Abstract
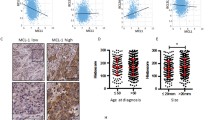
Only a minority of breast cancer patients responds to chemotherapy and we lack predictive biomarkers that help to select a patient-tailored therapy that takes into consideration the molecular heterogeneity of the cancer type. Responsiveness to the clinically important nucleoside analogs gemcitabine and decitabine may be critically determined by Deoxycytidine kinase (DCK) expression as this enzyme is required to convert the inactive prodrugs into their pharmacologically active forms. Here, we examined whether DCK is differentially expressed in breast cancer and evaluated whether DCK expression levels control responsiveness to these nucleoside analogs in vitro by experimentally modulating DCK expression levels. We examined DCK expression in gene expression data sets of breast tumors including the series of 295 consecutive patients that have been classified into low or high risk for recurrence using the MammaPrint 70 gene profile. We found that DCK is expressed at higher levels in patients having poor clinical outcome as judged by the MammaPrint assay. As such, patients that have a poor prognosis may thus be susceptible to treatment with nucleoside analogs. In support of this, we found a causal relationship between DCK levels and sensitivity to these nucleoside analogs in breast cancer cell lines. The data indicate that breast cancers that are at high risk of recurrence express higher levels of DCK, which we find to be strongly correlated to a favorable response to nucleoside analogs. The data suggest that DCK expression in breast cancer could be exploited to select patients that are likely to respond to treatment with nucleoside analogs.




Similar content being viewed by others
References
WHO (2009) Fact sheet 297
TG EBC (2005) Effects of chemotherapy and hormonal therapy for early breast cancer on recurrence and 15-year survival: an overview of the randomised trials. Lancet 365(9472):1687–1717. doi:10.1016/S0140-6736(05)66544-0
Clarke M, Collins R, Darby S, Davies C, Elphinstone P, Evans E, Godwin J, Gray R, Hicks C, James S, MacKinnon E, McGale P, McHugh T, Peto R, Taylor C, Wang Y (2005) Effects of radiotherapy and of differences in the extent of surgery for early breast cancer on local recurrence and 15-year survival: an overview of the randomised trials. Lancet 366(9503):2087–2106. doi:10.1016/S0140-6736(05)67887-7
Podsypanina K, Du YC, Jechlinger M, Beverly LJ, Hambardzumyan D, Varmus H (2008) Seeding and propagation of untransformed mouse mammary cells in the lung. Science 321(5897):1841–1844. doi:10.1126/science.1161621
Husemann Y, Geigl JB, Schubert F, Musiani P, Meyer M, Burghart E, Forni G, Eils R, Fehm T, Riethmuller G, Klein CA (2008) Systemic spread is an early step in breast cancer. Cancer Cell 13(1):58–68. doi:10.1016/j.ccr.2007.12.003
Bernards R (2010) It’s diagnostics, stupid. Cell 141(1):13–17. doi:10.1016/j.cell.2010.03.018
Tan DS, Gerlinger M, Teh BT, Swanton C (2010) Anti-cancer drug resistance: Understanding the mechanisms through the use of integrative genomics and functional RNA interference. Eur J Cancer. doi: 10.1016/j.ejca.2010.03.019
Berns K, Horlings HM, Hennessy BT, Madiredjo M, Hijmans EM, Beelen K, Linn SC, Gonzalez-Angulo AM, Stemke-Hale K, Hauptmann M, Beijersbergen RL, Mills GB, van de Vijver MJ, Bernards R (2007) A functional genetic approach identifies the PI3K pathway as a major determinant of trastuzumab resistance in breast cancer. Cancer Cell 12(4):395–402. doi:10.1016/j.ccr.2007.08.030
Farmer H, McCabe N, Lord CJ, Tutt AN, Johnson DA, Richardson TB, Santarosa M, Dillon KJ, Hickson I, Knights C, Martin NM, Jackson SP, Smith GC, Ashworth A (2005) Targeting the DNA repair defect in BRCA mutant cells as a therapeutic strategy. Nature 434(7035):917–921. doi:10.1038/nature03445
Sawyers CL (2008) The cancer biomarker problem. Nature 452(7187):548–552. doi:10.1038/nature06913
van t’Veer LJ, Dai H, van de Vijver MJ, He YD, Hart AA, Mao M, Peterse HL, van der Kooy K, Marton MJ, Witteveen AT, Schreiber GJ, Kerkhoven RM, Roberts C, Linsley PS, Bernards R, Friend SH (2002) Gene expression profiling predicts clinical outcome of breast cancer. Nature 415(6871):530–536. doi:10.1038/415530a
van de Vijver MJ, He YD, van t’Veer LJ, Dai H, Hart AA, Voskuil DW, Schreiber GJ, Peterse JL, Roberts C, Marton MJ, Parrish M, Atsma D, Witteveen A, Glas A, Delahaye L, van der Velde T, Bartelink H, Rodenhuis S, Rutgers ET, Friend SH, Bernards R (2002) A gene-expression signature as a predictor of survival in breast cancer. N Engl J Med 347(25):1999–2009. doi:10.1056/NEJMoa021967
Blackstock AW, Lightfoot H, Case LD, Tepper JE, Mukherji SK, Mitchell BS, Swarts SG, Hess SM (2001) Tumor uptake and elimination of 2′,2′-difluoro-2′-deoxycytidine (gemcitabine) after deoxycytidine kinase gene transfer: correlation with in vivo tumor response. Clin Cancer Res 7(10):3263–3268
Hapke DM, Stegmann AP, Mitchell BS (1996) Retroviral transfer of deoxycytidine kinase into tumor cell lines enhances nucleoside toxicity. Cancer Res 56(10):2343–2347
Qin T, Jelinek J, Si J, Shu J, Issa JP (2009) Mechanisms of resistance to 5-aza-2′-deoxycytidine in human cancer cell lines. Blood 113(3):659–667. doi:10.1182/blood-2008-02-140038
Beausejour CM, Gagnon J, Primeau M, Momparler RL (2002) Cytotoxic activity of 2′,2′-difluorodeoxycytidine, 5-aza-2′-deoxycytidine and cytosine arabinoside in cells transduced with deoxycytidine kinase gene. Biochem Biophys Res Commun 293(5):1478–1484. doi:10.1016/S0006-291X(02)00413-8
Stegmann AP, Honders MW, Hagemeijer A, Hoebee B, Willemze R, Landegent JE (1995) In vitro-induced resistance to the deoxycytidine analogues cytarabine (AraC) and 5-aza-2′-deoxycytidine (DAC) in a rat model for acute myeloid leukemia is mediated by mutations in the deoxycytidine kinase (DCK) gene. Ann Hematol 71(1):41–47
Brummelkamp TR, Bernards R, Agami R (2002) A system for stable expression of short interfering RNAs in mammalian cells. Science 296(5567):550–553. doi:10.1126/science.1068999
Glas AM, Floore A, Delahaye LJ, Witteveen AT, Pover RC, Bakx N, Lahti-Domenici JS, Bruinsma TJ, Warmoes MO, Bernards R, Wessels LF, Van’t Veer LJ (2006) Converting a breast cancer microarray signature into a high-throughput diagnostic test. BMC Genomics 7:278. doi:10.1186/1471-2164-7-278
McShane LM, Altman DG, Sauerbrei W, Taube SE, Gion M, Clark GM (2005) Reporting recommendations for tumor marker prognostic studies. J Clin Oncol 23(36):9067–9072. doi:10.1200/JCO.2004.01.0454
Rhodes DR, Yu J, Shanker K, Deshpande N, Varambally R, Ghosh D, Barrette T, Pandey A, Chinnaiyan AM (2004) ONCOMINE: a cancer microarray database and integrated data-mining platform. Neoplasia 6(1):1–6
Richardson AL, Wang ZC, De Nicolo A, Lu X, Brown M, Miron A, Liao X, Iglehart JD, Livingston DM, Ganesan S (2006) X chromosomal abnormalities in basal-like human breast cancer. Cancer Cell 9(2):121–132. doi:10.1016/j.ccr.2006.01.013
van’t Veer LJ, Bernards R (2008) Enabling personalized cancer medicine through analysis of gene-expression patterns. Nature 452(7187):564–570. doi:10.1038/nature06915
Stegmann AP, Honders MW, Willemze R, Landegent JE (1995) De novo induced mutations in the deoxycytidine kinase (DCK) gene in rat leukemic clonal cell lines confer resistance to cytarabine (AraC) and 5-aza-2′-deoxycytidine (DAC). Leukemia 9(6):1032–1038
Ohhashi S, Ohuchida K, Mizumoto K, Fujita H, Egami T, Yu J, Toma H, Sadatomi S, Nagai E, Tanaka M (2008) Down-regulation of deoxycytidine kinase enhances acquired resistance to gemcitabine in pancreatic cancer. Anticancer Res 28(4B):2205–2212
Galmarini CM, Clarke ML, Jordheim L, Santos CL, Cros E, Mackey JR, Dumontet C (2004) Resistance to gemcitabine in a human follicular lymphoma cell line is due to partial deletion of the deoxycytidine kinase gene. BMC Pharmacol 4:8. doi:10.1186/1471-2210-4-8
Stegmann AP, Honders WH, Willemze R, Ruiz van Haperen VW, Landegent JE (1995) Transfection of wild-type deoxycytidine kinase (dck) cDNA into an AraC- and DAC-resistant rat leukemic cell line of clonal origin fully restores drug sensitivity. Blood 85(5):1188–1194
Rejiba S, Bigand C, Parmentier C, Hajri A (2009) Gemcitabine-based chemogene therapy for pancreatic cancer using Ad-dCK:UMK GDEPT and TS/RR siRNA strategies. Neoplasia 11(7):637–650
Szatmari T, Huszty G, Desaknai S, Spasokoukotskaja T, Sasvari-Szekely M, Staub M, Esik O, Safrany G, Lumniczky K (2008) Adenoviral vector transduction of the human deoxycytidine kinase gene enhances the cytotoxic and radiosensitizing effect of gemcitabine on experimental gliomas. Cancer Gene Ther 15(3):154–164. doi:10.1038/sj.cgt.7701115
Sebastiani V, Ricci F, Rubio-Viqueira B, Kulesza P, Yeo CJ, Hidalgo M, Klein A, Laheru D, Iacobuzio-Donahue CA (2006) Immunohistochemical and genetic evaluation of deoxycytidine kinase in pancreatic cancer: relationship to molecular mechanisms of gemcitabine resistance and survival. Clin Cancer Res 12(8):2492–2497. doi:10.1158/1078-0432.CCR-05-2655
Stal O, Dufmats M, Hatschek T, Carstensen J, Klintenberg C, Rutqvist LE, Skoog L, Sullivan S, Wingren S, Nordenskjold B (1993) S-phase fraction is a prognostic factor in stage I breast carcinoma. J Clin Oncol 11(9):1717–1722
Marechal R, Mackey JR, Lai R, Demetter P, Peeters M, Polus M, Cass CE, Salmon I, Deviere J, Van Laethem JL Deoxycitidine kinase is associated with prolonged survival after adjuvant gemcitabine for resected pancreatic adenocarcinoma. Cancer. doi: 10.1002/cncr.25303
Fujita H, Ohuchida K, Mizumoto K, Itaba S, Ito T, Nakata K, Yu J, Kayashima T, Souzaki R, Tajiri T, Manabe T, Ohtsuka T, Tanaka M (2010) Gene expression levels as predictive markers of outcome in pancreatic cancer after gemcitabine-based adjuvant chemotherapy. Neoplasia 12(10):807–817
Acknowledgments
We thank Lucas Bruurs for technical assistance. The work of the authors was supported by the SPINOZA grant from the Netherlands Organization for Scientific Research (NWO).
Conflict of Interest
RB and ST are employees of Agendia BV, the company that markets the MammaPrint prognosis test for breast cancer.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Geutjes, EJ., Tian, S., Roepman, P. et al. Deoxycytidine kinase is overexpressed in poor outcome breast cancer and determines responsiveness to nucleoside analogs. Breast Cancer Res Treat 131, 809–818 (2012). https://doi.org/10.1007/s10549-011-1477-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10549-011-1477-3