651–5663
Giménez MJ, Serrano M, Valverde JM, Martínez-Romero D, Castillo S, Valero D, Guillén F (2017) Preharvest salicylic acid and acetylsalicylic acid treatments preserve quality and enhance antioxidant systems during postharvest storage of sweet cherry cultivars. J Sci Food Agric 97(4):1220–1228
Hackett CA, Bradshaw JE, Bryan GJ (2014) QTL mapping in autotetraploids using SNP dosage information. Theor Appl Genet 127:1885–1904
Hunter LJ, Brockington SF, Murphy AM, Pate AE, Gruden K, MacFarlane SA, Palukaitis P, Carr JP (2016) RNA-dependent RNA polymerase 1 in potato (Solanum tuberosum) and its relationship to other plant RNA-dependent RNA polymerases. Sci Rep 6:23082
Jung CS, Griffiths HM, De Jong DM, Cheng S, Bodis M, De Jong WS (2005) The potato P locus codes for flavonoid 3′, 5′-hydroxylase. Theor Appl Genet 110:269–275
Jung CS, Griffiths HM, De Jong DM, Cheng S, Bodis M, Kim TS, De Jong WS (2009) The potato developer (D) locus encodes an R2R3 MYB transcription factor that regulates expression of multiple anthocyanins structural genes in tuber skin. Theor Appl Genet 120:45–57
Lewis CE, Walker JRL, Lancaster JE, Sutton KH (1998) Determination of anthocyanins, flavonoids and phenolic acids in potatoes. I. Coloured cultivars of Solanum tuberosum L. J Sci Food Agric 77:45–57
Li W, Wang B, Wang M, Chen M, Yin JM, Kaleri GM, Zhang RJ, Zuo TN, You X, Yang Q (2014) Cloning and characterization of a potato StAN11 gene involved in anthocyanin biosynthesis regulation. J Integr Plant Biol 56(4):364–372
Liu Y, Lin-Wang K, Espley RV, Wang L, Yang H, Yu B, Dare A, Varkonyi-Gasic E, Wang J, Zhang J, Wang D, Allan AC (2016) Functional diversification of the potato R2R3 MYB anthocyanin activators AN1, MYBA1, and MYB113 and their interaction with basic helix-loop-helix cofactors. J Exp Bot 67(8):2159–2176
Lukaszewicz M, Matysiak-Kata I, Skala J, Fecka I, Cisowski W, Szopa J (2004) Antioxidant capacity manipulation in transgenic potato tuber by changes in phenolic compounds content. J Agric Food Chem 52(6):1526–1533
Olsen KM, Hehn A, Jugdé H, Slimestad R, Larbat R, Bourgaud F, Lillo C (2010) Identification and characterisation of CYP75A31, a new flavonoid 3′5′-hydroxylase, isolated from Solanum lycopersicum. BMC Plant Biol 10:21
Passeri V, Koes R, Quattrocchio FM (2016) New challenges for the design of high value plant products: stabilization of anthocyanins in plant vacuoles. Front Plant Sci 7:153
Rinaldi R, Van Deynze A, Portis E, Rotino GL, Toppino L, Hill T, Ashrafi H, Barchi L, Lanteri S (2016) New insights on eggplant/tomato/pepper synteny and identification of eggplant and pepper orthologous QTL. Front Plant Sci 7:1031
Salaman RN (1910) The inheritance of colour and other characters in the potato. J Genet 1:7–46
SGN, Solanaceae Genomics Network. https://solgenomics.net/
Śliwka J, Jakuczun H, Chmielarz M, Hara-Skrzypiec A, Tomczyńska I, Kilian A, Zimnoch-Guzowska E (2012) Late blight resistance gene from Solanum ruiz-ceballosii is located on potato chromosome X and linked to violet flower colour. BMC Genet 13(1):11
Śliwka J, Sołtys-Kalina D, Szajko K, Wasilewicz-Flis I, Strzelczyk-Żyta D, Zimnoch-Guzowska E, Jakuczun H, Marczewski W (2016) Mapping of quantitative trait loci for tuber starch and leaf sucrose contents in diploid potato. Theor Appl Genet 129(1):131–140
Spooner DM, Alvarez N, Peralta IE, Clausen AM (2016) Taxonomy of wild potatoes and their relatives in Southern South America (Solanum sect. Petota and Etuberosum). Syst Bot Monogr 100:240
SPUD Database—Potato Genomics Resource at Michigan University. http://solanaceae.plantbiology.msu.edu/new.shtml
StatSoft Inc (2011) STATISTICA (data analysis software system), version 10. www.statsoft.com
Toppino L, Barchi L, Lo Scalzo R, Palazzolo E, Francese G, Fibiani M, D’Alessandro A, Papa V, Laudicina VA, Sabatino L, Pulcini L, Sala T, Acciarri N, Portis E, Lanteri S, Mennella G, Rotino GL (2016) Mapping Quantitative Trait Loci affecting biochemical and morphological fruit properties in eggplant (Solanum melongena L.). Front. Plant Sci 7:256
Van Eck HJ, Jacobs JME, van Dijk J, Stiekema WJ, Jacobsen E (1993) Identification and mapping of three flower colour loci of potato (S. tuberosum L.) by RFLP analysis. Theor Appl Genet 86:295–300
Van Eck HJ, Jacobs JME, van den Berg PMMM, Stiekema WJ, Jacobsen E (1994) The inheritance of anthocyanin pigmentation in potato (Solanum tuberosum L.) and mapping of tuber skin colour loci using RFLPs. Heredity 73:410–421
Van Ooijen JW (2006) JoinMap® 4, Software for the calculation of the genetic linkage maps in experimental populations. Kyazma B.V., Wageningen, Netherlands
Van Ooijen JW (2009) MapQTL® 6, software for mapping of quantitative trait loci in experimental populations of diploid species. Kyazma B.V, Wageningen
Wei Q, Wang QY, Feng ZH, Wang B, Zhang YF, Yang Q (2012) Increased accumulation of anthocyanins in transgenic potato tubers by overexpressing the 3GT gene. Plant Biotechnol Rep 6:69–75
Willmann MR, Endres MW, Cook RT, Gregory BD (2011) The functions of RNA-dependent RNA polymerases in Arabidopsis. Arabidopsis Book 9:e0146
Wu F, Tanksley S (2010) Chromosomal evolution in the plant family Solanaceae. BMC Genom 11:182
Zhang Y, Cheng S, De Jong D, Griffiths H, Halitschke R, De Jong W (2009a) The potato R locus codes for dihydroflavonol 4-reductase. Theor Appl Genet 119:931–937
Zhang Y, Jung CS, De Jong WS (2009b) Genetic analysis of pigmented tuber flesh in potato. Theor Appl Genet 119:143–150
Zoteyeva N, Chrzanowska M, Flis B, Zimnoch-Guzowska E (2012) Resistance to pathogens of the potato accessions from the collection of N.I. Vavilov Institute of Plant Industry (VIR). Am J Pot Res 89:277–293
Acknowledgements
We thank Professor Herman van Eck (Wageningen University, the Netherlands) for inspiring this work and Dr. Dariusz Mańkowski (Plant Breeding and Acclimatization Institute – National Research Institute, Radzików, Poland) for advice and verification of the statistical analyses used in the article. The research was funded within a statutory dotation 1-3-00-3-06 from the Polish Ministry of Science and Higher Education and the G2P-SOL project (Title: Linking genetic resources, genomes and phenotypes of Solanaceous crops) which has received funding from the European Union’s Horizon 2020 research and innovation programme under grant agreement No. 677379. The re-identification of S. ruiz-ceballosii clone 99-10/36 was done as part of a visit to the Natural History Museum (BM) UK funded by the SYNTHESYS Project http://www.synthesys.info/ which is financed by European Community Research Infrastructure Action under the FP7 Integrating Activities Programme. We thank Dr. Sandra Knapp for hosting a visit of Jadwiga Śliwka in London and the curators of the herbarium BM for logistical support.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Śliwka, J., Brylińska, M., Stefańczyk, E. et al. Quantitative trait loci affecting intensity of violet flower colour in potato. Euphytica 213, 254 (2017). https://doi.org/10.1007/s10681-017-2049-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10681-017-2049-3