Abstract
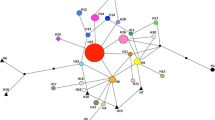
Unraveling the origin and colonization history of invasive plants is a long-standing challenge in evolutionary and conservation biology. The knowledge of the origin of the invasive plants in Europe is often confounded by limited sampling in the source region. We determined the extent of genetic structuring in the native range and reconstructed the origin and the invasion history of Persian hogweed, Heracleum persicum, into Europe. We used allelic polymorphism of microsatellite markers obtained from 36 Iranian populations from Middle East combined with data from 38 European populations representing the major native and introduced ranges, respectively. Comprehensive sampling in the native range covered 97% of allelic diversity found in the introduced range, and showed that allelic variation, heterozygosity and population differentiation are significantly reduced in the introduced populations. Results from Bayesian structure, neighbor-net, and ordination analyses showed that populations in the native range consist of three distinct genetic clusters: Eastern, Central, and Western. Although we observed high genetic differentiation among these groups, the Western cluster was genetically closer to the Central cluster. Approximate Bayesian Computation (ABC) analysis supports at least three independent origins for European populations. No European population originated from the Eastern cluster within the native range. Danish populations originated from the Central cluster, whereas the UK and Finland populations originated independently from the Western cluster. Norwegian populations originated from UK, and subsequently established the FI-Kar population in Finland. Our results shed light on the complex origin and history of an aggressive invasive plant species in Europe, supporting contribution from multiple genetic lineages in recent ancestry of introduced populations, thus suggesting the potential for ecological diversification within the introduced range.





Similar content being viewed by others
References
Austerlitz F, Mariette S, Machon N, Gouyon PH, Godelle B (2000) Effects of colonization processes on genetic diversity: differences between annual plants and tree species. Genetics 154:1309–1321
Baker HG (1955) Self-compatibility and establishment after “long-distance” dispersal. Evolution 9:347–349
Balloux F (2004) Heterozygote excess in small populations and the heterozygote-excess effective population size. Evolution 58:1891–1900
Bazin É, Mathé-Hubert H, Facon B, Carlier J, Ravigné V (2014) The effect of mating system on invasiveness: some genetic load may be advantageous when invading new environments. Biol Invasions 16:875–886
Bellard C, Bertelsmeier C, Leadley P, Thuiller W, Courchamp F (2012) Impacts of climate change on the future of biodiversity. Ecol Lett 15:365–377
Bossdorf O, Auge H, Lafuma L, Rogers WE, Siemann E, Prati D (2005) Phenotypic and genetic differentiation between native and introduced plant populations. Oecologia 144:1–11
Bryant D, Moulton V (2004) Neighbor-Net: an agglomerative method for the construction of phylogenetic networks. Mol Biol Evol 21:255–265
Buckley YM, Catford J (2016) Does the biogeographic origin of species matter? Ecological effects of native and non-native species and the use of origin to guide management. J Ecol 104:4–17
Calcagno V, Jarne P, Loreau M, Mouquet N, David P (2017) Diversity spurs diversification in ecological communities. Nat Commun 8:15810
Cannings C, Edwards AW (1969) Expected genotypic frequencies in a small sample: deviation from Hardy-Weinberg equilibrium. Am J Hum Genet 21:245–247
Colautti RI, Alexander JM, Dlugosch KM, Keller SR, Sultan SE (2017) Invasions and extinctions through the looking glass of evolutionary ecology. Philos Trans R Soc Lond B Biol Sci 372:20160031
Cornuet J-M, Pudlo P, Veyssier J, Dehne-Garcia A, Gautier M, Leblois R, Marin J-M, Estoup A (2014) DIYABC v2.0: a software to make approximate Bayesian computation inferences about population history using single nucleotide polymorphism. DNA Seq Microsatellite Data Bioinform 30:1187–1189
Didham RK, Tylianakis JM, Hutchison MA, Ewers RM, Gemmell NJ (2005) Are invasive species the drivers of ecological change? Trends Ecol Evol 20:470–474
Djamali M, Brewer S, Breckle SW, Jackson ST (2012) Climatic determinism in phytogeographic regionalization: a test from the Irano-Turanian region, SW and Central Asia. Flora 207:237–249
Dlugosch KM, Parker IM (2008) Founding events in species invasions: genetic variation, adaptive evolution, and the role of multiple introductions. Mol Ecol 17:431–449
Durka W, Bossdorf O, Prati D, Auge H (2005) Molecular evidence for multiple introductions of garlic mustard (Alliaria petiolata, Brassicaceae) to North America. Mol Ecol 14:1697–1706
EPPO (2009) EPPO data sheet on invasive alien plants: Heracleum mantegazzianum, Heracleum sosnowskyi and Heracleum persicum. OEPP/EPPO Bull 39:489–499
Excoffier L, Lischer H (2010) Arlequin suite ver 3.5: a new series of programs to perform population genetics analyses under Linux and Windows. Mol Ecol Resour 10:564–567
Falahati-Anbaran M, Mohammadi Bazargani M, Rohloff J (2018) Large scale geographical mapping of essential oil volatiles in Heracleum (Apiaceae): identification of novel compounds and unraveling cryptic variation. Chem Biodivers 15:e1800230
Falush D, Stephens M, Pritchard JK (2003) Inference of population structure using multilocus genotype data: linked loci and correlated allele frequencies. Genetics 164:1567–1587
Franklin J, Serra-Diaz JM, Syphard AD, Regan HM (2016) Global change and terrestrial plant community dynamics. Proc Natl Acad Sci U S A 113:3725–3734
Fröberg L (2010) Heracleum L. In: Jonsell B, Karlsson T (eds) Flora Nordica (Thymelaeaceae to Apiaceae). Stockholm, Sweden, The Swedish Museum of Natural History, pp 224–234
Gederaas L, Moen TL, Skjelseth S, Larsen LMK (2012) Alien species in Norway - with the Norwegian Black List 2012. Trondheim Norway, Norwegian Biodiversity Infomation Centre (NBIC)
Genton BJ, Shykoff JA, Giraud T (2005) High genetic diversity in French invasive populations of common ragweed, Ambrosia artemisiifolia, as a result of multiple sources of introduction. Mol Ecol 14:4275–4285
Gioria M, Pyšek P, Moravcová L (2012) Soil seed banks in plant invasions: promoting species invasiveness and long-term impact on plant community dynamics. Preslia 84:327–350
Hagenblad J, Hülskötter J, Acharya KP, Brunet J, Chabrerie O, Cousins SAO, Dar PA, Diekmann M, De Frenne P, Hermy M, Jamoneau A, Kolb A, Lemke I, Plue J, Reshi ZA, Graae BJ (2015) Low genetic diversity despite multiple introductions of the invasive plant species Impatiens glandulifera in Europe. BMC Genet 16:103
Hamelin RC, Roe AD (2019) Genomic biosurveillance of forest invasive alien enemies: a story written in code. Evol Appl 13:95–115
Henry P, Le Lay G, Goudet J, Guisan A, JahodovÁ Š, Besnard G (2009) Reduced genetic diversity, increased isolation and multiple introductions of invasive giant hogweed in the western Swiss Alps. Mol Ecol 18:2819–2831
Hornoy B, Atlan A, Roussel V, Buckley YM, Tarayre M (2013) Two colonisation stages generate two different patterns of genetic diversity within native and invasive ranges of Ulex europaeus. Heredity 111:355–363
Jahodová Š, Trybush S, Pyšek P, Wade M, Karp A (2007) Invasive species of Heracleum in Europe: an insight into genetic relationships and invasion history. Divers Distrib 13:99–114
Kalinowski ST (2005) hp-rare 1.0: a computer program for performing rarefaction on measures of allelic richness. Mol Ecol Notes 5:187–189
Kelager A, Pedersen JS, Bruun HH (2013) Multiple introductions and no loss of genetic diversity: invasion history of Japanese Rose, Rosa rugosa, in Europe. Biol Invasions 15:1125–1141
Kopelman NM, Mayzel J, Jakobsson M, Rosenberg NA, Mayrose I (2015) Clumpak: a program for identifying clustering modes and packaging population structure inferences across K. Mol Ecol Resour 15:1179–1191
Krinke L, Moravcová L, Pyšek P, Jarošík V, Pergl J, Perglová I (2005) Seed bank of an invasive alien, Heracleum mantegazzianum, and its seasonal dynamics. Seed Sci Res 15:239–248
Langella O (1999) Populations, 1.2.32, http://bioinformatics.org/~tryphon/populations/. CNRS UPR9034
Lau JA, Schultheis EH (2015) When two invasion hypotheses are better than one. New Phytol 205:958–960
Marrs RA, Sforza R, Hufbauer RA (2008a) Evidence for multiple introductions of Centaurea stoebe micranthos (spotted knapweed, Asteraceae) to North America. Mol Ecol 17:4197–4208
Marrs RA, Sforza R, Hufbauer RA (2008b) When invasion increases population genetic structure: a study with Centaurea diffusa. Biol Invasions 10:561–572
McCarty JP (2001) Ecological consequences of recent climate change. Conserv Biol 15:320–331
Mooney HA, Cleland EE (2001) The evolutionary impact of invasive species. Proc Natl Acad Sci U S A 98:5446–5451
Nei M, Maruyama T, Chakraborty R (1975) The bottleneck effect and genetic variability in populations. Evolution 29:1–10
Nei M, Tajima F, Tateno Y (1983) Accuracy of estimated phylogenetic trees from molecular data. J Mol Evol 19:153-170
Nielsen C, Ravn HP, Skov L, Nentwig W, Wade M (2005) The giant Hogweed best practice manual: guidelines for management and control of an invasive weed in Europe. Forest & Landscape Denmark, Hørsholm Kongevej 11, DK-2970 Hørsholm, Denmark
Novo M, Cunha L, Maceda-Veiga A, Talavera JA, Hodson ME, Spurgeon D, Bruford MW, Morgan AJ, Kille P (2015) Multiple introductions and environmental factors affecting the establishment of invasive species on a volcanic island. Soil Biol Biochem 85:89–100
Ouborg NJ, Piquot Y, Van Groenendael JM (1999) Population genetics, molecular markers and the study of dispersal in plants. J Ecol 87:551–568
Pairon M, Petitpierre B, Campbell M, Guisan A, Broennimann O, Baret PV, Jacquemart A-L, Besnard G (2010) Multiple introductions boosted genetic diversity in the invasive range of black cherry (Prunus serotina; Rosaceae). Ann Bot 105:881–890
Pannell JR, Barrett SCH (1998) Baker’s law revisited: reproductive assurance in a metapopulation. Evolution 52:657–668
Pantoja PO, Paine CET, Vallejo-Marín M (2018) Natural selection and outbreeding depression suggest adaptive differentiation in the invasive range of a clonal plant. Proc R Soc B 285:20181091
Pantoja PO, Simón-Porcar VI, Puzey JR, Vallejo-Marín M (2017) Genetic variation and clonal diversity in introduced populations of Mimulus guttatus assessed by genotyping at 62 single nucleotide polymorphism loci. Plant Ecol Divers 10:5–15
Pergl J, Müllerová J, Perglová I, Herben T, Pyšek P (2011) The role of long-distance seed dispersal in the local population dynamics of an invasive plant species. Divers Distrib 17:725–738
Perglová I, Pergl J, Pyšek P (2007) Reproductive ecology of Heracleum mantegazzianum. In: Pyšek P, Ravn HP, Nentwig W, Cock MJW (eds) Ecology and management of giant Hogweed (Heracleum mantegazzianum). CABI, Wallingford, pp 55–73
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Pudovkin AI, Zaykin DV, Hedgecock D (1996) On the potential for estimating the effective number of breeders from heterozygote-excess in progeny. Genetics 144:383–387
Pyšek P, Lambdon PW, Arianoutsou M, Kühn I, Pino J, Winter M (2009) Alien vascular plants of Europe. Handbook of Alien Species in Europe. Springer, Netherlands, Dordrecht, pp 43–61
Pyšek P, Pergl J, JahodovÁ Š, Moravcová L, Müllerová J, PerglovÁ I, Wild J (2010) The hogweed story: Invasion of Europe by large Heracleum species. In: Settele (ed) Atlas of Biodiversity Risk. Pensoft pp. 150–151
Rijal DP, Alm T, Jahodová Š, Stenøien HK, Alsos IG (2015a) Reconstructing the invasion history of Heracleum persicum (Apiaceae) into Europe. Mol Ecol 24:5522–5543
Rijal DP, Falahati-Anbaran M, Alm T, Alsos IG (2015b) Microsatellite markers for Heracleum persicum (Apiaceae) and allied taxa: application of next-gneration sequencing to develop genetic resources for invasive species management. Plant Mol Biol Rep 33:1381–1390
Rijal DP, Alm T, Nilsen L, Alsos IG (2017) Giant invasive Heracleum persicum: Friend or foe of plant diversity? Ecol Evol 7:4936–4950
Rousset F (2008) genepop’007: a complete re-implementation of the genepop software for Windows and Linux. Mol Ecol Resour 8:103–106
Sakai AK, Allendorf FW, Holt JS, Lodge DM, Molofsky J, With KA, Baughman S, Cabin RJ, Cohen JE, Ellstrand NC, McCauley DE, O’Neil P, Parker IM, Thompson JN, G. WS, (2001) The population biology of invasive species. Annu Rev Ecol Syst 32:305–332
Schirmel J, Bundschuh M, Entling MH, Kowarik I, Buchholz S (2016) Impacts of invasive plants on resident animals across ecosystems, taxa, and feeding types: a global assessment. Global Change Biol 22:594–603
Shirk RY, Hamrick JL, Zhang C, Qiang S (2014) Patterns of genetic diversity reveal multiple introductions and recurrent founder effects during range expansion in invasive populations of Geranium carolinianum (Geraniaceae). Heredity 112:497–507
Slatkin M, Excoffier L (2012) Serial founder effects during range expansion: a spatial analog of genetic drift. Genetics 191:171–181
Stenøien HK, Såstad SM (1999) Genetic structure in three haploid peat mosses (Sphagnum). Heredity 82:391–400
Stout JC, Duffy KJ, Egan PA, Harbourne M, Hodkinson TR (2015) Genetic diversity and floral width variation in introduced and native populations of a long-lived woody perennial. AoB Plants 7: plu087
Theoharides KA, Dukes JS (2007) Plant invasion across space and time: factors affecting nonindigenous species success during four stages of invasion. New Phytol 176:256–273
Trottier N, Groeneveld E, Lavoie C (2017) Giant hogweed at its northern distribution limit in North America: experiments for a better understanding of its dispersal dynamics along rivers. River Res Appl 33:1098–1106
Vilà M, Espinar JL, Hejda M, Hulme PE, Jarošík V, Maron JL, Pergl J, Schaffner U, Sun Y, Pyšek P (2011) Ecological impacts of invasive alien plants: a meta-analysis of their effects on species, communities and ecosystems. Ecol Lett 14:702–708
Walker NF, Hulme PE, Hoelzel AR (2003) Population genetics of an invasive species, Heracleum mantegazzianum: implications for the role of life history, demographics and independent introductions. Mol Ecol 12:1747–1756
Weir BS, Cockerham CC (1984) Estimating F-statistics for the analysis of population structure. Evolution 38:1358–1370
Wellborn GA, Langerhans RB (2015) Ecological opportunity and the adaptive diversification of lineages. Ecol Evol 5:176–195
Zhao J, Solis-Montero L, Lou A, Vallejo-Marin M (2013) Population structure and genetic diversity of native and invasive populations of Solanum rostratum (Solanaceae). PLoS ONE 8:e79807
Zhao SY, Sun SG, Dai C, Gituru RW, Chen JM, Wang QF (2015) Genetic variation and structure in native and invasive Solidago canadensis populations. Weed Res 55:163–172
Acknowledgements
We thank the NTNU University Museum and the Department of Biology at NTNU for providing laboratory resources for microsatellite genotyping. We also appreciate valuable comments suggested by two independent reviewers on the previous version of the manuscript.
Funding
This work was supported by a grant from Iran National Science Foundation (grant number 92038838).
Author information
Authors and Affiliations
Contributions
MFA designed research; MFA performed research; MFA and HKS contributed new reagents or analytical tools; MFA analyzed data; MFA wrote the paper with contribution from co-authors.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Data availability
The data that support the findings of this study will openly be available in Dryad at https://doi.org/10.5061/dryad.pzgmsbchn
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Falahati-Anbaran, M., Rijal, D.P., Lundemo, S. et al. Disentangling the genetic origin of Heracleum persicum (Apiaceae) in Europe: multiple introductions from multiple source populations. Biol Invasions 23, 3871–3890 (2021). https://doi.org/10.1007/s10530-021-02618-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10530-021-02618-0