Abstract
Purpose
Tobacco smoke exposure has been associated with altered DNA methylation. However, there is a paucity of information regarding tobacco smoke exposure and DNA methylation of breast tumors.
Methods
We conducted a case-only analysis using breast tumor tissue from 493 postmenopausal and 225 premenopausal cases in the Western New York Exposures and Breast Cancer (WEB) study. Methylation of nine genes (SFN, SCGB3A1, RARB, GSTP1, CDKN2A, CCND2, BRCA1, FHIT, and SYK) was measured with pyrosequencing. Participants reported their secondhand smoke (SHS) and active smoking exposure for seven time periods. Unconditional logistic regression was used to estimate odds ratios (OR) of having methylation higher than the median.
Results
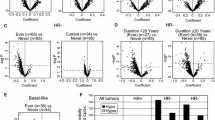
SHS exposure was associated with tumor DNA methylation among postmenopausal but not premenopausal women. Active smoking at certain ages was associated with increased methylation of GSTP1, FHIT, and CDKN2A and decreased methylation of SCGB3A1 and BRCA1 among both pre- and postmenopausal women.
Conclusion
Exposure to tobacco smoke may contribute to breast carcinogenesis via alterations in DNA methylation. Further studies in a larger panel of genes are warranted.
Similar content being viewed by others
References
Macacu A, Autier P, Boniol M, Boyle P (2015) Active and passive smoking and risk of breast cancer: a meta-analysis. Breast Cancer Res Treat 154:213–224
Rosenberg L, Boggs DA, Bethea TN, Wise LA, Adams-Campbell LL, Palmer JR (2013) A prospective study of smoking and breast cancer risk among African American women. Cancer Causes Control. https://doi.org/10.1007/s10552-013-0298-6
Catsburg C, Miller AB, Rohan TE (2015) Active cigarette smoking and risk of breast cancer. Int J Cancer 136:2204–2209
Dossus L, Boutron-Ruault MC, Kaaks R et al (2014) Active and passive cigarette smoking and breast cancer risk: results from the EPIC cohort. Int J Cancer 134:1871–1888
Gram IT, Park SY, Kolonel LN et al (2015) Smoking and risk of breast cancer in a racially/ethnically diverse population of mainly women who do not drink alcohol: the MEC study. Am J Epidemiol 182:917–925
Hecht SS (2002) Tobacco smoke carcinogens and breast cancer. Environ Mol Mutagen. 39:119–126
MacNicoll A, Easty G, Neville A, Grover P, Sims P (1980) Metabolism and activation of carcinogenic polycyclic hydrocarbons by human mammary cells. Biochem Biophys Res Commun 95:1599–1606
McCabe MT, Brandes JC, Vertino PM (2009) Cancer DNA methylation: molecular mechanisms and clinical implications. Clin Cancer Res 15:3927–3937
Johnson KC, Koestler DC, Cheng C, Christensen BC (2014) Age-related DNA methylation in normal breast tissue and its relationship with invasive breast tumor methylation. Epigenetics 9:268–275
Zhu X, Li J, Deng S et al (2016) Genome-wide analysis of DNA methylation and cigarette smoking in a Chinese population. Environ Health Perspect 124:966–973
Freeman JR, Chu S, Hsu T, Huang YT (2016) Epigenome-wide association study of smoking and DNA methylation in non-small cell lung neoplasms. Oncotarget 7(43):69579
Weisenberger DJ, Levine AJ, Long TI et al (2015) Association of the colorectal CpG island methylator phenotype with molecular features, risk factors, and family history. Cancer Epidemiol Biomark Prev 24:512–519
Limsui D, Vierkant RA, Tillmans LS et al (2010) Cigarette smoking and colorectal cancer risk by molecularly defined subtypes. J Natl Cancer Inst 102:1012–1022
Shui IM, Wong CJ, Zhao S et al (2016) Prostate tumor DNA methylation is associated with cigarette smoking and adverse prostate cancer outcomes. Cancer 122:2168–2177
Reynolds LM, Magid HS, Chi GC et al (2016) Secondhand tobacco smoke exposure associations with DNA methylation of the aryl hydrocarbon receptor repressor. Nicot Tob Res 19:442–451
Scesnaite A, Jarmalaite S, Mutanen P et al (2012) Similar DNA methylation pattern in lung tumours from smokers and never-smokers with second-hand tobacco smoke exposure. Mutagenesis 27:423–429
Ambatipudi S, Cuenin C, Hernandez-Vargas H et al (2016) Tobacco smoking-associated genome-wide DNA methylation changes in the EPIC study. Epigenomics 8:599–618
Ivorra C, Fraga MF, Bayon GF et al (2015) DNA methylation patterns in newborns exposed to tobacco in utero. J Trans Med 13:25
Rzehak P, Saffery R, Reischl E et al (2016) Maternal smoking during pregnancy and DNA-methylation in children at age 5.5 years: epigenome-wide-analysis in the European Childhood Obesity Project (CHOP)-Study. PLoS ONE 11:e0155554
Ladd-Acosta C, Shu C, Lee BK et al (2016) Presence of an epigenetic signature of prenatal cigarette smoke exposure in childhood. Environ Res 144:139–148
Lee KW, Richmond R, Hu P et al (2015) Prenatal exposure to maternal cigarette smoking and DNA methylation: epigenome-wide association in a discovery sample of adolescents and replication in an independent cohort at birth through 17 years of age. Environ Health Pers 123:193–199
White AJ, Chen J, Teitelbaum SL et al (2016) Sources of polycyclic aromatic hydrocarbons are associated with gene-specific promoter methylation in women with breast cancer. Environ Res 145:93–100
Pirouzpanah S, Taleban FA, Atri M, Abadi A-R, Mehdipour P (2010) The effect of modifiable potentials on hypermethylation status of retinoic acid receptor-beta2 and estrogen receptor-alpha genes in primary breast cancer. Cancer Causes Control 21:2101–2111
Johnson KC, Koestler DC, Fleischer T et al (2015) DNA methylation in ductal carcinoma in situ related with future development of invasive breast cancer. Clin Epigen 7:015–0094
Muggerud AA, Ronneberg JA, Warnberg F et al (2010) Frequent aberrant DNA methylation of ABCB1, FOXC1, PPP2R2B and PTEN in ductal carcinoma in situ and early invasive breast cancer. Breast Cancer Res 12:7
Tao MH, Marian C, Shields PG et al (2013) Exposures in early life: associations with DNA promoter methylation in breast tumors. J Dev Orig Health Dis 4:182–190
Bonner MR, Nie J, Han D et al (2005) Secondhand smoke exposure in early life and the risk of breast cancer among never smokers (United States). Cancer Causes Control 16:683–689
Tao MH, Shields PG, Nie J et al (2009) DNA hypermethylation and clinicopathological features in breast cancer: the Western New York Exposures and Breast Cancer (WEB) Study. Breast Cancer Res Treat. 114: 559–568
Callahan CL, Wang Y, Marian C et al (2016) DNA methylation and breast tumor clinicopathological features: The Western New York Exposures and Breast Cancer (WEB) study. Epigenetics. 11:1–10
Xu X, Gammon MD, Jefferson E et al (2011) The influence of one-carbon metabolism on gene promoter methylation in a population-based breast cancer study. Epigenetics 6:1276–1283
Feng W, Orlandi R, Zhao N et al (2010) Tumor suppressor genes are frequently methylated in lymph node metastases of breast cancers. BMC Cancer 10:1471–2407
Jeong YJ, Jeong HY, Lee SM, Bong JG, Park SH, Oh HK (2013) Promoter methylation status of the FHIT gene and Fhit expression: association with HER2/neu status in breast cancer patients. Oncol Rep 30:2270–2278
Marzese DM, Hoon DS, Chong KK et al (2012) DNA methylation index and methylation profile of invasive ductal breast tumors. J Mol Diagn 14:613–622
Hsu NC, Huang YF, Yokoyama KK, Chu PY, Chen FM, Hou MF (2013) Methylation of BRCA1 promoter region is associated with unfavorable prognosis in women with early-stage breast cancer. PLoS ONE 8:6
Klajic J, Fleischer T, Dejeux E et al (2013) Quantitative DNA methylation analyses reveal stage dependent DNA methylation and association to clinico-pathological factors in breast tumors. BMC Cancer 13:1471–2407
Arai T, Miyoshi Y, Kim SJ, Taguchi T, Tamaki Y, Noguchi S (2006) Association of GSTP1 CpG islands hypermethylation with poor prognosis in human breast cancers. Breast Cancer Res Treat 100:169–176
Zhang X, Shrikhande U, Alicie BM, Zhou Q, Geahlen RL (2009) Role of the protein tyrosine kinase Syk in regulating cell–cell adhesion and motility in breast cancer cells. Mol Cancer Res 7:634–644
Cheol Kim D, Thorat MA, Lee MR et al (2012) Quantitative DNA methylation and recurrence of breast cancer: a study of 30 candidate genes. Cancer Biomark 11:75–88
Tao MH, Marian C, Nie J et al (2011) Body mass and DNA promoter methylation in breast tumors in the Western New York Exposures and Breast Cancer Study. Am J Clin Nutr 94:831–838
Tao MH, Marian C, Shields PG et al (2011) Alcohol consumption in relation to aberrant DNA methylation in breast tumors. Alcohol. 45:689–699
Zeilinger S, Kühnel B, Klopp N et al (2013) Tobacco Smoking Leads to Extensive Genome-Wide Changes in DNA Methylation. PLoS ONE 8:e63812
Dogan MV, Shields B, Cutrona C et al (2014) The effect of smoking on DNA methylation of peripheral blood mononuclear cells from African American women. BMC Genom 15:151
Gao X, Jia M, Zhang Y, Breitling LP, Brenner H (2015) DNA methylation changes of whole blood cells in response to active smoking exposure in adults: a systematic review of DNA methylation studies. Clin Epigenet 7:113
Joubert BR, Haberg SE, Nilsen RM et al (2012) 450K epigenome-wide scan identifies differential DNA methylation in newborns related to maternal smoking during pregnancy. Environ Health Persp 120:1425–1431
Sadikovic B, Rodenhiser DI (2006) Benzopyrene exposure disrupts DNA methylation and growth dynamics in breast cancer cells. Toxicol Appl Pharmacol. 216:458–468
Peplonska B, Bukowska A, Wieczorek E, Przybek M, Zienolddiny S, Reszka E (2017) Rotating night work, lifestyle factors, obesity and promoter methylation in BRCA1 and BRCA2 genes among nurses and midwives. PLoS One 12:e0178792
Kim H, Kwon YM, Kim JS et al (2004) Tumor-specific methylation in bronchial lavage for the early detection of non-small-cell lung cancer. J Clin Oncol 22:2363–2370
D’Agostini F, Izzotti A, Balansky R, Zanesi N, Croce CM, De Flora S (2006) Early loss of Fhit in the respiratory tract of rodents exposed to environmental cigarette smoke. Cancer Res 66:3936–3941
Savitz DA, Olshan AF (1995) Multiple comparisons and related issues in the interpretation of epidemiologic data. Am J Epidemiol 142:904–908
Disclaimer
This article was prepared while Catherine Callahan and Youjin Wang were employed at the University at Buffalo. The opinions expressed in this article are the authors’ own and do not reflect the view of the National Institutes of Health, the Department of Health and Human Services, or the United States government.
Funding
Catherine L. Callahan was supported by the National Cancer Institute (NCI) Grant R25CA113951.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no potential conflict of interest.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Callahan, C.L., Bonner, M.R., Nie, J. et al. Active and secondhand smoke exposure throughout life and DNA methylation in breast tumors. Cancer Causes Control 30, 53–62 (2019). https://doi.org/10.1007/s10552-018-1102-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10552-018-1102-4