Abstract
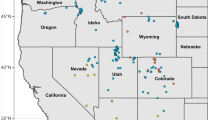
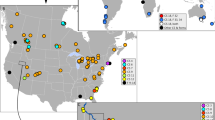
Establishing conservation priorities for invertebrate groups has proven difficult as many proposed units of diversity are based on morphological features that do not reflect evolutionary history. This confusion is confounded by poorly defined ranges of proposed endemic and endangered groups, leading to problems formulating adequate conservation management strategies. We examined one such group, Oreohelix peripherica wasatchensis, a land-snail located in the Wasatch Front Range of Utah that is a candidate for protection under the Endangered Species Act. We employed a broad sampling approach to determine the current range of O. p. wasatchensis, including the type locality, several localities in the surrounding Wasatch Mountains, and localities throughout Utah, Nevada, and Colorado. From these samples, we sequenced the mitochondrial DNA loci 12S and Cytochrome Oxidase I (COI) and nuclear loci Internal Transcribed Spacers 1 and 2 (ITS1 and ITS2) and 5.8S. We estimated phylogenetic relationships using maximum likelihood and Bayesian analyses. Molecular data and radula morphology data indicate that the group identified as O. p. wasatchensis falls into two distinct clades. We recommend further ecological and population assessments of these two distinct mitochondrial clades based on the newly defined range to evaluate its endangered status under the IUCN criteria.


Similar content being viewed by others
References
Binney WG (1886) A second supplement to the fifth volume of the terrestrial air breathing molluscs of the United States and adjacent territories. Bull Mus Comp Zool 13(second suppl):25–48
Boulding EG (1990) Are the opposing selection pressures on exposed and protected shores sufficient to maintain genetic differentiation between gastropod populations with high intermigration rates? Hydrobiologia 193:41–52
Clarke A (1993) Final report: status survey of fifteen species and subspecies of aquatic and terrestrial mollusks from Utah, Colorado, and Montana. US Fish and Wildlife Service Contract No 14-16-0006-91-046 (Revised)
Crandall KA, Bininda-Emonds ORP, Mace GM, Wayne RK (2000) Considering evolutionary processes in conservation biology. TREE 15:290–295
Emberton KC (1995a) When shells do not tell: 145 million years of conchological evolution in North America’s polygyrid land snails. Malacologia 37:69–110
Emberton KC (1995b) Sympatric convergence and environmental correlation between two land-snail species. Evolution 49:469–475
Felsenstein J (1981) Evolutionary trees from DNA sequences: a maximum likelihood approach. J Mol Evol 17:368–376
Felsenstein J (1985) Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39:783–791
Folmer O, Black M, Hoeh W, Lutz R, Vrijenhoek R (1994) DNA primers for amplification of mitochondrial cytochrome C oxidase subunit I from diverse metazoan invertebrates. Mol Mar Biol Biotechnol 3:294–299
Guindon S, Gascuel O (2003) A simple, fast, and accurate algorithm to estimate large phylogenies by maximum likelihood. Syst Biol 52:696–704
Guralnick RP (2005) Combined molecular and morphological approaches for documenting regional biodiversity and ecological patterns in problematic taxa: a case study utilizing the bivalve group Cyclocalyx (Sphaeriidae, Bivalvia) from western North America. Zool Scr 34:469–482
Henderson J (1924) Mollusca of Colorado, Utah, Montana, Idaho, and Wyoming. Library of the University of Colorado, Boulder Colorado
Hershler R, Liu HP (2004) A molecular phylogeny of aquatic gastropods provides a new perspective on biogeographic history of the Snake River Region. Mol Phylogenet Evol 32:927–937
Huelsenbeck JP, Ronquist F (2001) MRBAYES: Bayesian inference for phylogenetic trees. Bioinformatics 17:754–755
International Union for Conservation of Nature and Natural Resources (IUCN) (2002) IUCN red list of threatened species (online at http://www.redlistorg). IUCN, Gland, Switzerland, Cited 15 Sept 2006
Johnson JB, Jordan S (2000) Phylogenetic divergence in leatherside chub (Gila copei) inferred form mitochondrial cytochrome b sequences. Mol Ecol 9:1029–1035
Kane RA, Rollinson D (1994) Repetive sequences in the ribosomal DNA internal transcribed spacer of Schistosoma haematobium, Schistosoma intercalatum and Schistosoma mattheei. Mol Biochem Parasitol 63:153–156
Kocher TD, Thomas WK, Meyer A, Edwards AM, Paabo S, Villablanca FX, Wilson AC (1989) Dynamics of mitochondrial DNA evolution in animals: amplification and sequencing with conserved primers. PNAS 86:6196–6200
Meadows D (2002) Abundance, distribution and dispersal of the Ogden mountainsnail (Oreohelix peripherica wasatchensis). Western American Naturalist, Brigham Young University, Provo, Utah, 25 pp
Miller MP, Stevens LE, Busch JD, Sorensen JA, Keim P (2000) Amplified fragment length polymorphism and mitochondrial sequence data detect genetic differentiation and relationships in endangered southwestern USA amber snails (Oxyloma spp). Can J Zool 78:1845–1854
Morando M, Avila L, Sites JW Jr (2003) Sampling strategies for delimiting species: genes, individuals, and populations in the Liolaemus elongatus-kriegi complex (Squamata: Liolaemidae) in Andean-Patagonian South America. Syst Biol 52:159–185
Moritz C (1994) Defining ‘evolutionarily significant units’ for conservation. TREE 9:373–375
Pfenninger M, Cordellier M, Streit B (2006) Comparing the efficacy of morphologic and DNA-based taxonomy in the freshwater gastropod genus Radix (Basommatophora, Pulmonata). BMC Evol Biol 6:100
Pilsbry HA (1916) Notes on the anatomy of Oreohelix, with a catalog of the species. Proc Acad Nat Sci Philadelphia 68:340–359, pls 19–22
Pilsbry HA (1939) Land Mollusca of North America (North of Mexico). Monographs number 3, vol 1. Academy of Natural Sciences, Philadelphia, PA USA, 1003 pp, 580 figs
Ports MA (2004) Biogeographic and taxonomic relationships among the mountain snails (Gastropoda: Oreohelicidae) of the central Great Basin. West N Am Nat 64:145–154
Posada D, Crandall KA (1998) MODELTEST: testing the model of DNA substitution. Bioinformatics 14:817–818
Ronquist F, Huelsenbeck JP (2003) MrBayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 19:1572–1574
Shimodaira H, Hasegawa M (1999) Multiple comparisons of log-likelihoods with applications to phylogenetic inference. Mol Biol Evol 16:1114–1116
Sites JW Jr, Crandall KA (1997) Testing species boundaries in biodiversity studies. Conserv Biol 11:1289–1297
Sites JW Jr, Marshall JC (2004) Operational criteria for delimiting species. Ann Rev Ecol Evol Syst 35:199–227
Swofford DL (2002) PAUP*: phylogenetic analysis using parsimony, version 4. Sinauer Associates, Sunderland, MA
Templeton AR (2004) Statistical phylogeography: methods of evaluating and minimizing inference errors. Mol Ecol 13:789–810
Thiele J (1866–1893) Docoglossa. In: Troschel FH (eds) Das Gebiss der Schnecken, zur Begründung einer natürlichen classification. Nicolaische Verlags-Buchandlung, Berlin, pp 305–352
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The CLUSTAL_X windows interface: flexible strategies for multiple sequence alignment aided by quality analyses tools. Nucleic Acid Res 25:4876–4882
Turgeon DD, Quinn JF Jr, Bogan AE, Coan EV, Hochberg FG, Lyons WG, Mikkelsen PM, Neves RJ, Roper CFE, Rosenberg G, Roth B, Scheltema A, Thompson FG, Vecchione M, Williams JD (1998) Common and scientific names of aquatic invertebrates from the United States and Canada: Mollusks, 2nd edn. American Fisheries Society Special Publication 26, Bethesda, Maryland, 526 pp
Acknowledgments
This study was supported by the U.S. Forest Service, Wasatch-Cache National Forest, and the Brigham Young University Office of Research and Creative Activities. We would like to thank all those USFS employees who collected samples to complete this work. We thank George Oliver for his help in identification of Oreohelix specimens in the lab and helpful discussions on Oreohelix biology. We thank the anonymous reviewers for their helpful suggestions in improving our manuscript.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Weaver, K.F., Perez-Losada, M., Guralnick, R.P. et al. Assessing the conservation status of the land snail Oreohelix peripherica wasatchensis (Family Oreohelicidae). Conserv Genet 9, 907–916 (2008). https://doi.org/10.1007/s10592-007-9415-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10592-007-9415-y