Abstract
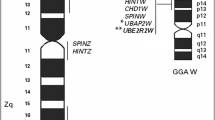
The chicken genome, like those of most avian species, contains numerous microchromosomes that cannot be distinguished by size alone. Unique properties attributed to the microchromosomes include high GC content and gene density, and an enhanced recombination rate. Previously, microchromosome GGA 17 was shown to align with the consensus genetic linkage group E41W17, and bacterial artificial chromosome (BAC) clones containing E41W17 markers were isolated and assigned on the physical BAC map as well as the recently assembled draft chicken genome sequence. For this study, these same BACS were utilized as probes for fluorescence in-situ hybridization (FISH) to develop the GGA 17 cytogenetic map. Here we detail the chromosome order of ten BAC DNAs, thereby deriving a cytogenetic map of GGA 17 that is simultaneously integrated with both the linkage map and genome sequence. The location of the FISH probes together with the morphological appearance of the chromosome suggested that GGA 17 is an acrocentric chromosome whose cytogenetic map orientation is reversed from that currently indicated by the linkage map and draft genome sequence. The reversed orientation and the centromere location of GGA 17 were confirmed experimentally by dual-colour FISH hybridization using terminal BACs and the centromere-specific CNM oligonucleotide as probes. An advantage of this cyto-genomic approach is the improved alignment of the sequence and linkage maps with cytogenetic features such as the centromere, telomeres, p and q arms, and staining patterns indicating GC versus AT content.
Similar content being viewed by others
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Romanov, M.N., Daniels, L.M., Dodgson, J.B. et al. Integration of the cytogenetic and physical maps of chicken chromosome 17. Chromosome Res 13, 215–222 (2005). https://doi.org/10.1007/s10577-005-1506-3
Received:
Revised:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/s10577-005-1506-3