Abstract
Especially in urbanized landscapes, habitat fragmentation and increasing numbers of infrastructural features may limit genetic exchange among wildlife populations. Yet, whether this results in genetic differentiation among individuals in different habitat fragments will depend on both the species studied and the composition of the landscape. European pine martens (Martes martes) show clear spatial structure at a Europe-wide scale, but whether gene flow among habitat patches can be maintained at a more local scale in intensively urbanized areas remained unclear. Here, we analysed genotypic data from 270 pine martens sampled from locations scattered across the Netherlands, one of the most densely populated areas in the world. Using Bayesian clustering models we show that most likely maximum two large subpopulations exist in the Netherlands. We observed relatively low levels of genetic differentiation and genetic evidence of regular long-distance dispersal by juveniles that must have crossed one or multiple major highways. Our results suggest that genetic exchange among Dutch pine martens has, until 2010, not been impacted severely by the countries’ dense infrastructural network. Furthermore this species seems to have maintained its genetic diversity despite a recent demographic bottleneck. These conclusions support the idea that the effects of habitat fragmentation may strongly differ between (groups of) species, and that prioritization and optimization of management decisions thus requires direct study of the targeted species.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Forested areas are becoming increasingly fragmented as a result of land exploitation and the development of infrastructural features such as railways and highways. Worldwide, fragmentation and loss of natural habitat forms the most important threat for wild species (Vitousek et al. 1997). Effects on biodiversity may be especially severe in strongly urbanized areas like north-western Europe (Mortelliti et al. 2010). Decreased size of forest patches may reduce habitat quality and maximum local population sizes of forest dwelling animals, which may increase extinction risks due to both demographic and genetic effects (Fahrig 2003). From a genetic point of view, small populations may show reduced adaptability due to lack of variation, as well as reduced fitness due to inbreeding depression (DiBattista 2008). At the same time, larger distances between patches or the presence of dispersal barriers may reduce connectivity and thereby opportunities for gene flow (Frankham et al. 2010). Understanding the relative importance of these effects is of great importance for conservation management, as it helps to decide whether mitigating measures should focus on habitat reconstruction or primarily on the development of dispersal corridors. Yet, responses are variable among groups of species (Blanchet et al. 2009). Many carnivorous mammals are disproportionately affected by habitat destruction, as they typically occur in low densities and thus require large areas to sustain a genetically viable population (Henle et al. 2004). Yet, if connectivity between patches is sufficiently high, individuals in various interconnected fragments may effectively function as a single population. Assessing levels of connectivity thus is essential, but not straightforward in complex urbanized landscapes. While local barriers may constrain genetic exchange between habitats on opposite sides, such barriers may occasionally be passed or circumvented, especially by carnivorous mammals showing high juvenile dispersal over potentially large distances (Gehrt et al. 2010). Depending on the behaviour and demography of the study species, and the composition of the landscape, urbanization may or may not result in a subdivision of a population into multiple genetically isolated groups. Whether this is the case for relatively small mammals, such as mustelids, in one of the most urbanized areas in the world, the Netherlands, remained largely unclear until now. In the Netherlands, strong urbanization and mass motorization have occurred after the second world war. A dense network of motorways was constructed in a relatively short period of time, with a strong peak in the 1960’s to 1980’s and a more gradual intensification afterwards (e.g. Staal 2003).
European pine martens (Martes martes) are an excellent example of a mid-sized mustelid still found to inhabit fragmented habitats in urban landscapes, while its low population densities and reproductive rates may make it particularly vulnerable to loss and change of habitat (Pertoldi et al. 2008; but see Mergey et al. 2012). Pine martens show strong territoriality, resulting in large home ranges and high dispersal of, at least in the Netherlands (Broekhuizen and Müskens 2000), especially the juvenile males. Natal dispersal distances of up to 9 km in females and up to 181 km in males were reported for the related American marten (Martes americana; Johnson et al. 2009), suggesting that occasional dispersal over >100 km may also occur in pine martens.
Due to a combination of habitat loss, persecution and traffic accidents the European pine marten experienced drastic population declines over the last century (e.g. Jordan et al. 2012). Several genetic studies were conducted to quantify effects on genetic variation. Pertoldi et al. (2008) showed a clear decline in genetic variation over time in Danish pine marten populations. Kyle et al. (2003) showed stronger spatial structure among populations of M. martes than among populations of Martes americana (Kyle and Strobeck 2002), which they attributed to a higher antropogenic pressure on the European continent. Widespread fragmentation of the European forests likely resulted in increased isolation and consequential differentiation between different European populations. Yet, as the populations sampled by Kyle et al. (2003) were located hundreds of kilometres apart, it remains unsure what happens at a finer scales in urbanized landscapes.
In this study, we used samples gathered from localities scattered throughout the Netherlands to assess to what extent the urbanization of the Dutch landscape in the 20th century has effected the genetic exchange and maintenance of variation among the pine martens living in its fragmented forests. Due to declines in the 1970’s and 1980’s (Müskens et al. 2000), the number of Dutch pine martens was estimated to be reduced to about 400 in the 1990’s (Thissen et al. 2010). Most pine martens occurred and reproduced in three key forest areas (Veluwe, Utrechtse Heuvelrug and Drents-Friese Wold) and a swamp forest in the north of the country (Wieden-Weerribben). Population sizes started to increase again after 2000 (see Figure S1 of the Electronic Supplementary Material), but the genetic consequences of the past declines remained unknown (Jansman and Broekhuizen 2000). We hypothesized that the demographic bottleneck in the 20th century has reduced genetic variation and increased spatial differentiation between key habitats (Barton and Charlesworth 1984), and that the intensification of traffic networks that occurred in parallel will have hampered the potential of recent increases in population size to restore gene flow. To test these hypotheses, the following questions were addressed: (1) To what extent are pine martens in different habitat patches genetically differentiated and acting as individual populations?; (2) Does the Dutch urbanized landscape impose genetic barriers to pine martens?; and (3) How much genetic variation do(es) the Dutch population(s) comprise?
Materials and methods
Sampling strategy
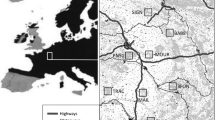
In the 1980’s, Alterra started the collection of detailed data on the occurrence and distribution of pine martens in the Netherlands (e.g. Müskens and Broekhuizen 1986). From 1991 onwards, this survey was intensified in collaboration with the Dutch Pine marten Working Group (e.g. Thissen et al. 2010), and starting from 1992 it included the collection of tissue samples from dead individuals (mainly traffic casualties). Between 1992 and 2010, a total of 251 dead individuals were collected throughout the country (Fig. 1) and delivered to Alterra, where a small sample of tongue tissue was removed for DNA extraction during post-mortem investigation. Samples of 127 individuals were collected from 1992 to 2000, mainly in the four core habitat areas (see Fig. S2 of the Online Electronic Materials). The remaining 124 tissue samples were collected from 2001 to 2010 and included samples from other parts of the country where numbers of pine martens had been increasing, such as the coastal area (see Figure S3). Additionally, 66 faecal samples were collected between April 2009 and May 2010 in the Wieden-Weerribben swamp forest. Samples were directly submerged in 96 % ethanol and stored at −21 °C in the lab.
Spatial distribution of locations from which samples were obtained. Dark grey dots indicate dead individuals (tissue samples), light grey triangles indicate faecal samples. Forested areas are indicated in green. The main forest habitats inhabited by pine martens are encircled on the map (DFW Drents-Friese Wold; UH Utrechtse Heuvelrug; V Veluwe; WW Wieden-Weerribben). (Color figure online)
Laboratory procedures
The DNeasy Blood and Tissue kit and the QIAamp DNA Stool Mini Kit (both QIAGEN) were used to extract DNA from tissue and faecal samples, respectively. Amplification was performed for nine microsatellite loci: GG454 (Walker et al. 2001), MA1, MA2 and MA4 (Davis and Strobeck 1998), MEL1 and MEL6 (Bijlsma et al. 2000), MEL10 (Domingo-Roura 2002), MVI20 and MVI57 (O’Connell et al. 1996). For DNA extracts from faeces, a tenth locus (DBY7Ggu, located on the Y-chromosome) was amplified for sex determination. After prior tests during a pilot study, optimized PCR reactions were performed in volumes of 10 µl, including 0.3 Units of Taq (Invitrogen Taq DNA polymerase), amounts of PCR buffer and W-1 according to the Invitrogen protocol, 100 nM of both primers, 200 µM of each dNTP, 3 mM MgCl2 and 320 µg/ml BSA. 2 µl of DNA-template was used per reaction, containing between 4 and 40 ng DNA in case of extracts from tissues. The following PCR programme was used for tissue samples: 94 °C for 2 min, 30 cycles at 94 °C for 30 s, 56–64 °C for 30 s, 72 °C for 30 s and a final extension step of 1 min at 72 °C. For faecal samples, a different programme was used: 94 °C for 2 min, 36 cycles at 94 °C for 30 s, 56–64 °C for 30 s, 72 °C for 1 min and a final extension step of 20 min at 72 °C. Forward primers were labelled with either IRD-700 or IRD-800 for analysis on a Li-Cor 4300 platform. Samples were first amplified for the locus MEL10, which can be used to distinguish pine martens from stone martens (Martes fiona; Pilot et al. 2007), and samples that did not amplify or that were identified as stone marten were discarded. Genetic profiles of tissue samples were based on a single PCR-reaction per locus. Genotypes with missing data for >1 locus were considered unreliable and were discarded from the data analysis. For faecal samples, a multi-tube approach with three independent PCR reactions per locus per sample was used to minimize genotyping errors (allelic dropout and false alleles). In accordance with the approach described in Koelewijn et al. (2010), we first amplified a single locus (MEL10) for all samples. The other loci were only amplified for samples that showed three times the same genotype for MEL10. Any samples that did not show the same genotype at all loci were considered unreliable and were discarded from the data analysis.
Tests for genotypic linkage disequilibrium in the total dataset were performed for all possible pairs of loci using a log-likelihood ratio G-test and Bonferroni correction in FStat (v.2.9.3; Goudet 1995). The same software was applied to test for deviations from Hardy–Weinberg equilibrium (HWE) per locus among samples. We used MicroChecker v.2.2.3 (Van Oosterhout et al. 2004) to check for null alleles, genotyping errors and allelic dropout per locus.
Inference of population structure
Although the distribution of pine martens in the Netherlands centres around a few core habitats, sampling localities were scattered all over the country (Fig. 1). No detailed data were available on the occurrence of reproduction outside the core habitats. As one of the primary goals of this study was to assess the presence of geographical barriers separating different mating groups, we chose not to subdivide the dataset into a priori defined populations. Rather, we applied individual-based Bayesian clustering methods to define the number of populations and to assign each individual to a population, and afterwards assessed levels of genetic variation within and gene flow between the inferred populations (Ball et al. 2010). Two complementary Bayesian clustering programs were applied. We first used STRUCTURE (v.2.3.3; Pritchard et al. 2000), a program that estimates the probability Pr (X|K) of the data to conform to a predefined number populations (K) and estimates per individual the probability q to belong to each of the clusters. We ran STRUCTURE for K = 1 to K = 10, under the admixture model. Although using a model with correlated allele frequencies would be in line with the hypothesized subdivision due to habitat fragmentation, we could not exclude the possibility that a substructure was already present before this fragmentation took place, and therefore repeated the analysis assuming uncorrelated allele frequencies. Runs were replicated five times for each K value, using 500.000 MCMC iterations and a 50.000 iterations burn-in period. We then determined ∆K (Evanno et al. 2005) using STRUCTURE HARVESTER (Earl and vonHoldt 2011) to infer the optimal number of clusters. Secondly, we applied GENELAND (v.4.0.0; Guillot et al. 2008), again testing both the correlated and uncorrelated allele frequency models, while incorporating spatial data. 10 replicate runs were performed (settings: 500.000 iterations per run, burn-in period of 50.000, thinning rate of 100, maximum number of nuclei of 300, and assuming individual admixture and filtering for null alleles). K was defined based on the result of run with the highest average log posterior density (ALPP; Guillot et al. 2012), and results for the five best runs with equal K-value were selected for further analysis.
For each model, we tested the repeatability of the cluster assignments and matched the results of the five replicate runs via CLUMPP (Jakobsson and Rosenberg 2007). Per individual, the modal cluster assigment was based on the population with the highest q-value in the analysis.
The inference of the number of clusters via the ∆K method may only detect the uppermost level of hierarchical structure (Evanno et al. 2005). We therefore applied a hierarchical framework, in which the dataset was split along the K detected clusters, and then the same Bayesian clustering analyses were repeated on each of the clusters to test for any further substructure (Evanno et al. 2005).
Genetic differentiation between the core habitat areas was quantified as G ST (Nei 1987). SMOGD (v.1.2.5; Crawford 2010) was used to calculate unbiased estimates of H S and H T (Nei 1973) per locus. Values for both metrics were averaged across loci before calculating G ST .
Principal component analysis (PCA), based on binary presence data per allele per locus, was applied in order to assess the level of variation in allelic composition among individuals in the four core habitat areas, as well as within and between the inferred clusters. The added value of this analysis is that it answers a different question than the Bayesian algorithms, as it groups individuals based on similarity in allelic variants rather than based on HWE (De Groot et al. 2012). Therefore, it allows assessment of the extent to which the clusters inferred via Bayesian algorithms contain different combinations of alleles instead of simply differing in allele frequencies. The analysis was conducted in PCord (v.6.0; McCune and Mefford 1999). Geographic X and Y coordinates were incorporated as quantitative variables in a secondary matrix to check for the effect of spatial distribution on genetic composition.
Influence of potential confounding factors
The presence of full siblings among the genotyped individuals was inferred via COLONY (Wang and Santure 2009), assuming male polygamy and female monogamy, a diploid dioecious species, inbreeding, and unknown population allele frequencies. We only accepted full-sib dyads for which the maximum likelihood analysis reported a probability of 1. Identified sibs were then removed prior to the Bayesian clustering analyses, as the presence of close relatives may result in ambiguous clustering output (Rodríguez-Ranilo and Wang 2012).
Furthermore, we are aware that sampling was performed over a time interval of 18 years (1992–2010), a period during which we know that the urbanization of the Dutch landscape has to some extent continued, and distribution patterns of pine martens may have changed (Thissen et al. 2010). To check for potential differences in the distribution of genetic variation over time, we split the dataset into subsets of samples collected before and after 1 January 2001. For both time periods, the suite of Bayesian clustering analyses described above was then repeated for both subsets to check whether the same clusters were obtained. We checked for a temporal change in spatial differentiation by calculating G ST between the main geographical regions identified in the clustering analyses, based on the total dataset as well as for the subsets of samples collected before and after 1 January 2001.
Likewise, we sampled mainly from traffic casualties, thereby potentially oversampling dispersing individuals. Since male-biased dispersal has been observed in the Netherlands, we reanalysed the dataset while including either only females (36 % of the individuals) or only adults (52 % of the individuals). Both sets were again analysed using all four Bayesian clustering models.
Analyses of genetic variation
To check for a recent genetic bottleneck as a result of reduced population sizes in the 20th century, we tested for deviations from a normal L-shaped allele frequency distribution using the software package BOTTLENECK, assuming an I.A.M. mutation model (Cornuet and Luikart 1996).
Various measures of genetic diversity were calculated for the total set of samples, as well as for the inferred clusters. The mean number of alleles per locus (A) and the mean number of private alleles per locus (A p ) were calculated by hand. Allelic richness (A- r ) was calculated by means of rarefaction in Fstat. Observed heterozygosity (Ho) was calculated in Genepop (Raymond and Rousset 1995). Expected heterozygosity (H e ; Nei 1987), and the fixation index F is were calculated using Fstat. Randomization tests were applied to test for significant deviation from HWE within clusters. We assessed levels of isolation by distance (IBD), by performing an individual-based analysis in Genepop. We tested for a difference in Ar between the first and second decade of sampling via a paired-sample t test in SPSS (v.22; IBM Corp.), using markers are replicates.
Results
Marker performance and total variation
Reliable genetic fingerprints were obtained from 24 of the 66 faecal samples (39.4 %). PI(SIB) was 0.0035, indicating only 0.35 % chance even for full siblings to have identical genotypes at the nine loci applied. Therefore, in case multiple scats from the same area showed the same multilocus genotype, these were interpreted as originating from the same individual. In this way, 19 individuals were detected based on faecal samples. This resulted in a total dataset of 270 individuals, of which 170 were males and 95 were females. For 5 individuals the sex could not be determined. Locus GG454 showed a significant non-random association with two other loci (MA2 and MVI57; P < 0.001). A significant homozygote excess was detected for locus MVI20 (Fis = 0.692; P < 0.001), likely due to the presence of null alleles. Both loci were discarded from further analysis.
A characterization of the genetic variation within the total set of Dutch pine martens is presented in Table 1. On average, 5.4 alleles were observed per locus (values ranging from 2–7; see Table S1 of the Electronic Supplementary Material). Mean observed heterozygosity (0.629) was slightly lower than the mean expected heterozygosity (0.662), resulting in a slightly positive FIS, but variation among loci was high (Table S1) and no significant deficit or excess of heterozygotes was detected. BOTTLENECK results showed no deviation from the L-shaped allele frequency distribution expected under a mutation-drift equilibrium.
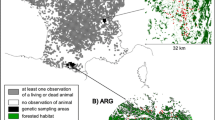
Using COLONY, based on the dataset with seven loci, 16 sets of individuals (all juveniles or subadults) were identified as full siblings (Table 2). Most sets concerned pairs, and set members were sampled 0–4 years apart. In 10 cases siblings were found >25 km apart, with a maximum of 87.6 km (Table 2). As shown in Fig. 2 and Table 2, in nine occasions the sampling locations of these siblings were separated by one or more major highways, indicating successful dispersal across these potential barriers by at least one of them.
Geographic location of the 16 sets of full siblings detected via COLONY. Sibs share the same set number. Males are depicted by triangles, females are depicted by circles. Fat yellow lines are highways, thin black lines are railways. Forests are indicated in green, urban areas are indicated in orange. (Color figure online)
Spatial genetic structure
Principle component analysis (PCA) based on genotypes of the individuals inhabiting the four core habitat areas indicated strong overlap between the two northern habitats (Drents-Friese Wold and Wieden-Weerribben) as well as between the two central habitats (Utrechtse Heuvelrug and Veluwe), but more limited overlap between the northern and central habitats (Fig. 3a). This was supported by G ST values for genetic differentiation, which were <0.03 between the northern and between the central habitats, but were >0.06 between habitats in the north and habitats in the south (Table 3(a)).
Results of a principal components analysis (PCA) based on presence data per allele per locus, for individuals inhabiting the four core habitat areas (a 19.7 % explained variation along the first two axes) and for all individuals in the total dataset (b 18.6 % explained variation along the first two axes). The red vectors in b depict the strength of the correlation between the axes of variation and geographical position with respect to X and Y coordinates. (Color figure online)
A PCA based on the total dataset (Fig. 3b) showed that the geographical position of the individuals along the X and Y coordinates explained respectively 13.2 and 34.9 % of the genetic variation along the first two axes. Furthermore, considerable and significant correlation was detected between pairwise geographic and genetic distances among individuals based on a Mantel test (R = 0.134; P < 0.001), indicating an isolation by distance effect at a country-wide scale.
The ∆K value calculated based on the output of the correlated allele frequencies models in STRUCTURE showed a clear optimum at K = 2 (see Fig. 4). At K = 2, a distinction between a northern (“North”) and central (“Central”) cluster could be observed when plotting modal cluster assignments on a map (Fig. 5a). Some overlap was present and many individuals showed an admixed cluster membership (Fig. 6a). Under the uncorrelated model in STRUCTURE, highly similar patterns were found and model cluster assignments were the same for all but three individuals. To avoid redundancy in the graphs, only the results of the correlated model are presented in Figs. 5 and 6.
Maps showing the spatial distribution of cluster assignments (a and c), and results of principal components analyses (PCA; b and d). Colours represent modal cluster assignments per individual. a and b show the results as inferred using STRUCTURE (correlated allele frequencies model); c and d show the results as inferred using the uncorrelated frequency model in GENELAND. Forested areas are indicated in green on the maps. PCA plots show the position of each individual along the first two axes of variation, which together explained 18.6 % of the total variation in the dataset. (Color figure online)
Plots of cluster membership per individual, calculated via CLUMPP based on the five best replicate runs of the correlated allele frequencies model in STRUCTURE (a), and the uncorrelated (b) and correlated (c) allele frequencies model in GENELAND. Each individual is represented by a small vertical bar, divided in one or more coloured segments. Colours represent different clusters, segment lengths indicate relative cluster memberships. In a and b, blue color represents the “North” cluster and red represents the “Central” cluster. The correlated and uncorrelated models in STRUCTURE showed highly similar results. Under the correlated model in GENELAND, the optimal number of clusters K was seven, yet each individual was assigned in more or less equal proportions to all seven clusters, thus not allowing any spatial interpretation (see main text for more information). (Color figure online)
Under the uncorrelated frequencies model in GENELAND, all runs consistently again detected two clusters (ALPP ranging from −9487.8 to −9400.9). However, 22 % of the individuals were assigned differently compared to STRUCTURE, resulting in a much clearer geographic distinction between the “North” and “Central” cluster (Fig. 5c). Only a small number of admixed individuals was detected (Fig. 6b). No interpretable spatial structure was present under the correlated frequencies model in GENELAND. The various model runs consistently suggested the presence of 7 clusters, yet individual assignments strongly varied among runs. As a result, the integrated cluster assignments as calculated via CLUMPP resulted in a situation where all individuals were assigned with almost equal probabilities to all 7 clusters (Fig. 6c), thus not allowing any valid spatial interpretation.
A second round of analyses was applied to each of the two clusters initially inferred under the STRUCTURE and uncorrelated GENELAND models. Using GENELAND, this resulted in K = 1 for both groups, thus suggesting no further substructure. Using STRUCTURE, this resulted in K = 1 for the “North” cluster, but K = 4 for the “Central” cluster. These subclusters were, however, fully spatially intermixed and did not have any apparent biological meaning. We therefore accepted K = 2 as the most meaningful level of spatial structure for our dataset.
Genetic differentiation between pine martens in the North and Central area was very limited (G ST = 0.034, when based on the total dataset; Table 3(b)).
Genetic variation within and between the inferred populations
Table 1 shows the level of genetic diversity expressed as either the allelic diversity (A and A r ) or the expected heterozygosity (H e ), for the total dataset as well as for each of the two clusters, as inferred via either STRUCTURE or GENELAND. The almost complete similarity in individual assignments under the correlated and uncorrelated STRUCTURE models resulted in equal values for the various parameters. The output of the correlated GENELAND model was ignored, as individuals could not be assigned to clusters.
Both in terms of allelic diversity (A and A r ) and in terms of expected heterozygosity, little difference was observed between the “North” and “Central” clusters (Table 1). Both contained most of the variation present in the total dataset. In STRUCTURE, the “North” cluster contained somewhat less individuals, which resulted in a bit less variation for this cluster, and a higher number of private alleles (A p ) for the “Central” cluster. This was also reflected by the PCA results, which showed less overlap in allelic composition between clusters based on the STRUCTURE output (Fig. 5b and d). In agreement with model assumptions, no significant deviation from HWE was detected. FIS values were close to zero for both clusters under both models, but especially in STRUCTURE.
Changes in genetic variation and spatial structure during the sampling period
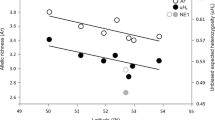
When analysing two subsets of the data for respectively the sampling period before and after 1-1-2001, allelic richness did not show a significant difference (Ar = 4.8 before and 5.2 after 1-1-2001; t = −2.218; P = 0.063). Bayesian clustering using the uncorrelated and correlated frequency models in STRUCTURE and the uncorrelated model in GENELAND in most cases supported a division into two clusters, largely resembling the “North” and “Central” clusters found for the total dataset (see Table 4 for optimal K-values, and Figure S2 and S3 of the Online Electronic Supplements for maps of cluster assignments per individual). For the dataset consisting of only samples taken before 2001, GENELAND assigned the southernmost samples of the “Central” cluster to a separate group. The correlated STRUCTURE model showed a third cluster, which was however spatially intermixed with the “North” and “Central” cluster.
Genetic differentiation (G ST ) between the northern and central core areas seemed slightly larger for the individuals sampled before 1-1-2001 than for the individuals sampled after this date (Table 3(b)), but this difference was not significant (t = −1.221; P = 0.262).
Potential confounding factors
In case subsets of the data representing either only the female individuals or only the adult individuals were used as input for the Bayesian clustering models, GENELAND consistently suggested a division into again a “North” and “Central” cluster. As for the subsets for different sampling periods, STRUCTURE models sometimes suggested a third group that was spatially intermixed with the “North” and “Central” cluster. Optimal K-values per model per subset are given in Table 4; maps of spatial distribution of the model cluster assignments are provided in Figures S4 and S5 of the Electronic Supplementary Material.
Discussion
Population structure as inferred by Bayesian clustering models
Our results consistently suggested that the Dutch pine martens can be subdivided into a northern and a central subpopulation. This may reflect a relatively limited dispersal between the central and northern core habitats. These two habitats are separated by the river IJssel, and a relatively open landscape on both sides of this river, which may have prevented regular dispersal across this area. Yet, based on our sibling analysis we have indications for a least two such events (sibling groups 8 and 9 in Fig. 2). Moreover, drawing conclusions about the exact location of the population boundary is dangerous, given the extent of the spatial overlap in the two clusters shown in the Structure output, but especially given the clear IBD pattern that was present along a north–south axis. Strong IBD has been shown to result in overestimation of the number of clusters by Bayesian techniques in some conditions (Frantz et al. 2009). We do however consider a subdivision in two subpopulations to be likely, as a difference in genetic composition between them was also detected via PCA. The presence of a few private alleles suggest that, besides limited interdispersal, the observed subdivision may also reflect a different origin. Exchange between the central Dutch population and forest patches in the south of the Netherlands and Belgium may exist (Koelewijn, unpublished data), but remains uncertain without the future inclusion of more samples from these areas, where the species is relatively rare (Van den Berge et al. 2000). Populations of moderate size seem present in North-Western Germany and individuals from these populations may have reached the north and east of the Netherlands.
Limited evidence for habitat isolation and dispersal barriers
Our observation that the Dutch population of pine martens did, after at least half a century of heavy urbanization in the Dutch landscape, not split up into more than two large subgroups is surprising. Given the relatively recent origin of some of the landscape barriers, it is possible that during our sampling period any genetic differentiation caused by these barriers was still too weak to be picked up by the applied clustering methods (Murphy et al. 2008). A stronger signal may build up over time, potentially subdividing the observed clusters into multiple groups in the future. However, most of the major highways intersecting or separating the core habitats for pine martens in the Netherlands were already constructed between 1955 and 1975. Given an average lifespan of wild pine martens of 3–8 years, this suggests that between two and ten generations will have passed between the construction of these roads and the onset of our sampling. Various studies have shown that a few generations may be enough to be able to observe negative effects on genetic differentiation and diversity (see e.g. Holderegger and Di Giulio 2010), although it must me noted that the number of microsatellite loci applied in our study (seven) was limited compared to some other studies which potentially may have caused us to overlook a weak signal of genetic structure (e.g. Lowe et al. 2004). In fact, we observed a putative (not significant) upward trend over time in genetic differentiation between pine martens in the northern and central core habitats. Although the observed level of differentiation was still low for the second decade of sampling (G ST = 0.032; roughly 30 years after the construction of the main highways separating the northern and central habitats), this observation may be an indication that gene flow is to some extent affected and that a stronger spatial genetic differentiation may arise in the future. We recommend periodic monitoring of the genetic structure of the Dutch pine martens over the next decades to check for such an effect.
The observed lack of stronger differentiation until the end of our sampling period could be explained from the contemporary re-expansion of the population (Thissen et al. 2010). Pine marten abundance has been increasing in all four core habitats, and dispersal into the surrounding more open habitats has been observed (see also the historic and current distribution patterns in Figure S1). Furthermore, pairwise estimates of differentiation were low even between the two subpopulations detected in our dataset, and both subpopulations harboured a large part of the total genetic variation present in the Netherlands. Private alleles occurred, but were rare. We therefore conclude that remaining connectivity between the Dutch pine marten habitats is the most likely explanation for the observed spatial structure.
Pine martens have been shown to often be tree-dependent rather than forest-dependent, and may disperse along, or occasionally even occupy, the tiny forest fragments and bushes still remaining throughout the Dutch landscape (e.g. Mergey et al. 2012). Evidence for successful dispersal over long-distances and/or across potential barriers is provided by the genetic identification of nine sets of full siblings that were sampled at locations lying >25 km apart and/or were separated by up to three major highways (and railways; see Fig. 2). This included individuals sampled at regular time intervals from 1995 (set 14; Table 2) up to 2008 (set 9; Table 2), indicating that occasional long-distance dispersal seems to have been possible throughout the sampling period of 20 years. Moreover, it included arterial motorways like the A1/E231 and A12/E35, which run from the German border to respectively Amsterdam and The Hague. Both highways intersect the core forest habitats Utrechtse Heuvelrug and Veluwe (Fig. 2) and frequently cause traffic casualties among pine martens. Additional evidence for long-distance dispersal comes from the observation of individuals assigned to the northern cluster in the coastal area of Noord-Holland, suggesting that pine martens from the northern habitats may have been able to walk along one of the two dikes that cross the IJsselmeer lake in the middle of the country.
Our results do not show any indications that open agricultural areas or large bodies of water have been functioning as barriers for gene flow. Likewise, the expected genetic divergence between the Utrechtse Heuvelrug and the Veluwe (Broekhuizen and Müskens 2000), which are separated by a busy highway (A30) surrounded by open terrain, was not (yet) supported by our data. These results are in contrast with those of various other studies on genetic structure in pine martens (e.g. Ruiz-González et al. 2014) and studies on different animal species, which reported a genetic divergence between populations on opposite sides of highways of similar age (e.g. Coulon et al. 2006; Lesbarrères et al. 2006). Clearly, effects of landscape elements on landscape genetic patterns will vary from site to site, as well as among species.
Levels of genetic variation
Field records have reported a strong decline in population size for the Dutch pine martens in the 1970’s and 1980’s (Müskens et al. 2000). Yet, our data did not show the heterozygote excess or abnormal allele frequency distribution expected in case this population reduction would have resulted in a fast and recent loss of genetic diversity in the population (i.e. a genetic bottleneck). This suggests that the population was able to maintain most of its genetic diversity even during this period. Furthermore, no loss of diversity was observed between the first and second decade of sampling. This, together with the observed evidence for regular long-distance dispersal, including migration between the two subpopulations, suggests that the total population size of >400 pine martens in the Netherlands may well be sufficient to maintain a high diversity in each of the habitat patches inhabited (e.g. Frankham et al. 2010). It is important to consider, however, that even if genetic exchange was effectively maintained until now, these roads clearly cause large numbers of traffic casualties each year, and thereby do function as a barrier for dispersal. As such, it may influence spatial demographic patterns, for instance by hampering recolonization of locally extinct habitat patches. Such potential effects may influence gene flow and (local) diversity in the future. Close monitoring of both demographic and genetic characteristics of the Dutch pine martens will remain important in years to come.
Conclusions
In marked contrast to our expectations, our results show that past population reductions have not seriously impacted levels of genetic variation among Dutch pine martens and that potential barriers for dispersal seem not to impact genetic exchange to such an extent that the Dutch population is split up into many isolated subgroups. Given this limited isolation, we expect the Dutch pine martens to be able to maintain most of their current diversity, even in this densely populated and highly urbanized landscape. This surprising conclusion gives hope for the survival of pine martens and related mustelids in our human-dominated world, although ongoing urbanization may already have started to impact gene flow at a yet still undetectable level and may impact genetic diversity in the future. Besides, our results reinforce the statement of Blanchet et al. (2009) that the effects of habitat fragmentation may strongly differ between (groups of) species, and should therefore be assessed separately for organisms with different life histories. While any potential negative impact of infrastructural works on wildlife connectivity is best avoided, detailed and landscape-scale information on multiple species would strongly aid managers to prioritize and optimize measures to mitigate the effects of fragmentation.
References
Ball MC, Finnegan L, Manseau M, Wilson P (2010) Integrating multiple analytical approaches to spatially delineate and characterize genetic population structure: an application to boreal caribou (Rangifer tarandus caribou) in central Canada. Conserv Genet 11:2131–2143
Barton NH, Charlesworth B (1984) Genetic revolutions, founder effects, and speciation. Annu Rev Ecol Evol Syst 15:133–164
Bijlsma R, Van De Vliet M, Pertoldi C, Van Apeldoorn RC, Van De Zande L (2000) Microsatellite primers from the Eurasian badger, Meles meles. Mol Ecol 9:2215–2216
Blanchet S, Rey O, Etienne R, Lek S, Loot G (2009) Species-specific responses to landscape fragmentation: implications for management strategies. Evol Appl 3:291–304
Broekhuizen S, Müskens GJDM (2000) Geslachtsafhankelijke dispersie bij boommarters (Martes martes) in Midden- en Noord-Nederland. Lutra 43:109–117
Cornuet JM, Luikart G (1996) Description and power analysis of two tests for detecting recent population bottlenecks from allele frequency data. Genetics 144:2001–2014
Coulon A, Guillot G, Cosson JF, Angibault MA, Aulagnier S, Cargnelutti B, Galan M, Hewison JM (2006) Genetic structure is influenced by landscape features: empirical evidence from a roe deer population. Mol Ecol 15:1669–1679
Crawford NG (2010) SMOGD: software for the measurement of genetic diversity. Mol Ecol Resour 10:556–557
Davis CS, Strobeck C (1998) Isolation, variability, and cross-species amplification of polymorphic microsatellite loci in the family Mustelidae. Mol Ecol 7:1776–1778
De Groot GA, During HJ, Ansell SW, Schneider H, Bremer P, Wubs ERJ, Maas JW, Korpelainen H, Erkens RHJ (2012) Diverse spore rains and limited exchange shaped fern genetic diversity in a recently created habitat colonized by long-distance dispersal. Ann Bot 109:965–978
DiBattista JD (2008) Patterns of genetic variation in anthropogenically impacted populations. Conserv Genet 9:141–156
Domingo-Roura X (2002) Genetic distinction of marten species by fixation of a microsatellite region. J Mamm 83:907–912
Earl DA, VonHoldt BM (2011) STRUCTURE HARVESTER: a website and program for visualizing STRUCTURE output and implementing the Evanno method. Conserv Genet Resour. doi:10.1007/s12686-011-9548-7
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol Ecol 14:2611–2620
Fahrig L (2003) Effects of habitat fragmentation on biodiversity. Annu Rev Ecol Evol Syst 34:487–515
Frankham R, Ballou JD, Briscoe DA (2010) Introduction to conservation genetics, 2nd edn. Cambridge University Press, Cambridge
Frantz AC, Cellina S, Krier A, Schley L, Burke T (2009) Using spatial Bayesian methods to determine the genetic structure of a continuously distributed population: clusters or isolation by distance? J Appl Ecol 46:493–505
Gehrt SD, Riley SPD, Cypher BL (2010) Urban carnivores—ecology, conflict and conservation. John Hopkins University Press, Baltimore
Goudet J (1995) FSTAT (vers. 1.2): a computer program to calculate F-statistics. J Hered 86:485–486
Guillot G, Santos F, Estoup A (2008) Geneland: a program for landscape genetics. Mol Ecol Notes 5:1406–1407
Guillot G, Renaud S, Ledevin R, Michaux J, Claude J (2012) A unifying model for the analysis of phenotypic, genetic and geographic data. Syst Biol 61:897–911
Henle K, Davies KF, Kleyer M, Margules C, Settele J (2004) Predictors of species sensitivity to fragmentation. Biodivers Conserv 13:207–251
Holderegger R, Di Giulio M (2010) The genetic effects of roads: a review of empirical evidence. Basic Appl Ecol 11:522–531
Jakobsson M, Rosenberg NA (2007) CLUMPP: a cluster matching and permutation program for dealing with label switching and multimodality in analysis of population structure. Bioinformatics 23:1801–1806
Jansman HAH, Broekhuizen S (2000) Vormen de lokale Nederlandse populaties van de boommarter (Martes martes) een metapopulatie? Lutra 43:101–108
Johnson CA, Fryxell JM, Thompson ID, Baker JA (2009) Mortality risk increases with natal dispersal distance in American martens. Proc R Soc B 276:3361–3367
Jordan NR, Messenger J, Turner P, Croose E, Birks J, O’Reilly C (2012) Molecular comparison of historical an contemporary pine marten (Martes martes) populations in the British Isles: evidence of different origins and fates, and implications for conservation management. Conserv Genet 13:1195–1212
Koelewijn HP, Pérez-Haro M, Jansman HAH, Boerwinkel MC, Bovenschen J, Lammertsma DR, Niewold FJJ, Kuiters AT (2010) The reintroduction of the Eurasian otter (Lutra lutra) into the Netherlands: hidden life revealed by noninvasive genetic monitoring. Conserv Genet 11:601–614
Kyle CJ, Strobeck C (2002) Genetic homogeneity of Canadian mainland marten populations underscores the distinctiveness of Newfoundland pine martens (Martes americana atrata). Can J Zool 81:57–66
Kyle CJ, Davison A, Strobeck C (2003) Genetic structure of European pine martens (Martes martes), and evidence for introgression with M. americana in England. Conserv Genet 4:179–188
Lesbarrères D, Primmer CR, Lodé T, Merilä J (2006) The effects of 20 years of highway presence on the genetic structure of Rana dalmatina populations. Ecoscience 13:531–538
Lowe A, Harris S, Ashton P (2004) Ecological genetics. Blackwell, Malden
McCune B, Mefford MJ (1999) PC-ORD. Multivariate analysis of ecological data. Version 5.0. MjM Software, Oregon
Mergey M, Larroque J, Ruette S, Vandel J-M, Helder R, Queney G, Devillard S (2012) Linking habitat characteristics with genetic diversity of the European pine marten (Martes martes) in France. Eur J Wildl Res 58:909–922
Mortelliti A, Amori G, Capizzi D, Rondinini C, Boitani L (2010) Experimental design and taxonomic scope of fragmentation studies on European mammals: current status and future priorities. Mamm Rev 40:125–154
Murphy MA, Evans JS, Cushman SA, Storfer A (2008) Representing genetic variation as continuous surfaces: an approach for identifying spatial dependency in landscape genetic studies. Ecography 31:685–697
Müskens GJDM, Broekhuizen S (1986) De verspreiding van de boommarter Martes martes (L., 1758) in Nederland. Lutra 29:81–98
Müskens GJDM, Broekhuizen S, Wijsman HJW (2000) De verspreiding van de boommarter Martes martes in Nederland, in het bijzonder in de periode 1989–1999. Lutra 43:81–91
Nei M (1973) Analysis of gene diversity in subdivided populations. PNAS 70:3321–3323
Nei M (1987) Molecular evolutionary genetics. Columbia University Press, New York
O’Connell M, Wright JM, Farid A (1996) Development of PCR primers for nine polymorphic American mink Mustela vison microsatellite loci. Mol Ecol 5:311–312
Pertoldi C, Barker SF, Madsen AB, Jørgensen H, Randi E, Muñoz J, Baagoe HJ, Loeschcke V (2008) Spatio-temporal population genetics of the Danish pine marten (Martes martes). Biol J Linnean Soc 93:457–464
Pilot M, Gralak B, Goszczyński J, Posłuszny M (2007) A method of genetic identification of pine marten (Martes martes) and stone marten (Martes foina) and its application to faecal samples. J Zool 271:140–147
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Raymond M, Rousset F (1995) GENEPOP (version 1.2): population genetics software for exact tests and ecumenicism. J Hered 86:248–249
Rodríguez-Ranilo ST, Wang J (2012) The effect of close relatives on unsupervised Bayesian clustering algorithms in population genetic structure analysis. Mol Ecol Resour 12:873–884
Ruiz-González A, Gurrutxaga M, Cushman SA, Madeira MJ, Randi E, Gómez-Moliner BJ (2014) Landscape genetics for the empirical assessment of resistance surfaces: the European pine marten (Martes martes) as a target species of a regional ecological network. PLoS One 9:e110552
Staal PE (2003) Automobilisme in Nederland. Walburg Pers, Zutphen
Thissen JBM, Achterberg C, Bekker DL (2010) Verspreidingsonderzoek Nederlandse zoogdieren VONZ 2009. Zoogdiervereniging VZZ, Arnhem
Van den Berge K, Broekhuizen S, Müskens GJDM (2000) Voorkomen van de boommarter Martes martes in Vlaanderen en het zuiden van Nederland. Lutra 43:125–136
Van Oosterhout C, Hutchinson WF, Wills DPM, Shipley P (2004) MICRO-CHECKER: software for identifying and correcting genotyping errors in microsatellite data. Mol Ecol Notes 4:535–538
Vitousek PM, Mooney HA, Lubchenco J, Melillo JM (1997) Human domination of Earths ecosystems. Science 277:494–499
Walker CW, Vila C, Landa A, Linden M, Ellegren H (2001) Genetic variation and population structure in Scandinavian wolverine (Gulo gulo) populations. Mol Ecol 10:53–63
Wang J, Santure AW (2009) Parentage and sibship inference from multilocus genotype data under polygamy. Genetics 181:1579–1594
Acknowledgments
S. Broekhuizen and G. Müskens initiated the project, aided in sample collection and administration, and provided valuable comments on a previous version of the manuscript. Special thanks go to the numerous volunteers throughout the Netherlands who also helped to collect samples. We thank R. Bijlsma, C. Pertoldi and L. van der Zande for their aid in setting up the genetic analyses. Thanks to M. van Oene en S. van Gils for help with GIS-maps.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
de Groot, G.A., Hofmeester, T.R., La Haye, M. et al. Hidden dispersal in an urban world: genetic analysis reveals occasional long-distance dispersal and limited spatial substructure among Dutch pine martens. Conserv Genet 17, 111–123 (2016). https://doi.org/10.1007/s10592-015-0765-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10592-015-0765-6