Abstract
Cells have stringent DNA repair pathways that are specific for each different set of DNA lesions which is accomplished through the integration of complex array of proteins. However, BRCA-mutated breast cancer (BC) has defective DNA repair mechanisms. This study aims to investigate differential expression of a large panel of DNA repair markers to characterise DNA repair mechanisms in BRCA-associated tumours compared to sporadic tumours in an attempt to characterise these tumours in routine practice. Immunohistochemistry and tissue microarray technology were applied to a cohort of clinically annotated series of sporadic (n = 1849), BRCA1-mutated (n = 48), and BRCA2-mutated (n = 27) BC. The following DNA damage response (DDR) markers are used; BRCA1, BRCA2, RAD51, Ku70/Ku80, BARD, PARP1 (cleaved), PARP1 (non-cleaved), and P53 in addition to basal cytokeratins, ER, PR, and HER2. A significant proportion of BRCA1 tumours were positive for PARP1 (non-cleaved), and negative for BARD1 and RAD51 compared with sporadic BC. BRCA2 tumours were significantly positive for PARP1 (non-cleaved) compared with sporadic tumours. RAD51 was significantly higher in BRCA1 compared with BRCA2 tumours (p = 0.005). When BRCA1/2 BCs were compared to triple-negative (TN) sporadic tumours of the studied DDR proteins, BARD1 (p < 0.001), PARP1 (non-cleaved) (p < 0.001), and P53 (p = 0.002) remained significantly different in BRCA1/2 tumours compared with TN BC. DNA repair markers showed differential expression in BRCA-mutated tumours, with a substantial degree of disruption of DNA repair pathways in sporadic BC especially TN BC. DNA double-strand break (DSB) repair is assisted by PARP1 expression in BRCA-mutated tumours, whereas the loss of DSB repair via RAD51 is predominant in BRCA1 rather than BRCA2 BC.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
DNA repair deficiencies are well-known risk factors for a variety of cancers [1]. Cells have numerous DNA repair pathways that are specific for each different set of lesions. In each pathway, several proteins are involved that interact with each other in order to guarantee the repair of the damage. When one of the mechanisms becomes inefficient, often some others prosper, turning the DNA repair towards another pathway. When even the alternate mechanism is damaged, the consequent genetic instability leads to cell death [2]. Traditional chemotherapy often employs DNA-damaging agents whose success or failure depends on the DNA repair capacity of the cells [3]. Knowing the damaged pathway can help understand the interaction between the different DNA repair systems and find candidate targets for therapy through the use of the mechanism known as synthetic lethality [4]. For instance, the selective inhibition of PARP (Poly ADP-ribose polymerase, an enzyme involved in base excision repair) leads to the persistence of DNA lesions resulting in chromosomal instability, cell cycle arrest, and subsequent apoptosis leading ultimately to kill selectively the tumour cells [5].
BRCA1 and BRCA2 proteins have been implicated in the repair of double-strand DNA breaks (DSB) to maintain genomic stability by homologous recombination (HR) [6]. BRCA1 is important to recruit DNA repair proteins to the sites of damage, while BRCA2 catalyses the formation of RAD51 filaments on single-stranded DNA at the damaged sites. The BRCA2 homologue Brh2 nucleates RAD51 filament formation at a dsDNA–ssDNA junction [7]. In this study, the immunohistochemical (IHC) expression of a panel of DNA damage repair including BRCA1, BRCA2, RAD51, Ku70/Ku80, BARD, PARP1 (cleaved), and PARP1 (non-cleaved) is assessed in an invasive BC series including a test set of BRCA1/2 mutant cases and a control set of sporadic BC. This panel of markers includes molecular markers essential for both mechanisms of DNA DDR, namely HR and non-homologous end joining (NHEJ), and work in partnership with BRCA1 and BRCA1 [8]. While PARP1 is involved in base excision repair occurring in response to DNA damage [9], BARD1 functions in association with BRCA1 [10]. The RAD51 is a key component of DNA damage repair by the error-free HR mechanism associated with the activation of DSB DNA repair and works in association with BRCA1 and BRCA2 [11]. On the other hand, Ku70/Ku80 is a heterodimer playing crucial roles in the regulation of diverse cellular processes including NHEJ, transcription regulation, and DNA replication [12].
The expression of this selected panel of DNA repair-related proteins in selected tumours of BRCA1 and BRCA2 mutations is compared to sporadic BC, highlighting the differences between hereditary and sporadic cancers with the aim to recognise the specific profile expression pattern in each population. The more insights into the specific DNA repair mechanism in BRCA-mutated and sporadic invasive BC, the more the opportunities of opening new avenues for therapeutic strategies.
Materials and methods
Patient series
This retrospective study was performed on formalin-fixed paraffin-embedded (FFPE) archival tumour tissues from patients being treated at the Nottingham University Hospitals NHS Trust. Representative tumour tissues were prepared as tissue microarrays (TMA), which were subject to IHC using a broad panel of markers of close relevance to BC biology/classification in addition to markers of DNA damage repair. This study was conducted using two independent cohorts of patients, as follows:
BRCA tumours
A total of 75 tumour samples from 68 patients with confirmed germline mutations for BRCA1 or BRCA2, including seven bilateral cancers, were available for inclusion in this study. The BRCA-mutated tumours consisted of 44 BRCA1 mutations and 24 BRCA2 mutations. The average age of patients at diagnosis was 42 years: 40 for BRCA1 carriers and 45 for BRCA2 carriers. Fifteen patients had bilateral cancer, eleven (11/44, 25 %) were BRCA1 carriers, and four (6/24, 16.7 %) were BRCA2 carriers. Thirty-one patients (31/68, 45.6 %) had prophylactic mastectomy. Seven patients (10.3 %) also had ovarian cancer; with 48/68 (70.6 %) patients underwent prophylactic oophorectomy. The most common type of surgery was mastectomy (44 patients, 64.7 %), 22 of them were bilateral mastectomies. For the remaining 24 patients (35.3 %), wide-local excision was performed.
The mean overall survival following surgical intervention was 121.5 and 87.3 months in BRCA1 and BRCA2 patients, respectively, with a total of 11/44 (25 %) deaths occurred in the former, and 8/24 (33.3 %) occurred in the latter. Twelve patients (17.6 %) experienced a recurrence; seven (15.9 %) were BRCA1 carriers and 5 (20.8 %) BRCA2 carriers. Metastatic disease occurred in 19 patients (27.9 %), 12 (27.3 %) in the BRCA1 and 7 (29.2 %) in the BRCA2 population.
Sporadic tumours
The control population was based on a well-characterised consecutive series of early-stage (TNM Stage I–III) primary operable invasive BC from patients (age ≤ 70 years) enrolled into the Nottingham Tenovus Primary Breast Carcinoma Series presented at Nottingham City Hospital between 1986 and 1998 (n = 1844). Patients’ clinical history, tumour characteristics, and information on therapy and outcomes are prospectively maintained. Outcome data included survival status, survival time, cause of death and development, and time to local, regional recurrence or distant metastasis (DM).
Tissue arrays and immunohistochemistry
Tumour samples were arrayed as previously described [13]. Briefly, tissue cores with a diameter of 0.6 mm were punched from the representative tumour regions of each donor FFPE block. Cores were precisely arrayed into a recipient paraffin block using a tissue microarrayer (TMA Grand Master, 3DHistech Ltd).
Immunohistochemical staining was performed on 4 µm sections using Novolink polymer detection system (Leica Biosystems, RE7150-K). Supplementary Table 1 shows the dilution, source and clone of antibodies used in this study. Briefly, tissue slides were deparaffinised with xylene and rehydrated through three changes of alcohol. Antigen retrieval (except for EGFR and HER2) was performed in citrate buffer (pH 6.0) for 20 min using a microwave oven. Endogenous peroxidase activity was blocked by Peroxidase Block for 5 min. Slides were washed with TBS (pH 7.6), followed by the application of Protein Block for 5 min. Following another TBS wash, primary antibody, optimally diluted in Leica antibody diluent (RE7133), was applied and incubated for 60 min. Slides were washed with TBS followed by incubation with Post-Primary Block for 30 min followed by a TBS wash. Novolink polymer was applied for 30 min. DAB working solution made up of 1:20 DAB chromogen in DAB substrate buffer was prepared and applied for 5 min. Slides were counterstained with Novolink haematoxylin for 6 min, dehydrated, and cover-slipped. Negative (omission of the primary antibody) and positive controls were included according to manufacturer datasheet of each antibody.
Evaluation of immunohistochemical staining
The TMA slides were initially assessed by light microscope assessment of staining quality and specificity. Slides were then scanned into high-resolution digital images (0.45 µm/pixel) using a NanoZoomer slide scanner (Hamamtsu Photonics, Welwyn Garden City, UK) and accessed using a web-based interface (Distiller, SlidePath Ltd, Dublin, Ireland). They were scored at ×20 magnification using a minimum of 24″ high-resolution screen (×1920 1080). Cases were scored without knowledge of the BRCA status and patient outcome and were scored by three people (DC, MA, and ARG).
Assessment of staining was based on a semi-quantitative approach using a modified histochemical score (H-score) taking the intensity of staining and the percentage of stained cells into account [14]. For the intensity, a score index of 0, 1, 2, and 3 corresponding to negative, weak, moderate, and strong staining intensity was used, and the percentage of positive cells at each intensity was estimated subjectively. Cut-off points of the other biomarkers included in this study were chosen as per previous publications [15, 16]. BC molecular subtypes were classified based on their IHC expression profile for ER, PgR, HER2, CK5/6, CK14, and EGFR into (1) luminal (ER+ and/or PR+/HER2−); (2) HER2+ (HER2 positive); and (3) Triple-negative (TN; ER−, PR−, HER2−) including basal (TN-Basal, TN and positive for CK5/6, and/or EGFR and/or CK14) and non-basal (TN-non-basal) (negative for all five markers) [16].
Statistical analysis
Statistical analysis was performed using SPSS 17.0 statistical software. Univariate and multivariate analyses were performed by Chi squared and Log-rank tests. Kaplan and Meier (1958) plots were used to visualise the survival distribution of studied patients’ subgroups, with differences in survival estimated using Log-rank tests. A p value < 0.01 (two-tailed) was considered significant. This study adheres to REporting recommendations for tumour MARKer prognostic studies (REMARK) criteria [17].
Results
Clinico-pathological parameters between BRCA-mutated and sporadic breast tumours
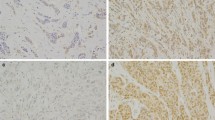
Supplementary Fig. 1 depicts representative examples of IHC expression of DNA damage response markers in invasive BC tissue cores. The histological parameters were compared among the sporadic, BRCA1, and BRCA2 tumours, which are summarised in Table 1. BRCA1 and BRCA2 tumours were of significantly higher grade than the sporadic tumours (p < 0.001) where 90 % of BRCA1 tumours were grade 3 compared with 47 % in the sporadic series. The same applies to the components of histological grade, where less tubule formation and more nuclear pleomorphism were similarly significantly associated with BRCA-mutated tumours. In addition, BRCA1 tumours had a higher mitotic frequency than sporadic and BRCA2 tumours (p < 0.01). Medullary-like tumours were significantly more frequent in BRCA1-mutated tumours compared with sporadic tumours (p < 0.001). There was no significant difference between BRCA2 tumours and sporadic cases in terms of histological type, but lobular tumours were more common in BRCA2 tumours than in BRCA1 tumours (p = 0.010). However, there were no significant differences in tumour size, stage, or presence of vascular invasion observed between the studied series.
Biological characteristics between BRCA-mutated and sporadic BC
A significantly higher proportion of BRCA1-mutated tumours showed a lack of ER expression (83.0 %) compared with the sporadic (22.3 %) and BRCA2 (14.8 %) tumours (p < 0.001, Table 2). PgR was similarly expressed to ER which was more prevalent in the sporadic and BRCA2 tumours than the BRCA1 tumours. Nevertheless, there was no significant difference in the expression of HER2 between the BRCA-mutated tumours and sporadic cases. Triple-negative tumours were highly represented in the BRCA1-mutated tumours (32/48, 67.4 %) compared with both sporadic (16.1 %, p < 0.001) and BRCA2 tumours (7.4 %, p < 0.001).
Likewise, positive basal cytokeratin (CK5/6 and CK14) and EGFR expression was significantly more common in BRCA1 tumours compared with sporadic or BRCA2 tumours. Consequently, basal phenotype was significantly more frequent in BRCA1 tumours (72.7 %) compared with sporadic (24.1 %, p < 0.001) and BRCA2 (24.0 %, p < 0.001) tumours. Additionally, TN-basal tumours were significantly associated with BRCA1-mutated tumours compared with sporadic or BRCA2 tumours (both p < 0.001).
The proliferation marker Ki67 was highly expressed in BRCA1 tumours (43/45, 95.6 %) compared with sporadic BC (920/1453, 63.3 %, p < 0.001). There was no difference between BRCA2 and BRCA1 tumours in terms of Ki67 expression (21/26, 84.0 %).
There was no significant difference in any of the clinico-pathological parameters between TN or TN-basal sporadic and TN BRCA1-mutated BC (data not shown).
Expression of DNA repair markers in BRCA-mutated BC
There was a significant difference in BRCA1 protein expression in both BRCA1- and BRCA2-mutated BC compared with sporadic cases (Table 3). In pairwise comparisons, BRCA1 protein showed significantly different expression between all the studied series. In addition, significant differences were observed between BRCA1/2-mutated and sporadic tumours regarding BRCA2 protein expression (p = 0.007, and p < 0.001, respectively).
PARP1 (non-cleaved) expression was significantly higher in BRCA1 (93.5 %) and BRCA2 (88 %) tumours compared with sporadic cases (48.8 %; p < 0.001). However, P53 expression was not significantly expressed between any of the studied series (p > 0.05).
Significantly lower RAD51 expression was observed in BRCA1-mutated tumours (8/48, 19.5 %) compared with BRCA2 (11/24, 45.8 %; p = 0.024) and sporadic breast cancers (429/896, 47.9 %, p < 0.001). Moreover, BARD1 was significantly less expressed in BRCA1/2-mutated tumours than sporadic tumours (21/45; 46.7 %, 18/24; 75 %, and 1113/1119; 99.5 %, respectively, p < 0.001), and BRCA1- than BRCA2-mutated tumours (p = 0.024), Table 3.
However, PARP1 (cleaved) and Ku70/Ku80 were similarly highly expressed in the majority of BRCA-mutated and sporadic BC (p > 0.05). Furthermore, P53 showed comparably low frequencies of expression in all the studied series with differences not reaching statistical significance (p > 0.05).
The expression of DNA repair markers in sporadic TN and BRCA1/2-mutated BC showed significant differences in the expression of BARD1, PARP1 (non-cleaved), and P53, and BRCA2 expression in BRCA2-mutated BC only (Supplementary Table 2). In this respect, the vast majority of TN BRCA1-mutated tumours were PARP1 positive compared to TN sporadic BC (93.5, and 52.7 % respectively, p < 0.001). On the contrary, 21/45 cases (46.7 %) of the BRCA1-mutated tumours and 18/27 (75 %) of BRCA2-mutated tumours were BARD1 positive compared to 297/297 (100 %) of the TN sporadic tumours (p < 0.001). In line with BARD1 expression, P53 was significantly more frequent in TN sporadic tumours than BRCA1- and BRCA2-mutated tumours (57 and 22.7 and 17.4 %, respectively, p = 0.002). However, none of the remaining studied DNA repair markers were significantly different between TN sporadic and BRCA1/2-mutated tumours (p > 0.05, Supplementary Table 2). To further scrutinise these associations, TN BRCA1 mutant cases were identified (n = 31 cases) and compared to TN sporadic BC regarding the expression of the studied DNA repair markers panel. Interestingly, the same significantly different markers (BARD1, P53, and PARP1 non-cleaved) maintained their significant expression between TN sporadic BC and TN BRCA1 mutant BC, Supplementary Table 3.
BRCA mutations and patient outcome
There was no significant difference in overall survival between BRCA1 and BRCA2 tumours or between BRCA1 versus sporadic or BRCA2 versus sporadic tumours (data not shown). Likewise, there was no significant difference in patient survival between BRCA1 or BRCA2 tumours and sporadic tumours taking into consideration DNA repair markers (data not shown).
Discussion
Etiologically, BC is classified into sporadic and familial forms, with the latter forming up to 5–10 % of invasive BC. BRCA-related familial BC is caused by germline mutations either in the BRCA1 or BRCA2 genes and is associated with an increased risk of developing breast and other cancers [18]. However, in sporadic BC, mutational inactivation of BRCA1/2 is a rare occurrence, as inactivation requires both gene alleles to be mutant or totally deleted. However, non-mutational functional suppression of BRCA1/2 could result from various mechanisms, such as hypermethylation of the BRCA1 promoter or silencing of BRCA2 by other proteins [19, 20]. Moreover, BRCA genes function in a highly coordinated manner in concert with a complex array of genes to carry out high-fidelity repair of DNA damages. However, the expression of these DNA damage repair genes in clinical BC samples and their associations with clinico-pathological parameters have so far yielded controversial findings.
We have used two well-characterised cohorts of invasive BC with updated comprehensive biomarkers, including DNA damage response proteins, and outcome data to assess the pathobiological and clinical features of tumours harbouring BRCA1 and BRCA2 mutations compared to sporadic BC.
Regarding their clinico-pathological criteria, BRCA1/2 tumours when compared with sporadic tumours were significantly poorly differentiated/higher grade with less tubule formation and more nuclear pleomorphism. BRCA1 tumours had a higher mitotic frequency than sporadic and BRCA2 tumours. The vast majority of BRCA1 tumours showed higher proliferative index as assessed by Ki67LI compared to sporadic cancers. However, BRCA2 and BRCA1 tumours were neither different from each other regarding their Ki67LI, nor the BRCA2 tumours were significantly different from the sporadic tumours. These findings go in line with previous reports [21, 22]. Moreover, medullary-like tumours were significantly more frequent in BRCA1-mutated tumours. Although the latter has long been recognised [23], it underscores the potential significance of histologic observations, medullary histologic criteria reported by pathologists in patients’ clinical care [24]. Nevertheless, BRCA2 tumours were not significantly different from sporadic cases in terms of histological type. However, lobular tumours were more common in BRCA2 tumours than in BRCA1 tumours.
The majority of BRCA1-mutated tumours were ER negative compared to sporadic and BRCA2 tumours with similar expression pattern observed for PgR. Moreover, TN tumours were significantly more represented in the BRCA1-mutated tumours compared with both sporadic and BRCA2 tumours. Furthermore, BRCA1 tumours displayed significantly higher proportions of basal BC than sporadic and BRCA2 tumours as evidenced by significant session of the basal markers CK5/6, CK14, and EGFR. Accordingly, TN-basal tumours were significantly associated with BRCA1-mutated tumours compared with sporadic or BRCA2 tumours. These findings are in agreement with our previous report [16] and those of gene microarray studies [25]. Additionally, BRCA1-mutated BCs were not significantly different from TN-non-basal or TN-basal sporadic tumours in any of the known clinico-pathological parameters. These findings describing the clinico-pathological associations of BRCA1/2 tumours are in agreement with those previously described [26]. BRCA2 tumours appear to show a phenotype between sporadic and BRCA1‐associated BC. In other words, the IHC profile of BRCA1-mutated BC is distinctively different from sporadic BC more than BRCA2-mutated tumours.
Regarding the biomarkers’ profile of DNA damage repair markers in the studied series, there was, as expected, a significantly reduced expression of BRCA1 and BRCA2 proteins in both BRCA1- and BRCA2-mutated BC compared with sporadic cases. Moreover, BRCA1 protein was significantly more expressed in BRCA1-mutated tumours compared to BRCA2-mutated tumours. However, BRCA2 protein did not show significant difference in expression between BRCA1- versus BRCA2-mutated tumours.
PARP1 is a known key facilitator of DNA repair and is implicated in pathways of carcinogenesis. It is a 113 kDa nuclear enzyme which is cleaved into two fragments (89 and 24 kDa) during apoptotic cell death [27]. In this study, only non-cleaved PARP1 showed significantly different expression between the studied series, while the cleaved PARP1, whose levels are known to be increased in apoptosis, did not show any significantly different expression. The vast majority of BRCA1 (93.5 %) and BRCA2 (88.5 %) tumours were significantly positive for PARP1 (non-cleaved) expression compared to only half of the sporadic cases. With BRCA1 or BRCA2 loss of function due to mutations, cells become deficient in DNA DSB repair. This in turn activates PARPs whose catalytic activity is immediately stimulated by DNA breaks. It is not, therefore, surprising for the BRCA1/2 cases in our series overexpressing PARP1 (non-cleaved), which appears to be the reactive response of cells when DNA damage repair by BRCA gene is defective, and has been recently reported in BRCA1 mutant breast cancer cell lines and BRCA1-mutated BC cases [28]. Our results are in line with previous studies [27, 29, 30]. The relatively high percentages of sporadic BC positive for PARP1 (non-cleaved) prompted some authorities to suggest the potential therapeutic benefits of PARP1 inhibitors not only in familial BRCA-mutated BC but in sporadic cancers as well [31]. On the other hand, PARP1 cleaved isoform which has been regarded as a useful hallmark of activated cellular apoptotic machinery [9] did not show any significantly different expression between the studied series, with the vast majority of cases in all series showing positive expression. In our recent report, cleaved PARP1 was found to be highly significantly associated with other DNA repair proteins including RAD51, CHK1, CHK2, and others [32]. This overexpression of cleaved PARP1 in our series could point out to its roles in DNA repair, in addition to the recently reported functions of transcriptional regulation of other molecular regulators [33].
BARD1 gene encodes a protein that forms heterodimers with BRCA1 N-terminal region. This stable BARD1/BRCA1 complex is crucial for BRCA1 tumour suppression and coordinates a diverse range of cellular pathways such as DNA repair, transcriptional regulation to maintain genomic stability, and others [34]. BARD1 in this study was significantly down-regulated in the BRCA-mutated series compared to sporadic BC series. Nearly all cases of the latter showed positive expression, while three-quarters and up to half of the BRCA1-mutated cases and BRCA2-mutated cases were BARD1 positive, respectively. The concomitant reduction of BARD1 expression in BRCA1 mutant cases underscores that the participation of BARD1 may be required for proper functioning of BRCA1-mediated tumour suppressor activity aiming ultimately to maintain chromosome integrity through HR mechanism [10].
RAD51 is a key component of DNA damage repair by HR mechanism associated with the activation of DSB DNA repair. It binds to single- and double-stranded DNA giving rise to a RAD51 nucleoprotein filament, which is essential for strand-pairing reactions during DNA recombination [11, 35]. Both BRCA1 and BRCA2 co-localise with RAD51 at sites of DNA damage and activate HR repair of DSB mediated by RAD51 [36]. In this study, significantly lower RAD51 expression was observed in BRCA1-mutated tumours relative to BRCA2 and sporadic BC. Thus, BRCA1 dysfunction caused by inactivating mutations appears to deregulate nuclear RAD51 levels. Similar findings were reported in BRCA1 mutant ovarian cancer cell lines which displayed lowest levels of BRCA1 and RAD51 [37]. Moreover, lower RAD51 nuclear expression was observed in prostatic carcinomas associated with BRCA1/2 mutations than sporadic cancers [38]. Reduced nuclear expression of RAD51 has been reported concomitantly with reduced nuclear expression of BRCA1 and BRCA2 protein expression in early invasive BC [39]. However, in our study, BRCA2 mutated tumours were not significantly different from the sporadic BC. Although BRCA2 is needed for RAD51 nuclear localisation [40], other BRCA2-independent mechanisms involving other proteins, for instance RAD51C, have also been proposed [6].
The tumour suppressor P53 showed closely comparable low expression in BRCA1/2 and sporadic tumours with no significant differences between the studied series. In other words, up to three-quarters to around four-fifths of the studied series have negative P53 expression. These results of low P53 expression in BRCA mutant BC are in agreement with the findings reported by Zakhartseva and co-authors in invasive BC [41]. However, they are contradicting other studies describing the collaborative synergistic functionality of P53 mutations for the tumourigenic influence of BRCA2 loss [42] and BRCA1 loss [43]. P53 inactivation through protein-truncating mutations, rather than hotspot mutations, has been reported to be one of the mechanisms which accompany BRCA loss, which could not be always detected by IHC expression of P53 protein. This could, at least in part, explain the low levels of P53 expression, and hence inactivation, in our BRCA mutant series. Accordingly, this has prompted some authorities to report on the inherent weakness of immunohistochemical detection of TP53 inactivation that could lead to misdiagnosis in significant proportions of BRCA1 mutant tumours [43].
Ku70/Ku80 is a heterodimer known to play crucial roles in regulating diverse cellular processes including NHEJ, transcription regulation, and DNA replication [12]. In this study, Ku70/Ku80 was similarly highly expressed in the studied series, where all BRCA2-mutated and 95.6 % of BRCA1-mutated BC cases showed positive expression. These results are in agreement with our recent report using IHC on invasive BC cases and reverse phase protein array on cell lysates from BRCA1-deficient BC cell lines [44]. These figures might indicate over-activation of non-homologous end joining (NHEJ) known to be mediated by Ku70/Ku80 as a back-up alternative mechanism for DNA DSB repair in cases when BRCA1/2 are mutated with altered HR DNA repair pathway.
Currently, it is widely accepted that the biologic processes carried out by BRCA1 are disrupted by numerous mechanisms in sporadic cancers especially the TN and basal-like BC. Therefore, we have restricted the analysis to compare TN sporadic BC with BRCA1/2-mutated tumours. Expectedly, there were no statistically significant differences between the TN and basal-like phenotypes regarding any of the clinico-pathological parameters. Interestingly, both had large proportions of large tumour size and the vast majority were high-grade and moderate/poor NPI. Of the DNA repair markers studied, PARP1 (non-cleaved) showed significantly more positivity in BRCA1-mutated tumours. Although more than 90 % of the latter showed PARP1 overexpression, up to 53 % of the TN sporadic tumours were also PARP1 (non-cleaved) positive. Moreover, both phenotypes showed more BRCA1/2-negative, RAD51-negative, and Ku70/Ku80-positive expression. The same pattern of association were maintained when the analyses were further restricted between TN BRCA1-mutated cases and TN sporadic BC. In other terms, in both phenotypes, the HR protein RAD51 showed reduced expression, while the NHEJ protein Ku70/Ku80 was similarly overexpressed. Furthermore, BARD1 and the tumour suppressor P53 showed significantly lower expression in BRCA1/2 mutant and TN BRCA1/2 mutant tumours compared with TN sporadic BC cases. Collectively, these findings refer to the properties that define ‘BRCAness’ of sporadic BC. These are the traits that some sporadic cancers could share with those occurring in either BRCA1 or BRCA2 mutation carriers [45, 46]. These shared properties between the TN sporadic and familial cancers might have important implications in clinical management of these cancers. This highlights the potential therapeutic benefit of PARP1 inhibitors, based on the hypothesis of synthetic lethality, in TN sporadic as well as BRCA1-mutated BC [47].
In conclusion, the results of this study demonstrate, at a translational level, the complexity of DNA repair mechanisms in BRCA-mutated tumours and the presence of a degree of disruption of these pathways especially in TN BC. Moreover, our results support the hypothesis that DSBs are repaired by one or more alternative pathways and they are not independent of each other as evidenced by the reciprocal relationship between markers of HR and NHEJ of DNA DSB. Furthermore, DSB DNA repair is assisted by PARP1 expression in BRCA-mutated tumours, whereas the loss of DSB repair via RAD51 is predominant in BRCA1 rather than BRCA2 BC.
References
Wu P-E, Shen C-Y (2011) ‘Hide-then-hit’ to explain the importance of genotypic polymorphism of DNA repair genes in determining susceptibility to cancer. J Mol Cell Biol 3(1):59–65
Polo SE, Jackson SP (2011) Dynamics of DNA damage response proteins at DNA breaks: a focus on protein modifications. Genes Dev 25(5):409–433
Cheung-Ong K, Giaever G, Nislow C (2013) DNA-damaging agents in cancer chemotherapy: serendipity and chemical biology. Chem Biol 20(5):648–659
Hosoya N, Miyagawa K (2014) Targeting DNA damage response in cancer therapy. Cancer Sci 105(4):370–388
Morales J, Li L, Fattah FJ, Dong Y, Bey EA, Patel M, Gao J, Boothman DA (2014) Review of poly (ADP-ribose) polymerase (PARP) mechanisms of action and rationale for targeting in cancer and other diseases. Crit Rev Eukaryot Gene Expr 24(1):15–28
Gildemeister OS, Sage JM, Knight KL (2009) Cellular redistribution of Rad51 in response to DNA damage: novel role for Rad51C. J Biol Chem 284(46):31945–31952
Gudmundsdottir K, Ashworth A (2006) The roles of BRCA1 and BRCA2 and associated proteins in the maintenance of genomic stability. Oncogene 25(43):5864–5874
Tutt A, Ashworth A (2002) The relationship between the roles of BRCA genes in DNA repair and cancer predisposition. Trends Mol Med 8(12):571–576
Virag L (2005) Structure and function of poly(ADP-ribose) polymerase-1: role in oxidative stress-related pathologies. Curr Vasc Pharmacol 3(3):209–214
Westermark UK, Reyngold M, Olshen AB, Baer R, Jasin M, Moynahan ME (2003) BARD1 participates with BRCA1 in homology-directed repair of chromosome breaks. Mol Cell Biol 23(21):7926–7936
Forget AL, Kowalczykowski SC (2010) Single-molecule imaging brings Rad51 nucleoprotein filaments into focus. Trends Cell Biol 20(5):269–276
Skorokhod OM, Kravchuk IV, Teleheiev HD, Maliuta SS (2006) Role of Ku protein in normal and cancer cells. Ukr Biokhim Zh 78(5):5–15
Kononen J, Bubendorf L, Kallioniemi A, Barlund M, Schraml P, Leighton S, Torhorst J, Mihatsch MJ, Sauter G, Kallioniemi OP (1998) Tissue microarrays for high-throughput molecular profiling of tumor specimens. Nat Med 4(7):844–847
McCarty KS Jr, McCarty KS Sr (1984) Histochemical approaches to steroid receptor analyses. Semin Diagn Pathol 1(4):297–308
Rakha EA, El-Sayed ME, Green AR, Lee AH, Robertson JF, Ellis IO (2007) Prognostic markers in triple-negative breast cancer. Cancer 109(1):25–32
Rakha EA, Elsheikh SE, Aleskandarany MA, Habashi HO, Green AR, Powe DG, El-Sayed ME, Benhasouna A, Brunet JS, Akslen LA et al (2009) Triple-negative breast cancer: distinguishing between basal and nonbasal subtypes. Clin Cancer Res 15(7):2302–2310
McShane LM, Altman DG, Sauerbrei W, Taube SE, Gion M, Clark GM (2005) Reporting recommendations for tumor marker prognostic studies (REMARK). J Natl Cancer Inst 97(16):1180–1184
Edlich RF, Winters KL, Lin KY (2005) Breast cancer and ovarian cancer genetics. J Long Term Eff Med Implants 15(5):533–545
Hughes-Davies L, Huntsman D, Ruas M, Fuks F, Bye J, Chin SF, Milner J, Brown LA, Hsu F, Gilks B et al (2003) EMSY links the BRCA2 pathway to sporadic breast and ovarian cancer. Cell 115(5):523–535
Korkola J, Gray JW (2010) Breast cancer genomes––form and function. Curr Opin Genet Dev 20(1):4–14
van der Groep P, Bouter A, van der Zanden R, Siccama I, Menko FH, Gille JJ, van Kalken C, van der Wall E, Verheijen RH, van Diest PJ (2006) Distinction between hereditary and sporadic breast cancer on the basis of clinicopathological data. J Clin Pathol 59(6):611–617
Chappuis PO, Nethercot V, Foulkes WD (2000) Clinico-pathological characteristics of BRCA1- and BRCA2-related breast cancer. Semin Surg Oncol 18(4):287–295
Eisinger F, Jacquemier J, Charpin C, Stoppa-Lyonnet D, Bressac-de Paillerets B, Peyrat JP, Longy M, Guinebretiere JM, Sauvan R, Noguchi T et al (1998) Mutations at BRCA1: the medullary breast carcinoma revisited. Cancer Res 58(8):1588–1592
Kleer CG (2009) Carcinoma of the breast with medullary-like features: diagnostic challenges and relationship with BRCA1 and EZH2 functions. Arch Pathol Lab Med 133(11):1822–1825
Turner NC, Reis-Filho JS (2006) Basal-like breast cancer and the BRCA1 phenotype. Oncogene 25(43):5846–5853
Mangia A, Malfettone A, Simone G, Darvishian F (2011) Old and new concepts in histopathological characterization of familial breast cancer. Ann Oncol 22(Suppl 1):i24–i30
Ossovskaya V, Koo IC, Kaldjian EP, Alvares C, Sherman BM (2010) Upregulation of poly (ADP-Ribose) polymerase-1 (PARP1) in triple-negative breast cancer and other primary human tumor types. Genes Cancer 1(8):812–821
De Summa S, Pinto R, Pilato B, Sambiasi D, Porcelli L, Guida G, Mattioli E, Paradiso A, Merla G, Micale L et al (2014) Expression of base excision repair key factors and miR17 in familial and sporadic breast cancer. Cell Death Dis 5:e1076
Domagala P, Huzarski T, Lubinski J, Gugala K, Domagala W (2011) PARP-1 expression in breast cancer including BRCA1-associated, triple negative and basal-like tumors: possible implications for PARP-1 inhibitor therapy. Breast Cancer Res Treat 127(3):861–869
Domagala P, Huzarski T, Lubinski J, Gugala K, Domagala W (2011) Immunophenotypic predictive profiling of BRCA1-associated breast cancer. Virchows Arch 458(1):55–64
Frizzell K, Kraus WL (2009) PARP inhibitors and the treatment of breast cancer: beyond BRCA1/2? Breast Cancer Res 11(6):111
Green A, Caracappa D, Benhasouna A, Alshareeda A, Nolan C, Macmillan RD, Madhusudan S, Ellis I, Rakha E (2014) Biological and clinical significance of PARP1 protein expression in breast cancer. Breast Cancer Res Treat 1–10
Erener S, Pétrilli V, Kassner I, Minotti R, Castillo R, Santoro R, Hassa PO, Tschopp J, Hottiger MO (2012) Inflammasome-activated caspase 7 cleaves PARP1 to enhance the expression of a subset of NF-κB target genes. Mol Cell 46(2):200–211
Wu LC, Wang ZW, Tsan JT, Spillman MA, Phung A, Xu XL, Yang MC, Hwang LY, Bowcock AM, Baer R (1996) Identification of a RING protein that can interact in vivo with the BRCA1 gene product. Nat Genet 14(4):430–440
Daley JM, Kwon Y, Niu H, Sung P (2013) Investigations of homologous recombination pathways and their regulation. Yale J Biol Med 86(4):453–461
Cousineau I, Abaji C, Belmaaza A (2005) BRCA1 regulates RAD51 function in response to DNA damage and suppresses spontaneous sister chromatid replication slippage: implications for sister chromatid cohesion, genome stability, and carcinogenesis. Cancer Res 65(24):11384–11391
Sun C, Li N, Yang Z, Zhou B, He Y, Weng D, Fang Y, Wu P, Chen P, Yang X et al (2013) miR-9 regulation of BRCA1 and ovarian cancer sensitivity to cisplatin and PARP inhibition. J Natl Cancer Inst 105:1750–1758
Mitra A, Jameson C, Barbachano Y, Sanchez L, Kote-Jarai Z, Peock S, Sodha N, Bancroft E, Fletcher A, Cooper C et al (2009) Overexpression of RAD51 occurs in aggressive prostatic cancer. Histopathology 55(6):696–704
Söderlund K, Skoog L, Fornander T, Askmalm MS (2007) The BRCA1/BRCA2/Rad51 complex is a prognostic and predictive factor in early breast cancer. Radiother Oncol 84(3):242–251
Davies AA, Masson J-Y, McIlwraith MJ, Stasiak AZ, Stasiak A, Venkitaraman AR, West SC (2001) Role of BRCA2 in control of the RAD51 recombination and DNA repair protein. Mol Cell 7(2):273–282
Zakhartseva LM, Gorovenko NG, Podolskaya SV, Anikusko NF, Lobanova OE, Pekur KA, Kropelnytskyi VA, Shurygina OV (2009) Breast cancer immunohistochemical features in young women with BRCA 1/2 mutations. Exp Oncol 31(3):174–178
Jonkers J, Meuwissen R, van der Gulden H, Peterse H, van der Valk M, Berns A (2001) Synergistic tumor suppressor activity of BRCA2 and p53 in a conditional mouse model for breast cancer. Nat Genet 29(4):418–425
Holstege H, Joosse SA, van Oostrom CTM, Nederlof PM, de Vries A, Jonkers J (2009) High incidence of protein-truncating TP53 mutations in BRCA1-related breast cancer. Cancer Res 69(8):3625–3633
Alshareeda AT, Negm OH, Albarakati N, Green AR, Nolan C, Sultana R, Madhusudan S, Benhasouna A, Tighe P, Ellis IO et al (2013) Clinicopathological significance of KU70/KU80, a key DNA damage repair protein in breast cancer. Breast Cancer Res Treat 139(2):301–310
De Summa S, Pinto R, Sambiasi D, Petriella D, Paradiso V, Paradiso A, Tommasi S (2013) BRCAness: a deeper insight into basal-like breast tumors. Ann Oncol 24(Suppl 8):viii13–viii21
Turner N, Tutt A, Ashworth A (2004) Hallmarks of ‘BRCAness’ in sporadic cancers. Nat Rev Cancer 4(10):814–819
Dedes KJ, Wetterskog D, Weigelt B, Ashworth A, Reis-Filho JS (2011) Synthetic lethality of PARP inhibition in cancers lacking BRCA1 and BRCA2 mutations. (1551-4005 (Electronic)). Cell Cycle 10:1192–1199
Acknowledgments
DC was sponsored by The Vito Distante Project in Breast Cancer Clinical Research, Florence, Italy.
Conflict of interest
The authors have no conflict of interest to declare.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Supplementary material 1 (JPEG 128 kb)
Supplementary Fig. 1: Detection of DNA repair markers in invasive BC as determined by IHC. Representative examples of A) BRCA1, B) BRCA2, C) RAD51, D) BARD1, E) PARP-1 (noncleaved), F) PARP-1 (cleaved), G) Ku70/ku80, and H) p53, (Original magnification, 100 ×)
Supplementary material 2 (DOCX 15 kb)
Supplementary Table 1: Dilution, source/clone and pre-treatment for the antibodies used in this study
Supplementary material 3 (DOCX 17 kb)
Supplementary Table 2: Statistical associations of DNA repair markers in sporadic Triple Negative BC and BRCA1/2-mutated BC
Supplementary material 4 (DOCX 15 kb)
Supplementary Table 3: DNA repair markers in sporadic TN BC and BRCA1-mutated triple-negative breast cancer
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution Noncommercial License which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and the source are credited.
About this article
Cite this article
Aleskandarany, M., Caracappa, D., Nolan, C.C. et al. DNA damage response markers are differentially expressed in BRCA-mutated breast cancers. Breast Cancer Res Treat 150, 81–90 (2015). https://doi.org/10.1007/s10549-015-3306-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10549-015-3306-6