Abstract
Human-induced ecological and climatic changes have led to the decline and even local extinction of many formerly widely distributed temperate and cold-adapted species. Determining the exact causes of this decline remains difficult. Bryodemella tuberculata was a widely distributed orthopteran species before the mid-19th century. Since then, many European populations have suffered drastic declines and are now considered extinct or critically endangered. We used ecological niche modelling based on a large dataset of extant and extinct occurrence data to investigate whether poor climatic suitability in the periphery of its global range was a possible cause of the local extinction of the European populations of B. tuberculata. We also used population genetics based on the COI marker to estimate and compare the genetic diversity of extant populations. We found that Europe still provides highly suitable habitats close to the climatic optimum, contradicting the assumption of climate change as major driver of this decline. Instead, changes in land-cover and other anthropogenic modifications of the habitats at the local scale seem to be the major reasons for local extinctions. Genetic analysis suggests Central Asia as center of diversity with a stable population size, whereas the effective sizes of the remaining European populations are decreasing. We found European genetic lineages nested within Central Asian lineages, suggesting a Central Asian source distribution area. Our results suggest that the declining European populations represent relics of a formerly wider distribution, which was fragmented by changes in land-use. These relics are now threatened by limited connectivity and small effective population sizes. Specific conservation actions, such as the restoration of former or potential new habitats, and translocation of individuals from extant populations to these restored sites may help slow, stall, or even revert the extinction process.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Biodiversity decline, specifically insect decline, has been recognized as a major threat to global ecosystems (Dirzo et al. 2014; Conrad et al. 2006; Hallmann et al. 2017). Insects are key elements of most terrestrial trophic interactions, provide essential ecosystem services of global relevance, and generate significant economic and aesthetic benefits with cultural value to human society (Wagner 2019). At the same time, insect species and populations are threatened by a variety of factors, the most crucial being habitat loss, fragmentation and deterioration, followed by climate change (Franco et al. 2006; Potts et al. 2010; Vanbergen and the Insect Pollinators Initiative 2013).
Species are affected differently by environmental change depending on their ecological tolerance and other species-specific life history traits, determining the likelihood of decline and local extinction (Cahill et al. 2013). In most cases, the extinction of local populations depends on the combination of the geographic location of the population within the distribution range, the ecological tolerance of the species and shifts in habitat conditions (Lawton 1994; Cahill et al. 2013). Hutchinson's concept of the ecological niche (Hutchinson 1957) divides the multidimensional parameter space of habitat variation roughly into two regions: (1) the region in which the number of local births exceeds that of local deaths (source populations), close to the optimum niche of the species, and a source of emigration; and (2) the region in which the number of deaths exceeds that of local births (sink populations), typically far from the environmental optimum of the species, where populations are maintained by immigration from source populations (Holt 1996). Consequently, the proximity to the optimal environmental space (centroid) is expected to correspond to higher genetic diversity, growth rates and population stability (Pulliam 1988; Vanderwal et al. 2009). Known as center-periphery hypothesis, this concept has provided a baseline for many studies of population dynamics and genetic variability at species distribution limits.
We know of only few studies that have explored the center-periphery hypothesis regarding genetic variability in relation to environmental suitability or niche centrality—with contrasting results. Diniz-Filho et al. (2009), for example, related genetic diversity to average ecological niche model (ENM) suitability scores derived from multiple correlative niche modeling algorithms. Lira-Noriega and Manthey (2014) focused on testing linear regressions of genetic diversity measures (e.g., allelic richness, nucleotide diversity) in comparison with Euclidean distances between the environmental centroid of the suitable area estimated by ENM, and assumed to be the fundamental niche of the species. The correlations were tested on 40 species, including plants, birds, mammals, worms and insects. No clear relationship was recovered in any study, with the majority of cases showing negative correlations and a few cases displaying positive correlations (Lira‐Noriega and Manthey 2014). The mixed results might have been a product of the lack of a direct assessment of the species environmental centroid. Hence, a possible alternative to the approaches previously used is to characterize the ecological niche from a Grinnellian perspective—a set of environmental conditions that allow a species to maintain populations for long periods of time without immigration events (Peterson et al. 2011). This concept has been successfully applied to study the relationship between geographical distances of populations to the niche centroid and their abundances (Yañez‐Arenas et al. 2012; Urena-Aranda et al. 2015). However, no study has explored thus far the relationship of Grinnellian niche centroids with the geographic centroid and/or genetic diversity to explain patterns of population decline.
Due to well documented population declines within a huge part of its natural range, the speckled buzzing grasshopper Bryodemella tuberculata is well suited to investigate this relationship. This species occupies a wide global range across the northern Palearctic from Central Europe to eastern Asia (Bagachanova et al. 2011; Budrys et al. 2008; Budrys and Pakalniškis 2007; Fartmann et al. 2008; Reich 1991; Sergeev 1992; Srinivasan and Prabakar 2013). Up to the mid-20th century, B. tuberculata was still common in the European parts of its range, as documented by many specimens deposited in museum collections. It was frequently recorded from heath areas and river banks (Zacher 1919; Krauss 1883; Graber 1872). Over the course of the 20th century, many populations became extinct because their habitats (heath areas and unregulated river banks) were destroyed due to human demographic expansion, specifically a change in land-use due to intensified agriculture and shipping industry, and infrastructural developments (e.g. regulation of rivers; Laussmann et al. 2010; Bischoff 1997; Breunig and Thielmann 1992; Quinger 2014). Today, B. tuberculata has become a flagship species for the conservation of its threatened habitat types, mostly due to its large size for a grasshopper, its bright reddish hindwings and its characteristic buzzing flight noise. The most recent evaluation by the IUCN in 2016 (Zuna-Kratky et al. 2016) indicated that the populations in Denmark, Latvia, Poland and Switzerland are extinct (Berg et al. 2005; Budrys et al. 2008; Budrys and Pakalniškis 2007; Liana 2004; Maas et al. 2002; Monnerat et al. 2007), and the same probably applies to those in Italy (Massa et al. 2012). Furthermore, the relic populations of most other European countries have strongly declined despite frequent monitoring and legal protection (Binot-Hafke et al. 2011; Maas et al. 2011). In Central Europe, B. tuberculata is now restricted to small areas in Germany and Austria along the rivers Isar and Lech. In Northern Europe it is restricted to the island Øland in Sweden (Fig. 1; Bieringer and Weißmair 2017; Pfeuffer 2004; Voith et al. 2016). At the same time, the species remains common in North and Central Asia, occupying almost all suitable areas in the region (pers. observed by the authors).
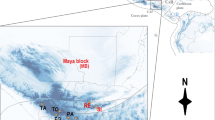
Distribution map based on all extinct and extant distribution records of Bryodemella tuberculata collected for this study from literature, museum vouchers, and private collections. Numbers 1 to 9 show the localities of the most recent records for countries (and regions of Germany) which B. tuberculata is now considered extinct. (Photograph of B. tuberculata, taken by Inci Livia Baez, Sylvenstein-Staumauer (Germany), August 2019)
This recent and steep decline of B. tuberculata populations offers an optimal setting to test the center-periphery hypothesis, and in connection with analysis of the land-use change potentially providing a framework to understand the factors leading to its population decline. Hence, we calculated ellipsoid envelope models based on a large dataset of present and past occurrences of the species to characterize the climatic niche of B. tuberculata and test the center-periphery hypothesis by comparing relationships between genetic diversity, climatic niche, and geographic centrality between extinct and extant populations. We further evaluated landscape change in the European area of the distribution and evaluate several extinct and extant locations for their habitat structure and suitability. We hypothesize that (1) changing climatic conditions over the last 100 years are not the reason for the decline of the species, but rather (2) changes in land-use and land-cover are responsible for the decline and local extinction. Assuming that genetic diversity and proximity to the niche centroid are related to fitness and survivability (Reed and Frankham 2003; Leimu et al. 2006), we specifically seek to understand whether the relationship between those factors could provide any insight into the susceptibility of populations (and possibly species) to decline and extinction. Based on this, we hypothesize that (3) the genetic diversity in the Asian core populations is higher than in the European relict populations.
Materials and methods
Distribution data
We assembled a set of 651 records of B. tuberculata obtained from fieldtrips, literature, and museum collections, complemented by data from the Global Biodiversity Information Facility (GBIF, www.gbif.org) and Naturbasen (https://www.naturbasen.dk/art?id=7888): based on fieldtrips, a total of 93 records were collected between 1873 and 2019 from Mongolia, Russia, Kazakhstan, Germany and China (see SI 1 for further information on collected material). Further 231 records were extracted from the literature (see SI 2 for consulted references) and 114 records were obtained from public databases (111 records from GBIF Global Biodiversity Information Center; GBIF.org (14 January 2020), GBIF Occurrence Download https://doi.org/10.15468/dl.hhsegk (extant dataset); GBIF.org (14 January 2020), GBIF Occurrence Download https://doi.org/10.15468/dl.pfaaea (extinct dataset); 13 records from Naturbasen (https://www.naturbasen.dk/art?id=7888)). Additionally, 213 records were obtained from museum specimens (see Supplementary Information SI1 for further information). We identified these museum specimens based on the key of Bey-Bienko (1930). Records with missing geographic coordinates were georeferenced using Google Earth (https://www.google.com/earth/) and checked for correctness following the data cleaning protocol by Cobos et al. (2018).
Due to the low dispersal capacity of B. tuberculata (Reinhardt et al. 2005) every single record was considered as a population. If several specimens from the same location were recorded, duplicates were removed. We considered any European population as extinct if it was listed as extinct in the relevant literature (Budrys and Pakalniškis 2007; Voith et al. 2016; Ruffo 2003; De Carlini 1889; Schmidt and Lilge 1997; Baur and Museum 2006; Bakker et al. 2015; Zuna-Kratky et al. 2017; Glowacinski and Nowacki 2006) or if no findings were documented after 1999 at the sampling location or within a 10 km buffer zone around the occurrence. All other populations from Europe and all known populations from Asia were considered extant. After data cleaning and trimming, 280 extant and 100 extinct locations were used to calculate the environmental distance to the centroid. All records used for modelling are supplied in Supplementary Information SI 3 and visualized in Fig. 1.
Environmental data
Environmental layers were obtained from Worldclim v. 1.4 (www.worldclim.org; Hijmans et al. 2005) in 2.5 arc-min resolution. WorldClim is based on interpolations of weather station data (i.e., monthly precipitation and minimum and maximum temperatures) over the period 1960–1990. From the 19 variables available, we excluded four (mean temperature of wettest quarter, mean temperature of driest quarter, precipitation of warmest quarter, precipitation of coldest quarter) a priori due to known spatial artifacts between adjacent grid cells (Escobar et al. 2014; Campbell et al. 2015). Seeking to avoid bias regarding the combination of variables used to characterize the species niche centrality, we tested three distinct environmental sets: ‘set 1’ included all 15 variables; ‘set 2’ included only temperature variables (i.e., annual mean temperature; mean diurnal range; isothermality; temperature seasonality; max. temperature of warmest month; min. temperature of coldest month; temperature annual range; mean temperature of warmest quarter and mean temperature of coldest quarter); and ‘set 3’ included only precipitation variables (i.e., annual precipitation; precipitation of wettest month; precipitation of driest month; precipitation seasonality; precipitation of wettest quarter; precipitation of driest quarter). To avoid overfitting, overly dimensional environmental space and collinearity among variables, we performed principal component analysis (PCA) using the function kuenm_rpca in the package ‘kuenm’ (Cobos et al. 2019b) in R 3.6.3 (R Core team 2020). For each set, we retained as many components as necessary to explain > 95% of the total variance in the dataset for model calibration. As a result, set 1 was reduced to five principal components (PCs) summarizing the 15 bioclimatic variables, while set 2 and 3 were reduced to three PCs, summarizing temperature and precipitation variables (see SI 6).
Ellipsoid models
To characterize the environmental niche of B. tuberculata we created an ellipsoid envelope model representing the niche shape assumed when multiple dimensions are considered (Jiménez et al. 2019) using the ‘ellipsenm’ package (Cobos et al. 2019a). Based on these models, we calculated the niche centroid for all collected extant records of the species and extracted the Mahalanobis distances between grid cells representing local environments of study populations and environmental conditions of the optimum (ellipsoid centroid). Models were calibrated using the 95% pairwise confidence region for the ellipsoid and were evaluated as candidate models using the function ‘ellipsoid_calibration’ of the ‘ellipsenm’ R package (Cobos et al. 2019a). Two distinct methods were employed to construct ellipsoid models: (1) ‘covmat’, which creates ellipsoids based on the centroid and a matrix of co-variances of the variables and (2) ‘mve1’, which generates an ellipsoid that reduces the volume contained in it without losing the data contained (i.e., minimum volume ellipsoid; Van Aelst and Rousseeuw 2009). Best model selection was based on statistical significance (partial ROC; Peterson et al. 2008); the proportion of testing data known to be in suitable areas and prediction of unsuitable areas was based on omission rates (E = 5%; Anderson et al. 2003) and prevalence. To calculate the partial ROC metric, we used 500 bootstrap iterations with 50% of testing data to be used in each bootstrapped process with 5% of testing data error in the data due to uncertainty. The calibration area (i.e., region accessible to the species; Barve et al. 2011) included Eurasia, except Southeast Asia, where the species is not found.
Final parameters were selected based on best evaluated models and used to create the final models using 10 replicates with bootstrapped subsamples of 75% of the data using the function ‘ellipsoid_model’ available in the ‘ellipsenm’ R package (Cobos et al. 2019a). The replicates were produced by excluding one occurrence record at a time. The confidence level of a pairwise confidence region for the ellipsoid was 99%, with 1% of the occurrence data considered as potential environmental outliers, not included in the ellipsoid envelope model for the species’ ecological niche.
Environmental and geographic distances
The environmental distance of each record (= population) to the ellipsoid centroid (= ecological optimum) was calculated using the Mahalanobis distance metric, which is considered wel- suited for non-spherically symmetric distributions and deals robustly with variables with different scales, such as the bioclimatic variables we used for characterizing niche space (De Maesschalck et al. 2000; Hijmans 2020). We generated histograms showing the Mahalanobis distance (D2) of all populations to the centroid.
To estimate the geographical distance of each population to the centroid, we created a minimum convex polygon based on all known records of B. tuberculata and then calculated the distances of the centroid to its edges. For that, we used the R packages ‘adehabitatHR’ (Calenge 2006); ‘geogrid’ (Bailey 2018); ‘sp’ (Bivand et al. 2008; Pebesma and Bivand 2005); ‘rgeos’ (Bivand and Rundel 2019); ‘rgdal’ (Bivand et al. 2015); ‘raster’ (Hijmans 2020) in R v. 3.6.3. (R Core Team 2020) under R Studio v. 1.1.463 (R Studio 2020). Histograms for both distance measures (i.e., environmental and geographic) were created with the R packages ‘ggplot2’ (Wickham 2016); ‘gridExtra’ (Auguie 2017); ‘plyr’ (Wickham 2011); ‘cowplot’ (Wilke 2019) and ‘grid’ (R Core Team 2020).
To test whether the distribution of Mahalanobis distances of extinct populations to the centroid are significantly distinct from the distances recovered for extant populations, we performed a non-parametric Mood’s Median test, which tests if two or more independent samples originate from populations with the same median. We used this to compare the shape and scale of distributions for all groups, based on the hypothesis that the Mahalanobis distances of extant populations are lower compared to those of extinct populations (p-value > 0.05). Analyses were performed in R. Distribution maps and Mahalanobis plots were created in QGIS v. 3.10.3 (QGIS Development Team 2020).
Land-cover change
Furthermore, we used the HILDA dataset v. 2 (HIstoric Land Dynamics Assessment 2—gross land-cover changes; Fuchs et al. 2013, 2014, 2015; http://www.geo-informatie.nl/fuchs003/#) to display landscape change through time from 1900 to 2000, the main timeframe of local extinctions in Europe. Based on this, we calculated the percent land-cover change for forest, grassland, settlement and cropland area from 1900 to 2000 using the QGis plugin LecoS—Landscape Ecology Statistics v. 3.0.0 (Jung 2016).
Molecular analyses
For this study, we tested several genetic markers. Most gene fragments were highly conserved and did not show any variability; genomic data is currently not available to us and remains challenging because of the extremely large genome sizes in Orthoptera and specifically Oedipodinae (Husemann et al. 2021). In turn, DNA barcode data is known to provide meaningful estimates of genetic diversity, and additional genetic data of this marker was available from other studies in BOLD and GenBank. Hence, we decided to use a fragment of the Cytochrome Oxidase 1 for the genetic comparison of the European and Asian populations. We compiled COI sequences of 34 specimens of B. tuberculata from populations across its global range (Table 1). DNA was extracted from muscle tissue of a hind femur of each specimen with a high salt extraction protocol (Paxton et al. 1996). We amplified the barcoding fragment of the mitochondrial Cytochrome C Oxidase subunit I (COI) gene using the primers provided by Pan et al. (2006). For PCR reactions we used DreamTaq DNA polymerase (Thermo Fischer Scientific, Schwerte, Germany) following the manufacturer’s protocol. Cycling conditions are shown in SI 4. PCR products were visualized using 1 µl of EZ-Vision Two DNA Dye (Amresco, Darmstadt, Germany); samples were run on 1% agarose gels. We purified successfully amplified samples using the Thermo Fisher protocol for PCR product clean-up prior sequencing (Werle et al. 1994). The samples were sequenced at Macrogen Europe (Macrogen, Amsterdam, Netherlands) using the EZ-Seq service. Further, sequences from earlier projects of MH and OH (Husemann et al. 2012; Hawlitschek et al. 2017) were obtained from BOLD (see Table 1).
Analysis of genetic diversity
The MUSCLE algorithm (Edgar 2004) as implemented in Geneious v. 10.0.9 (http://www.geneious.com, Kearse et al. 2012) was used to edit, trim and align the sequences, resulting in three datasets for molecular analyses. The first dataset included all 34 sequences (total alignment length 609 bp). The second dataset only comprised samples from Europe (Germany N = 15; Sweden N = 1; Lithuania N = 1) with a total alignment length of 609 bp. A third dataset was generated comprising the samples from Central Asia (Russia N = 9; Mongolia N = 5; China N = 3; Korea N = 1) with a total alignment length of 609 bp. Geneious was used to check for internal stop codons (none were present). All sequences were blasted in NCBI GenBank and BOLD systems v. 3 (Ratnasingham and Hebert 2007) to exclude rough sequencing mistakes. ModelFinder (Kalyaanamoorthy et al. 2017) implemented in IQTree (Nguyen et al. 2015) was used to statistically select the best model of nucleotide substitution for the datasets using the Akaike information criterion (AIC) (Akaike 1973). HKY was selected as best model for all datasets. Pairwise genetic distance between the Asian (samples from Russia, Mongolia, China and Korea) and European (samples from Germany, Sweden and Lithuania) populations was calculated using Arlequin v. 3.5.2.2 (Excoffier et al. 2005). We used PopArt (Leigh and Bryant 2015) to construct a TCS haplotype network (Clement et al. 2000) of the full dataset without outgroup and with seven geographical traits (Germany, Sweden, Lithuania, Russia, Mongolia, China. Korea).
To test for demographic stability we conducted Bayesian Skyline analyses using BEAST v. 2.5.1, based on the reduced datasets of Europe and Central Asia. HKY was used as substitution model. A relaxed log-normal clock using the substitution rate provided by Papadopoulou et al. (2010) was implemented as clock model to calculate a Coalescent Bayesian Skyline plot; the analysis was run for 100 million generations sampling every 1,000 generations. We ran BEAST v. 2.5.1 with automatic thread pool size and BEAGLE library (Ayres et al. 2012). Convergence was assessed with TRACER v. 1.7.1 to avoid insufficient sampling. TRACER was also used to calculate the Bayesian Skyline reconstruction (Bouckaert et al. 2014) discarding 10% of samples as burn-in.
Results
Species distribution
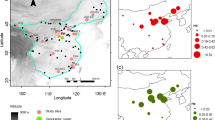
We assembled 280 extant distribution records from Austria (28), China (13), Finland (1), Germany (25), Kazakhstan (19), North Korea (2), South Korea (22), Lithuania (1), Mongolia (19), Romania (1), Russia (138), Sweden (8) and Ukraine (3) and 100 records of extinct populations from Austria (11), Denmark (12), Estonia (1), Germany (34), Italy (2), Latvia (5), Poland (31) and Switzerland (5). Location files including coordinate data are provided as supplementary information SI 3. Figure 1 shows all records we used for modelling. The northernmost location was found in Siberia (Russia) at 67°N; the southernmost location was in Tibet (China) at 28°N. Most of the occurrences were recorded between 57°N and 45°N. The longitudinal extension stretches across most of Eurasia from 152°E in Russia to 12°E in Germany over a range of around 7500 km.
Ellipsoid model
The best fitting method to construct the climatic ellipsoids was ‘covmat’; mean AUC, p-value of partial ROC and omission rates were significantly better than random expectations (p-value < 0.05; Table 2). The complete report of ellipsoid characteristics (e.g., centroid, covariance matrix, semi-axes length, etc.) can be found in SI 5.
Environmental and geographic distances
The geographic prediction of Mahalanobis distance (D2) in the calibration area is shown in Fig. 2 and SI 6 and SI 7. The comparison of D2 on all environmental sets shows all extinct populations, except for one in northern Austria (Ried im Innkreis: 48.222461°N 13.484835°E) to be as close to the climatic centroid as extant populations (Table 3). The environmental set of all bioclimatic variables yielded a D2 ratio similar between the investigated locations. Statistical results of Mood’s Median test show no significant difference between the distances of extinct and extant populations to the climatic niche centroid (Mood’s Median test: Set 1: p-value < 0.001; Fig. 3). Moreover, the average distances to the centroid were found to be higher for the extinct and extant European locations (distance: 38.00–66.28, Ø = 59.53 (± 6.08)) than for Asian populations, which were geographically closer to the centroid (distance: 3.86–80.42, Ø = 30.39 (± 18.27); overall p-value < 0.001; Fig. 3).
Histograms showing the distribution of Mahalanobis distances (D2) of extinct and extant populations. The centroid of the climatic niche was estimated using Set 1 including all bioclimatic variables. p-values included in graphs indicate results of Moods’ Median test, which tests the differences between the distribution of extinct and extant distances of the different parameters. Moreover, the figure shows the geographic distance from each location to the center of the distribution. Values associated with extinct populations are shown in red, while extinct are colored blue. The black arrow in the histogram “All bioclimatic variables (Set 1)” shows the outlier point in Austria close to Ried im Innkreis (D2 = 22.306), the grey arrow shows the outlierclose to the German-Austrian border in the Berchtesgaden national park (D2 = 17.142)
Land-cover change
Using the HILDA dataset we were able to track changes in land-cover in Central Europe from 1900 to 2000 (SI 8): a massive change from cropland (1900: 33.9% to 2000: 29.3%) and grassland (1900: 34% to 2000: 27.5%) to more settlement area (1900: 2.2% to 2000: 4.2%) and forests (1900: 25% to 2000: 34.3%) can be observed.
Analysis of genetic diversity
We included 34 COI sequences of B. tuberculata in the analyses. These represented a total of 28 haplotypes with a haplotype diversity of 0.978 (± 0.015) and a nucleotide diversity of 0.031 (± 0.015). The pairwise distance between central (Asia) and peripheral (Europe) populations was moderate, but significant (ΦST of 0.104; p-value < 0.001).
The haplotype network (Fig. 4) shows a close relationship of all haplotypes with few evolutionary steps between them. Two main groups are recognizable, albeit with only few separating mutations. These two groups do not reflect a split between European and Asian samples, as European samples are grouping with sequences from Central Asia. The samples from China are more closely related to those from Korea and to most of the Mongolian samples representing the first group; the other group is composed of the Russian, Mongolian and European samples (Germany, Sweden, Lithuania). Furthermore, the Russian, German, Lithuanian and one Mongolian sample share the same haplotype.
Bayesian Skyline plots tracing female effective population size through time were calculated for European and Central Asian specimens separately (Fig. 5). The analysis showed a clear trend of population decline during the last 1000 years for the European populations. The same analysis performed for the Central Asian samples yielded a 10 times higher estimate of female effective population size and demographic stability through time.
Discussion
In this study, we investigated potential reasons for the decline of the European populations of the formerly widely distributed band-winged grasshopper B. tuberculata using ecological niche modelling in combination with a basic population genetic assessment. Specifically, we aimed to test if (1) changing climatic conditions over the last 100 years, or (2) changes in land-use and land-cover are responsible for the decline and local extinction. For this we assessed the habitat requirements and the decline of the species and modelled the environmental niche. Furthermore, we tested if (3) the genetic diversity in the Asian core populations is higher than in the European relic populations, which may support the center-periphery hypothesis. We discuss the results in regards of conservation management of the species.
Climate models reveal suitable conditions in extinct locations
We used an ellipsoid envelope approach to estimate the climatic niche of B. tuberculata and measure the proximity of its populations to their optimum. All sets recovered low distances of the extinct populations to the climatic niche centroid, and none recovered any difference between extinct and extant populations (Figs. 2, 3; SI 6–7). Set 1 offered higher variation in distances to the centroid (range: 0.417–32.611), possibly due to the higher variation and multi-dimensionality of the principal components included (i.e., 15 variables).
While most extinct populations were located in areas with lower Mahalanobis distance (D2) to the centroid (i.e., high suitability), two extinct locations with higher values were recorded (D2 = 22.306 and 17.142; Fig. 2, Table 3). These locations represent a population close to the German-Austrian border in the Berchtesgaden national park (D2 = 17.142) and a population in Austria close to Ried im Innkreis (D2 = 22.306). The higher D2 at these two locations could have several reasons, e.g. imprecise locality data, errors in georeferencing, individuals dispersed from neighboring locations (e.g. migrating individuals; Barton and Hewitt 1982), artefacts of the bioclimatic layers used (e.g. Marchi et al. 2019; Poggio et al. 2018), or even just a high tolerance regarding abiotic conditions (e.g. Preston and Johnson 2020). The last reason is supported by data from Asia with similar D2 range. Since our models predict areas situated in the distribution gap between the European and Asian populations as climatically suitable, the destruction of natural habitats is a likely reason for the loss of stepping stones and for the extinction of any connecting populations in the area.
Habitat requirements and historical decline of B. tuberculata
Bryodemella tuberculata is a species of open landscapes, preferring habitats with sparse vegetation and open ground. According to our observations, all Russian collection sites in the Altay-Sayan Mountains, the Kulunda Plain (the south-eastern part of West Siberian Plain between the Irtysh and Ob Rivers) and within the south-eastern Ural Mountains (pers. obs. MGS; for further details, see SI 9) were mainly composed of dry meadows and steppes with sparse vegetation (~ 10–30 cm height) and spots of open ground (generally, the vegetation cover was less than 80%). The ground is mostly sandy to rocky, with smaller pebbles. In most cases occurrence sites were close to water systems. The species enters transformed ecosystems, such as the steppe variants with moderate livestock grazing and the dry hayfields. This type of habitat was found at almost all visited collection sites in Central Asia and Europe (for habitat images see Supplementary Information SI 10). In Mongolia, the species is found in low abundance, but in almost all habitats with open spaces and sparse vegetation, mainly covered by shorter plant species up to 30 cm and at the edges to forests (pers. observ., LSD 2015–2019; Dey et al. in press; Supplementary Information SI 10). Pastures for grazing represent the major agricultural use in the country, hence, the landscape is characterized by bare steppe habitats with low vegetation providing good conditions for B. tuberculata.
All extant populations of Central Europe (Southern Germany, Austria) inhabit alpine river valleys with unmodified or little modified flood regimes; specifically zones of alluvial gravel beds with sparse or without any vegetation. Occasionally, individuals may be observed in adjacent structures similar to those described for the Asian populations (Landmann 2017). Since the middle of the 19th century, populations of B. tuberculata in Europe have been declining (e.g. Reich 1991; Zuna-Kratky et al. 2017). Despite the availability of a large database, we are unable to make any statements about the exact timing of extinction of B. tuberculata at any particular location, but we provide the last recorded findings for several regions (Fig. 1; Supplementary Information SI 11). Altogether, our data suggest that most Central European extinction events took place between the 1920’s and 1960’s. We therefore suggest that each local extinction event in Central Europe may be closely connected with local landscape changes. Historically, at least in the mountainous regions, the naturally occurring annual floods following snowmelt removed most vegetation growing in the riparian habitats. Anthropogenic reductions of the flood volume, mostly due to the construction of hydroelectric dams, led to the establishment of shrub and tree vegetation in these habitats and a decline of the areas suitable for B. tuberculata (Reich et al. 2008; Juszczyk et al. 2020). A variety of restoration measures have been put in place over the last decades, including a reduced deduction of water for hydroelectric energy generation, mechanical removal of vegetation, and education of the public; the mid- and long-term efficiency of these measures remains to be demonstrated (Juszczyk et al. 2020). A different type of change to the habitat structure was recorded in Denmark. There, most heathlands in the vicinity of Abild (an extinct location) have been converted to agricultural areas or were overgrown by shrubs (pers. observ. LSD; SI 10). Although the agricultural land-use decreased from 74% in 1915 to 61% in 2015, afforestation and urban expansion still led to a decline of heath areas previously typical for many parts of Denmark (Pedersen and Møllenberg 2017). The species has been listed as extinct in Denmark since the 1950s. Similarly, in Northern Germany, restoration of some old heath landscapes has been performed. Based on literature and museum surveys, we were able to find several records from the Lüneburger Heide heathland areas, for which the gradual conversion to arable land and forests at the end of the 19th century is historically documented (Koopmann 2000). This change in land-use was mainly triggered by a decrease in the local production of wool and honey due to external competitors, making the maintenance of large pastures uneconomic (Naturpark-Lueneburger-Heide.de; Koopmann 2000). During the early 20th century, the first areas were restored to the original heath systems. Nowadays, the Lüneburger Heide is one of the largest heath areas in Northern and Central Germany. Although most heath areas in Central Europe declined through time, some of these cultural landscapes are still intact. Many of them are now under military use and kept open by military activity, which practically equals to legal protection of the landscape. As a result, B. tuberculata was rediscovered on 03 August 2008 in the Pabradė military training area in Lithuania, where it had been assessed as possibly extinct until 2008 (Budrys et al. 2008; Budrys and Pakalniškis 2007). This was the first record since the first half of the 20th century (Budrys and Pakalniškis 2007); yet, the status of the population remains unknown and it will have to be monitored to check its population establishment. In Northern Europe the population on Øland (Sweden) shows similar habitat preferences for barren land. The implementation of specific conservation measures appears to keep the population relatively stable.
Bryodemella tuberculata needs larger areas with open habitat, often with frequent natural disturbance. Anthropogenic changes of the European landscape in the last centuries (Plieninger et al. 2016; Hersperger and Bürgi 2009; Antrop 2004) likely led to the current patchy distribution of the species and will probably cause the extinction of further populations in the European range. The time series of land-cover plots (Supplementary Information SI 8) supports land-use change as a driver of the decline of B. tuberculata as it displays a rapid change from grassland and agricultural areas to settlement and reforestation in the middle of the 20th century and still ongoing. This change from open areas to more closed habitat types, and an increase of human pressure due to high-intensity land-use, may have led to the extinction of B. tuberculata at many Central European locations. Even though some habitats of extinct populations have now been restored, they are highly fragmented, and the dispersal capabilities of B. tuberculata appear insufficient to allow for quick colonization of these habitats without support. Similar scenarios of decline have been described for many other European species, e.g., carabid beetles with different ecological preferences in Belgium, Denmark and the Netherlands (Kotze and O’Hara 2003); common and widespread butterflies in the Netherlands (Van Dyck et al. 2009); or the ground nesting Black Grouse populations in Lower Saxony, Germany (Ludwig et al. 2009). In turn, the availability of large stretches of natural habitats suitable for B. tuberculata currently remains much higher in the Asian range. However, vast areas of habitat are also being destroyed in Russia, especially in the European part of the country, due to urbanization, agriculture and mining (Smelansky and Tishkov 2012). This process is much slower due to lower human activity in these areas and simply a larger area, but can also be expected to fragment populations and impact genetic diversity in the near future.
Distribution of genetic diversity and historical demography
While our genetic data is limited, also due to the rareness of the species, it suggests that the majority of genetic diversity is found in Central Asia. The European populations show lower diversity and are nested within the Asian populations. Hence, based on our COI data, European populations only represent a subset of the species’ genetic diversity, as expected for relic populations, which may have gone through a population bottleneck (e.g. Chen et al. 2016; Gaublomme et al. 2013; Hájková et al. 2007; Lucchini et al. 2004). We did not find any evidence supporting the status of these populations as distinct genetic lineages. This may suggest a previously more continuous, potentially panmictic distribution across much of Europe and Asia and a relatively recent fragmentation and decline in Central Europe, as also suggested by our historical distribution data.
The genetic patterns do not support the center-periphery hypothesis, i.e., low genetic variability in areas of low environmental suitability, as low genetic variability was also detected in areas of high environmental suitability. However, due to strong contemporary restrictions on migration, gene flow is likely completely interrupted and the European populations are likely under a drift regime, rendering them prone to extinction. On the other hand, Bayesian Skyline plots provide some evidence that supports the center-periphery hypothesis driven by geographic distance. Our results show that female effective population size is larger in Central Asia and also has been more stable in the last 10 ka (Fig. 5), while the European populations have smaller estimated population sizes and are in decline. The results we obtained match other taxa with similar distributions, e.g., the leaf-beetle Cheilotoma musciformis, which has some relic populations in Poland. These populations are declining, probably because habitat destruction caused a disjunction from the main distribution, resulting in an observed genetic bottleneck (Kajtoch et al. 2016). Such patterns of loss of genetic diversity associated with habitat destruction, which have led to the disjunction of populations, can also be seen in the decline of the Danish population of the otter Lutra lutra (Pertoldi et al. 2001), among others (e.g. Cremene et al. 2005; Kotze and O’Hara 2003).
Insufficient genetic samples and lack of more fine-scaled genome-wide markers inhibited the direct assessment of a relationship between genetic diversity and niche centrality. Nevertheless, our results support the findings of Lira‐Noriega and Manthey (2014), who equally failed to detect any clear relationship between genetic diversity, and geographic and environmental distance to the geographic centroid. However, they did find a strong relationship between genetic diversity and climatic niche centrality. In some taxa, Lira‐Noriega and Manthey (2014) describe a negative tendency for the relationship of genetic diversity and geographical distance to the source population, reflecting environmental impact on the population dynamics, rather than a fundamental ecological relationship. In case of B. tuberculata, we suggest a similar effect of habitat destruction in the periphery of the distribution supported by decreasing female effective population size in the European populations (geographically distant from the main distribution; source) in contrast to the stable demography in the Asian populations (geographically close to the source). A more detailed study of other habitat characteristics, especially land-use patterns, may shed light on the reasons for the local decline and could help to manage the conservation activities to save the remaining European populations from decline, which may also facilitate the recolonization of former locations.
Aspects of conservation management
Local conservation efforts for species with isolated and declining populations may often be seen as unsustainable, because these populations may be threatened by the effects of climate change, which cannot be counteracted by localized efforts (e.g. Sinervo et al. 2010, Malcom et al. 2006). Our results show that extinct or threatened European populations of B. tuberculata are in areas of optimal climatic conditions for this species, whereas some Asian populations appear to be thriving also in areas of poor climatic suitability. This suggests that B. tuberculata is most likely resilient to the purely climatic effects of global warming in the Central European part of its range. If our results hold true, this means that threatened populations may survive the next decades of changing climate as long as their habitats are locally protected, and that these local efforts can be considered sustainable. This includes not only threatened extant populations, but also means that reintroduction efforts into the localities of extinct populations may be a promising option as long as the habitat structures necessary for B. tuberculata have been restored and long-term management measures are in place. We suggest that monitoring of extant and potentially translocated populations should be accompanied by genetic studies using deeper population-level sequencing methods, such as microsatellites or RAD sequencing. Once a high-quality whole genome becomes available, genome resequencing may become a viable option.
The speckled buzzing grasshopper is just one species of an assembly of taxa sharing a habitat that is highly threatened by anthropogenic land-cover change. We believe that our study reinforces that status of B. tuberculata as a flagship and umbrella species for this organism community, and we hope that our results help making a case for increased efforts to their protection.
Data availability
All genetic data will be available in GenBank. All distribution data will be available in the supplementary section.
Code availability
The R code for bioclimatic classification of the region will be available on Github: https://github.com/msimoes123/NicheCentrality_Bryodemella; https://github.com/msimoes123/NicheCentrality_Bryodemella.
References
Akaike H (1973) Maximum likelihood identification of Gaussian autoregressive moving average models. Biometrika 60(2):255–265
Anderson RP, Lew D, Peterson AT (2003) Evaluating predictive models of species’ distributions: criteria for selecting optimal models. Ecol Model 162:211–232
Antrop M (2004) Landscape change and the urbanization process in Europe. Landsc Urban Plan 67:9–26
Auguie B (2017) gridExtra: miscellaneous functions for “Grid” graphics. Version R package version 2.3. https://CRAN.R-project.org/package=gridExtra. Accessed 9 Nov 2020
Ayres DL, Darling A, Zwickl DJ et al (2012) BEAGLE: an application programming interface and high-performance computing library for statistical phylogenetics. Syst Biol 61:170–173
Bagachanova AK, Vinokurov NN, Evdokarova TG et al (2011) Taxonomic diversity of insects from the relic steppes of the Mid-Lena River Valley (Central Yakutia). Arid Ecosyst 1:38
Bailey J (2018) geogrid: turn geospatial polygons into regular or hexagonal grids. Version R package version 0.1.1. https://CRAN.R-project.org/package=geogrid. Accessed 9 Nov 2020
Bakker W, Bouwman J, Brekelmans F et al (2015) De Nederlandse sprinkhanen en krekels (Orthoptera). Nederlandse Entomologische Vereniging, pp 248
Barton NH, Hewitt GM (1982) A measurement of dispersal in the grasshopper Podisma pedestris (Orthoptera: Acrididae). Heredity 48:237–249
Barve N, Barve V, Jiménez-Valverde A, Lira-Noriega A, Maher SP, Peterson AT, Soberón J, Villalobos F (2011) The crucial role of the accessible area in ecological niche modeling and species distribution modeling. Ecol Model 222(11):1810–1819
Baur B, Museum BN (2006) Die Heuschrecken der Schweiz. Haupt, Bern
Berg H-M, Bieringer G, Zechner L (2005) Rote liste der heuschrecken (Orthoptera). Österreichs. In: Zulka K-P (Red.) Rote Listen gefährdeter Tiere Österreichs. Grüne Reihe des Lebensministeriums, Band, vol 14, issue no 1, Böhlau Verlag, Wien pp 167–209
Bey-Bienko GY (1930) A monograph of the genus Bryodema Fieb. (Orthoptera, Acrididae) and its nearest allies. Ann Zool Mus Acad Leningr 31:71–127
Bieringer G, Weißmair W (2017) Gefährdung und Schutz der Heuschrecken in Österreich. Denisia 39:161–180
Binot-Hafke M, Balzer S, Becker N et al (2011) Rote liste gefährdeter tiere, pflanzen und pilze Deutschlands. Wirbellose Tiere (part 1). Bundesamt für Naturschutz, pp 716
Bischoff I (1997) Die Wahner Heide, ein Naturschutzgebiet und Truppenübungsplatz in Nordrhein-Westfalen. Brandenburgische Umwelt Berichte (BUB) 1:39–47
Bivand R, Rundel C (2019) Rgeos: interface to geometry engine—open source (‘GEOS’). Version R package version 0.5-2URL. https://CRAN.R-project.org/package=rgeos. Accessed 9 Nov 2020
Bivand RS, Pebesma EJ, Gomez-Rubio V, Pebesma EJ (2008) Applied spatial data analysis with R. Springer, Berlin
Bivand R, Keitt T, Rowlingson B et al (2015) Package ‘rgdal.’ Bindings for the Geospatial Data Abstraction Library. Available online https://cran.r-project org/web/packages/rgdal/index html. Accessed 21 Sept 2020
Bouckaert R, Heled J, Kühnert D et al (2014) BEAST 2: a software platform for Bayesian evolutionary analysis. PLoS Comput Biol. https://doi.org/10.1371/journal.pcbi.1003537
Breunig T, Thielmann G (1992) Binnendünen und Sandrasen. Landesanstalt für Umweltschutz Umweltschutz Baden-Württemberg (Hrsg.): Biotope in Baden-Württemberg 1. Karlsruhe, pp 36
Budrys E, Pakalniškis S (2007) The Orthoptera (insecta) of Lithuania. Acta Zool Litu 17:105–115. https://doi.org/10.1080/13921657.2007.10512821
Budrys E, Bačianskas V, Budrienė A et al (2008) Distribution of four species of Oedipodinae grasshoppers in Lithuania (Orthoptera: Acrididae). New Rare Lith Insect Species Rec Descr 20:14–19
Cahill AE, Aiello-Lammens ME, Fisher-Reid MC et al (2013) How does climate change cause extinction? Proc R Soc B 280:20121890
Calenge C (2006) The package adehabitat for the R software: tool for the analysis of space and habitat use by animals. Ecol Model 197:516–519
Campbell LP, Luther C, Moo-Llanes D et al (2015) Climate change influences on global distributions of dengue and chikungunya virus vectors. Philos Trans R Soc B 370:20140135
Chen N, Cosgrove EJ, Bowman R et al (2016) Genomic consequences of population decline in the endangered Florida scrub-jay. Curr Biol 26:2974–2979
Clement M, Posada D, Crandall KA (2000) TCS: a computer program to estimate gene genealogies. Mol Ecol 9:1657–1659
Cobos ME, Jiménez L, Nuñez-Penichet C et al (2018) Sample data and training modules for cleaning biodiversity information. Biodiv Inf 13:49–50
Cobos ME, Osorio-Olvera L, Soberon J et al (2019) Ellipsenm: an R package for ecological niche’s characterization using ellipsoids. Version R package version 0.3.4. https://github.com/marlonecobos/ellipsenm. Accessed 27 Oct 2020
Cobos ME, Peterson AT, Barve N, Osorio-Olvera L (2019b) kuenm: an R package for detailed development of ecological niche models using Maxent. PeerJ 7:e6281
Conrad KF, Warren MS, Fox R et al (2006) Rapid declines of common, widespread British moths provide evidence of an insect biodiversity crisis. Biol Conserv 132:279–291
Cremene C, Groza G, Rakosy L et al (2005) Alterations of steppe-like grasslands in Eastern Europe: a threat to regional biodiversity hotspots. Conserv Biol 19:1606–1618
De Carlini A (1889) Artropodi di Valtellina (Rincoti, Ortotteri, Aracnidi). Bull Soc Ent Ital 21:9–19
De Maesschalck R, Jouan-Rimbaud D, Massart DL (2000) The mahalanobis distance. Chemom Intell Lab Syst 50:1–18
Diniz-Filho JAF, Nabout JC, Bini LM et al (2009) Niche modelling and landscape genetics of Caryocar brasiliense (“Pequi” tree: Caryocaraceae) in Brazilian Cerrado: an integrative approach for evaluating central–peripheral population patterns. Tree Genet Genomes 5:617–627
Dirzo R, Young HS, Galetti M et al (2014) Defaunation in the Anthropocene. Science 345:401–406
Edgar RC (2004) MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res 32:1792–1797
Escobar LE, Lira-Noriega A, Medina-Vogel G, Peterson AT (2014) Potential for spread of the white-nose fungus (Pseudogymnoascus destructans) in the Americas: use of Maxent and NicheA to assure strict model transference. Geospat Health 9:221–229
Excoffier L, Laval G, Schneider S (2005) Arlequin (version 3.0): an integrated software package for population genetics data analysis. Evol Bioinform 1:117693430500100000
Fartmann T, Behrens M, Loritz H (2008) Orthopteran communities in the conifer-broadleaved woodland zone of the Russian Far East. Eur J Entomol 105:673–680
Franco AM, Hill JK, Kitschke C et al (2006) Impacts of climate warming and habitat loss on extinctions at species’ low-latitude range boundaries. Glob Change Biol 12:1545–1553
Fuchs R, Herold M, Verburg PH et al (2013) A high-resolution and harmonized model approach for reconstructing and analysing historic land changes in Europe. Biogeosciences 10(3):1543–1559. https://doi.org/10.5194/bg-10-1543-2013
Fuchs R, Herold M, Verburg PH et al (2014) Gross changes in reconstructions of historic land cover/use for Europe between 1900–2010. Glob Change Biol. https://doi.org/10.1111/gcb.12714
Fuchs R, Verburg PH, Clevers JGPW et al (2015) The potential of old maps and encyclopaedias for reconstructing historic continental land cover/use change. Appl Geogr 59:43–55. https://doi.org/10.1016/j.apgeog.2015.02.013
Gaublomme E, Maebe K, Van Doninck K et al (2013) Loss of genetic diversity and increased genetic structuring in response to forest area reduction in a ground dwelling insect: a case study of the flightless carabid beetle Carabus problematicus (C oleoptera, C arabidae). Insect Conserv Divers 6:473–482
Glowacinski Z, Nowacki J (2006) Polish red data book of animals: invertebrates. Website: http://www.iop.krakow.pl/pckz/default. Accessed 13 July 2020
Graber V (1872) Bemerkungen über die“ Gehör-und Stimmorgane“ der Heuschrecken und Cikaden. SB. Akad. Wissensch. Wien 66(Abth. I):205–213
Hájková P, Pertoldi C, Zemanová B et al (2007) Genetic structure and evidence for recent population decline in Eurasian otter populations in the Czech and Slovak Republics: implications for conservation. J Zool 272:1–9
Hallmann CA, Sorg M, Jongejans E et al (2017) More than 75 percent decline over 27 years in total flying insect biomass in protected areas. PLoS ONE 12:e0185809
Hawlitschek O, Morinière J, Lehmann GUC et al (2017) DNA barcoding of crickets, katydids and grasshoppers (Orthoptera) from Central Europe with focus on Austria, Germany and Switzerland. Mol Ecol Resour 17:1037–1053
Hersperger AM, Bürgi M (2009) Going beyond landscape change description: quantifying the importance of driving forces of landscape change in a Central Europe case study. Land Use Policy 26:640–648
Hijmans RJ (2020) raster: geographic data analysis and modeling. Version R package version 3.0-10/r3416. https://R-Forge.R-project.org/projects/raster/. Accessed 31 July 2020
Hijmans RJ, Cameron SE, Parra JL et al (2005) Very high resolution interpolated climate surfaces for global land areas. Int J Climatol 25:1965–1978. https://doi.org/10.1002/joc.1276
Holt RD (1996) Adaptive evolution in source-sink environments: direct and indirect effects of density-dependence on niche evolution. Oikos 75:182–192
Husemann M, Namkung S, Habel JC et al (2012) Phylogenetic analyses of band-winged grasshoppers (Orthoptera, Acrididae, Oedipodinae) reveal convergence of wing morphology. Zoolog Scr 41:515–526
Husemann M, Sadílek D, Dey LS, Hawlitschek O, Seidel M (2021) New genome size estimates for band-winged and slant-faced grasshoppers (Orthoptera: Acrididae: Oedipodinae, Gomphocerinae) reveal the so far largest measured insect genome. Caryologia. Int J Cytol Cytosyst Cytogenet 73:111–120. https://doi.org/10.13128/caryologia-966
Hutchinson GE (1957) Concluding remarks. Cold Spring Harb Symp Quant Biol 22:415–427. https://doi.org/10.1101/SQB.1957.022.01.039
Jiménez L, Soberón J, Christen JA, Soto D (2019) On the problem of modeling a fundamental niche from occurrence data. Ecol Model 397:74–83
Jung M (2016) LecoS—A python plugin for automated landscape ecology analysis. Ecol Inform 31:18–21
Juszczyk I, Egger G, Müller N, Reich M (2020) Auswirkungen der Ausleitung der Oberen Isar auf die Auenvegetation. Auenmagazin 17:28–37
Kajtoch Ł, Cieślak E, Varga Z et al (2016) Phylogeographic patterns of steppe species in Eastern Central Europe: a review and the implications for conservation. Biodivers Conserv 25:2309–2339
Kalyaanamoorthy S, Minh BQ, Wong TK et al (2017) ModelFinder: fast model selection for accurate phylogenetic estimates. Nat Methods 14:587
Kearse M, Moir R, Wilson A et al (2012) Geneious basic: an integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics 28:1647–1649
Koopmann A (2000) An Naturschutzzielen und historischer Heidebauernwirtschaft orientierte Landwirtschaft auf Sandböden: Fallstudie Landschaftspflegehof Tütsberg (Lüneburger Heide). Göttinger Bodenkundliche Berichte 114:1–210
Kotze DJ, O’hara RB (2003) Species decline—but why? Explanations of carabid beetle (Coleoptera, Carabidae) declines in Europe. Oecologia 135:138–148
Krauss HA (1883) Neuer beitrag zur orthopteren-fauna Tirols mit beschreibung zurier neuer Pezotettix-arten. Zool Bot Ges Österreich:219–224
Landmann A (2017) Gefleckte Schnarrschrecke Bryodemella tuberculatum (FABRICIUS, 1775). Die Heuschrecken Österreichs:621–624
Laussmann T, Radtke A, Wiemert T, Dahl A (2010) 150 years of butterfly monitoring in Wuppertal-Effects of climate and landscape change (Lepidoptera). Entomologische Zeitschrift Mit Insekten-Börse 120:269–277
Lawton JH (1994) Population dynamic principles. Philos Trans R Soc Lond B 344:61–68
Leigh JW, Bryant D (2015) popart: full-feature software for haplotype network construction. Methods Ecol Evol 6:1110–1116
Leimu R, Mutikainen PIA, Koricheva J, Fischer M (2006) How general are positive relationships between plant population size, fitness and genetic variation? J Ecol 94:942–952
Liana A (2004) Bryodema tuberculatum (Fabricius, 1775). Polish red data book invertebrates (Internet version). http://www.iop.krakow.pl/pckz/opis. Accessed 15 Oct 2020
Lira-Noriega A, Manthey JD (2014) Relationship of genetic diversity and niche centrality: a survey and analysis. Evolution 68:1082–1093
Lucchini V, Galov A, Randi E (2004) Evidence of genetic distinction and long-term population decline in wolves (Canis lupus) in the Italian Apennines. Mol Ecol 13:523–536
Ludwig T, Storch I, Gärtner S (2009) Large-scale land use change may explain bird species declines in semi-natural areas: the case of black grouse population collapse in Lower Saxony, Germany. J Ornithol 150:871–882
Maas S, Detzel P, Staudt A (2002) Gefährdungsanalyse der Heuschrecken Deutschlands: Verbreitungsatlas, Gefährdungseinstufung und Schutzkonzepte; Ergebnisse aus dem F+ E-Vorhaben 89886015 des Bundesamtes für Naturschutz. pp. 401
Maas S, Detzel P, Staudt A (2011) Rote Liste und Gesamtartenliste der Heuschrecken (Saltatoria) Deutschlands. Rote Liste Gefährdeter Tiere, Pflanzen Und Pilze Deutschlands 3:577–606
Malcolm JR, Liu C, Neilson RP, Hansen L, Hannah LEE (2006) Global warming and extinctions of endemic species from biodiversity hotspots. Conserv Biol 20(2):538–548
Marchi M, Sinjur I, Bozzano M, Westergren M (2019) Evaluating WorldClim Version 1 (1961–1990) as the baseline for sustainable use of forest and environmental resources in a changing climate. Sustainability 11:3043
Massa B, Fontana P, Buzzetti F et al (2012) Fauna d’Italia–Orthoptera. Calderini 48:563
Monnerat C, Thorens P, Walter T, Gonseth Y (2007) Rote Liste Heuschrecken. BAFU, Bern
Nguyen L-T, Schmidt HA, Von Haeseler A, Minh BQ (2015) IQ-TREE: a fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Mol Biol Evol 32:268–274
Pan C, Hu J, Zhang X, Huang Y (2006) The DNA barcoding application of mtDNA COI genes in seven species of Catantopidae (Orthoptera). Entomotaxonomia 28:103–110
Papadopoulou A, Anastasiou I, Vogler AP (2010) Revisiting the insect mitochondrial molecular clock: the mid-Aegean trench calibration. Mol Biol Evol 27:1659–1672
Paxton RJ, Thorén PA, Tengö J et al (1996) Mating structure and nestmate relatedness in a communal bee, Andrena jacobi (Hymenoptera, Andrenidae), using microsatellites. Mol Ecol 5:511–519
Pebesma E, Bivand RS (2005) S classes and methods for spatial data: the sp package. R News 5:9–13
Pedersen HB, Møllenberg S (2017) Agriculture and Danish farm returns through 100 years. Statistics Denmark, Denmark
Pertoldi C, Hansen MM, Loeschcke V et al (2001) Genetic consequences of population decline in the European otter (Lutra lutra): an assessment of microsatellite DNA variation in Danish otters from 1883 to 1993. Proc R Soc Lond B 268:1775–1781
Peterson AT, Papeş M, Soberón J (2008) Rethinking receiver operating characteristic analysis applications in ecological niche modeling. Ecol Model 213:63–72
Peterson AT, Soberón J, Pearson RG, Anderson RP, Martínez-Meyer E, Nakamura M, Araújo MB (2011) Ecological niches and geographic distributions (MPB-49). Princeton University Press, Princeton
Pfeuffer E (2004) Zur Heuschreckenfauna des Schwarzwassertales, eines Seitenzubringers des Oberen Lech (Tirol), unter besonderer BerUcksichtigung von Bryodemella tuberculata und Chorthippus pullus. Articulata 19(2):195–203
Plieninger T, Draux H, Fagerholm N et al (2016) The driving forces of landscape change in Europe: a systematic review of the evidence. Land Use Policy 57:204–214
Poggio L, Simonetti E, Gimona A (2018) Enhancing the WorldClim data set for national and regional applications. Sci Total Environ 625:1628–1643
Potts SG, Biesmeijer JC, Kremen C et al (2010) Global pollinator declines: trends, impacts and drivers. Trends Ecol Evol 25:345–353
Preston DB, Johnson SG (2020) Generalist grasshoppers from thermally variable sites do not have higher thermal tolerance than grasshoppers from thermally stable sites-A study of five populations. J Therm Biol 88:102527
Pulliam HR (1988) Sources, sinks, and population regulation. Am Nat 132:652–661
QGIS Development Team (2020) QGIS Geographic Information System v. 3.10.3. Open Source Geospatial Foundation Project. http://qgis.osgeo.org. Accessed 18 Sept 2020
Quinger B (2014) Sandmagerrasen, offene Sandfluren und Binnendünen. Handbuch Naturschutz Und Landschaftspflege. https://doi.org/10.1002/9783527678471.hbnl2000011
R Core Team (2020) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna
Ratnasingham S, Hebert PD (2007) BOLD: the barcode of life data system (http://www.barcodinglife.org). Mol Ecol Notes, 7: 355–364
Reed DH, Frankham R (2003) Correlation between fitness and genetic diversity. Conserv Biol 17:230–237
Reich M (1991) Struktur und Dynamik einer Population von Bryodema tuberculata (Fabricius, 1775) (Saltaroria, Acrididae). Universität Ulm, pp 105
Reich M, Bargiel D, Rühmkorf H (2008) Die Obere Isar zwischen Fkm 253 und Fkm 232: Veränderungen der Vegetationsverhältnisse zwischen 1858 und 2006, Auswirkungen der Hochwasser 1999 und 2005 und Situation und Perspektive ausgewählter Zielarten. Gutachten im Auftrag des Bayerischen Landesamtes für Umwelt und des Wasserwirtschaftsamtes Weilheim, Hannover
Reinhardt K, Köhler G, Maas S, Detzel P (2005) Low dispersal ability and habitat specificity promote extinctions in rare but not in widespread species: the Orthoptera of Germany. Ecography 28:593–602
RStudio Team (2020) RStudio: integrated development for R. RStudio, PBC, Boston
Ruffo S (2003) Checklist e distribuzione della fauna italiana. Memorie del Museo Civico di Storia Naturale di Verona, Series 2, Sect. Scienze della Vita, 16, pp 307
Schmidt G-H, Lilge R (1997) Geographische Verbreitung der Oedipodinae (Orthopteroidea, Caelifera, Acrididae) in Europa und Randgebieten. Verlag Dr Kovac, Hamburg
Sergeev MG (1992) Distribution patterns of Orthoptera in North and Central Asia. J Orthoptera Res. https://doi.org/10.2307/3503557
Sinervo B, Mendez-De-La-Cruz F, Miles DB, Heulin B, Bastiaans E, Villagrán-Santa Cruz M, Sites JW (2010) Erosion of lizard diversity by climate change and altered thermal niches. Science 328(5980):894–899
Smelansky IE, Tishkov AA (2012) The steppe biome in Russia: ecosystem services, conservation status, and actual challenges. Eurasian steppes. Ecological problems and livelihoods in a changing world. Springer, Berlin, pp 45–101
Srinivasan G, Prabakar D (2013) A pictorial handbook on grasshop-pers of Western Himalayas. Zoological Survey of India, Kolkata
Urena-Aranda CA, Rojas-Soto O, Martinez-Meyer E et al (2015) Using range-wide abundance modeling to identify key conservation areas for the micro-endemic Bolson tortoise (Gopherus flavomarginatus). PLoS ONE. https://doi.org/10.1371/journal.pone.0131452
Van Aelst S, Rousseeuw P (2009) Minimum volume ellipsoid. Wiley Interdisciplin Rev Comput Stat 1:71–82
Van Dyck H, Van Strien AJ, Maes D, Van Swaay CA (2009) Declines in common, widespread butterflies in a landscape under intense human use. Conserv Biol 23:957–965
Vanbergen AJ, The Insect Pollinators Initiative (2013) Threats to an ecosystem service: pressures on pollinators. Front Ecol Environ 11:251–259
Vanderwal J, Shoo LP, Johnson CN, Williams SE (2009) Abundance and the environmental niche: environmental suitability estimated from niche models predicts the upper limit of local abundance. Am Nat 174:282–291
Voith J, Beckmann A, Sachteleben J et al (2016) Rote Liste und Gesamtartenliste der Heuschrecken (Saltatoria) Bayerns. Bayerisches Landesamt für Umwelt (LfU), Augsburg
Wagner DL (2019) Global insect decline: comments on Sánchez-Bayo and Wyckhuys (2019). Biol Conserv 233:332–333
Werle E, Schneider C, Renner M et al (1994) Convenient single-step, one tube purification of PCR products for direct sequencing. Nucleic Acids Res 22:4354
Wickham H (2011) The split-apply-combine strategy for data analysis. J Stat Softw 40:1–29
Wickham H (2016) ggplot2: elegant graphics for data analysis. Springer-Verlag, New York
Wilke C (2019) cowplot: streamlined plot theme and plot annotations for “ggplot2.” Version R package version 1.0.0. https://CRAN.R-project.org/package=cowplot. Accessed 9 Nov 2020
Yañez-Arenas C, Martínez-Meyer E, Mandujano S, Rojas-Soto O (2012) Modelling geographic patterns of population density of the white-tailed deer in central Mexico by implementing ecological niche theory. Oikos 121:2081–2089
Zacher F (1919) Beitrage zur Kenntnis der Geradfluglerfauna des deutschen Alpengebietes. Entomologische Mitteilungen 8:96–99
Zuna-Kratky T, Fontana P, Roesti C et al (2016) Bryodemella tuberculata. IUCN Redlist. https://www.iucnredlist.org/species/16084462/70292211. Accessed 13 Mar 2020
Zuna-Kratky T, Landmann A, Illich I et al (2017) die Heuschrecken österreichs. denisia 39. Biologiezentrum des Oberösterreichischen landesmuseums, Linz
Acknowledgements
This work benefited from expertise sharing and discussions within the DFG priority program SPP 1991. We thank Benjamin Wildermuth and Sajad Noori for support in the molecular lab. Moreover, we would like to thank the German-Mongolian expeditions for the possibility of obtaining samples from many locations in Mongolia. All sampling in Mongolia was performed under government permit #200008 (02.01.2020). Sampling in Germany was performed with the permit granted on 15 July 2019 by the government of Upper Bavaria. Many thanks to Eduardas Budrys (Nature Research Centre Vilnius, Lithuania), Jürgen Deckert (MfN Berlin), Paolo Fontana (CTT San Michele All’Adige), Andrey Gorochov (ZIN St. Petersburg), Tanja Koskela (JYU Jyväskylä), Judith Marshall (BNHM London), Bruno Massa (SAF Palermo), Ralph Peters (ZFMK Bonn), Susanne Randolph (NHM Vienna), Thomas Schmitt (SDEI Müncheberg), Peter Schwendinger (MNHG Geneva), Villu Soon (UT Tartu), Arnold Staniczek (SMNS Stuttgart), Frank Steinheimer (ZNS Halle), Lars Vilhelmsen (SNM Copenhagen), Luc Willemse (NMNL Leiden) and Przemek Żurawlew (Projekt Orthoptera Polski) for providing data and to Thomas Wagner, Teresa Krebs, and students (TUM Weihenstephan; University of Hamburg) for field support. Thanks to the editor and two anonymous reviewers for valuable comments on a previous version of the paper. Moreover, we thank all citizen scientists who collected and published occurrence data to public databases.
Funding
Open Access funding enabled and organized by Projekt DEAL. Orthoptera Species File to MH and LSD; Deutsche Gesellschaft für Orthopterologie to LSD; Gesellschaft für Wild- und Jagdtierforschung to LSD; DAAD Promos to LSD; Risk and Innovation Fund of the University of Hamburg to MH, MS and OH; SNSB Innovativ to OH; BMBF Promotionsförderung of Heinrich-Böll-Stiftung to LSD; Russian Federal Fundamental Scientific Research Program (FWGS-2021-0002) to MGS; Deutsche Forschungsgemeinschaft DFG HA 7255/2-1 to OH.
Author information
Authors and Affiliations
Contributions
LSD, MS, MH, OH designed the study; LSD, MH, OH, MGS, DL, SQX collected data; LSD and MS performed analyses; LSD, MH, OH, MS, MGS, DL, SQX wrote the manuscript; LSD and MH coordinated the writing process.
Corresponding author
Ethics declarations
Conflicts of interest
All authors contributed significantly to the manuscript and declare no conflict of interest.
Additional information
Communicated by Akihiro Nakamura.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Dey, LS., Simões, M.V.P., Hawlitschek, O. et al. Analysis of geographic centrality and genetic diversity in the declining grasshopper species Bryodemella tuberculata (Orthoptera: Oedipodinae). Biodivers Conserv 30, 2773–2796 (2021). https://doi.org/10.1007/s10531-021-02221-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10531-021-02221-8