Abstract
Pellet counts are widely used to monitor ungulates but rely on the assumption that pellets of different species are correctly identified in the field. Recent studies question this assumption using DNA barcoding techniques to check field identification rates. For Europe, which is undergoing a rapid shift towards more diverse ungulate assemblages, such an assessment is still missing. Using DNA barcoding on 3889 fecal samples from nine ungulate species in four European countries, we found average field misidentification rates varied from 0.6% for horse (Equus ferus) to 41.1% for roe deer (Capreolus capreolus). Most identification errors occurred between similar-sized species from the same taxonomic family. For a subset of samples from Sweden, we looked at the effect of dung morphometry, observer experience, and season on species identification success. Dung morphometry clearly distinguished moose (Alces alces) but not red (Cervus elaphus), roe, and fallow deer (Dama dama). Experienced observers performed better than novices for red and fallow deer although still making significant identification errors (26% and 17% incorrectly identified). Identification success was higher during spring and winter (\( \overline{x} \)= 86%) than summer and autumn (\( \overline{x} \)= 74%). We question pellet counts as an accurate monitoring tool where similar-sized species coexist and monitoring relates to the whole community. For this increasingly common situation across Europe, DNA testing or camera traps may be a better alternative. Pellet counts remain useful where only few species with clearly different dung morphology coexist (e.g., moose and roe deer) or when focused on species with distinctive dung morphology (e.g., moose).
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The sustainable management of wild ungulate populations is of high priority in many European countries but is becoming increasingly challenging (Apollonio et al. 2017). Rapidly growing populations and a shift towards more diverse ungulate communities, where previously only one or two species occurred (Apollonio et al. 2010), present novel and sometimes complex situations, which calls for critical re-assessment of established management tools. A key part of this management is population monitoring. The use of feces is a common monitoring approach for the study of many terrestrial mammals (Chame 2003; Rouco et al. 2009; Lonsinger et al. 2015), including ungulates (Cromsigt et al. 2009). Fecal pellets can be collected non-invasively in large numbers and with relative ease. This opens numerous avenues of investigation, including diet, disease, stress level, habitat use, and population dynamics (Kohn and Wayne 1997). The counting of pellet groups is the most commonly used method for estimating ungulate abundances and has been described as very accurate due to the well-established correlation between number of pellet groups and individuals (Campbell et al. 2004; Putman 1984). Species-level identification of fecal pellets has traditionally relied upon assessment of morphometric variables like length, width, weight, the number of pellets per group, and various derivatives, e.g., the length/width ratio and volume. Less common metrics include the proportion of adherence (Yamashiro et al. 2013), pH (Elliott and Longhurst 1984), or near infrared reflectance spectra (Tolleson et al. 2005). Several studies have shown that morphometric measurements can work well for distinguishing ungulate pellet groups at the intraspecific level, especially if pellet size scales with body size. For example, MacCracken and Van Ballenberge (1987) were able to correctly identify 91% of moose pellet groups as originating from adult males, adult females, and yearlings based on pellet volume. Ball (2010) successfully used pellet length to distinguish calf and yearling age-classes from adults in caribou (Rangifer tarandus caribou), and Woodruff et al. (2016) found a 98% probability of correctly classifying fawns versus adult Sonoran pronghorn (Antilocapra americana sonoriensis) using pellet width.
However, at the interspecific level, concerns about misidentifications arose early (Seton 1925). Alvarez (1994) found that only 78% of fallow deer and 81% of red deer pellets collected in central Spain could be correctly assigned by discriminant analysis. In a study on sympatric black-tailed deer (Odocoileus hemionus columbianus), axis deer (Cervus axis), and fallow deer, Elliott and Longhurst (1984) unsuccessfully tried to circumnavigate the problem of discriminating morphologically similar pellets by measuring fecal pH. In recent years, the advancement of molecular methods, which allow for species identification based on DNA markers, and the sharp decline in associated costs have enabled researchers to investigate misclassification rates in multi-species ungulate systems more comprehensively. The results indicate that the reliance on fecal morphology can lead to alarmingly erroneous outcomes. For example, Yamashiro et al. (2013) found that 41% of sika deer (Cervus nippon) pellets were misidentified as Japanese serow (Capricornis crispus). A study involving six neotropical and largely sympatric deer species found correct dung identification rates to vary from 56 to 92% depending on the location (Costa et al. 2017), and Bowkett et al. (2013) reported an overall identification accuracy of only 59% for five forest-associated antelopes in Tanzania.
Even though feces are widely used for ungulate monitoring in Europe (Plhal et al. 2014; Rönnegård et al. 2008; Prokesová et al. 2006; Marques et al. 2001; Lioy et al. 2015), the problem of misidentification bias has rarely been addressed, even in studies recommending the use of dung counts for estimating abundances (Tottewitz et al. 1996). With a few exceptions, e.g., Alvarez (1994), studies assessing the efficacy of dung morphometry for species identification in the increasingly diverse European ungulate communities are largely missing. Here, we used a large dataset of DNA-validated fecal samples comprising nine free-ranging ungulate species to:
-
1.
Quantify species misidentification rates in five different European multi-species ungulate systems
-
2.
Test the efficacy of dung morphometry to discriminate between moose, red deer, fallow deer, and roe deer as these species increasingly co-occur across Europe, are frequently being monitored through dung counts, and produce similarly shaped pellets
-
3.
Test the effect of ungulate species, observer experience, and season on species identification rates
We expected misidentification rates to be highest between ecologically similar species of approximately the same size, owing to their presumably similar dung morphologies. Because the appearance of ungulate feces can vary, even at the intraspecific level, we expected identification success to be higher among experienced observers than novices and also for misidentification errors to increase during the growing season. The latter corresponds to the time when calves, which produce small pellets, enter the population. We discuss sources of bias and make situation-specific recommendations that would aid in the monitoring of multi-species ungulate systems.
Methods
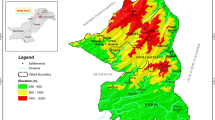
Fecal samples were collected in Sweden (two areas), the Netherlands (two areas), Germany (one area) and Poland (one area, Fig. 1). DNA identification of ungulate species was standardized for all fecal samples, but the number of ungulate species, community composition, sampling period, and available meta-data varied among countries. As a consequence, different subsets of the data were used in the subsequent analyses. All statistical tests were carried out with R (R Core Team 2017) at a significance level of alpha = 0.05.
Sample collection and storage
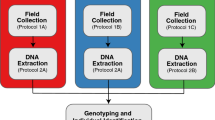
For DNA analysis, we collected approximately 2 g of fresh ungulate feces into sterile, airtight 20 ml scintillation tubes filled with silica gel desiccant (~ 1–3 mm, with indicator (orange gel), Merck KGaA, Germany) (Taberlet et al. 2018; DeMay et al. 2013). We classified feces as fresh if they still had a wet, shiny surface and showed no signs of infestation by insects or microorganisms (Hemami and Dolman 2005). To avoid contamination, we either used disposable plastic spoons or nudged fecal pellets directly into the scintillation bottle, avoiding all contact with the collector or other samples. The silica-dried samples were then stored in the dark at room temperature until further processing.
In Sweden, we collected samples in two study areas, Nordmaling (northern Sweden) and Öster Malma (central Sweden). The Nordmaling area is dominated by boreal forests of Scots pine (Pinus sylvestris), Norway spruce (Picea abies), and birch (Betula sp.), interspersed with mires and some agricultural lands. Öster Malma lies in the boreal-nemoral transition zone characterized by a milder climate, higher proportions of deciduous trees, including pedunculate oak (Quercus robur), and more extensive agricultural land with more diversified crops. Moose, red deer, fallow deer, and roe deer occur sympatrically in parts of both study areas. Wild boar (Sus scrofa) is currently only present in Öster Malma whereas reindeer (Rangifer tarandus tarandus) occurs sporadically during the winter in Nordmaling. Most samples were collected just after snowmelt (central: March–April, northern: May–June) in 2015, 2016, and 2017 as part of annual pellet count surveys. These are carried out on established sampling grids of 50 square transects (1 × 1 km), spaced several kilometers apart, in each study area. In addition, we collected samples on a subset of the transects in both areas on a monthly basis from September 2016 until November 2017.
In the Netherlands, sample collections were carried out during the period from May 2017 until January 2018 in two nature reserves: (1) Kraansvlak is a 330-ha fenced area within the National park Zuid-Kennemerland. This heterogeneous coastal dune landscape is characterized by open sandy dune areas, dry and wet grasslands, both deciduous and coniferous forest patches, and shrubland characterized by a continuous grass layer and shrubs, including spindle tree (Euonymus europaeus), sea-buckthorn (Hippophae rhamnoides), hawthorn (Crataegus monogyna), and creeping willow (Salix repens). The area is home to European bison (Bison bonasus), cattle (Bos taurus), horse, fallow deer, and roe deer. (2) Maashorst encompasses 3500 ha and is located in the Dutch province of North Brabant. The area consists of both coniferous and deciduous forests, heathlands, dunes, grasslands, and fens. Different herbivore assemblages are found in different fenced areas within the reserve. We collected samples of European bison, exmoor pony, and roe deer in a fenced area of 170 ha, while Tauros cattle samples were collected in an adjacent compartment of 250 ha. Samples were collected twice a month along transect lines that covered the habitat variability offered in each area. Deer species can move through the fences and no supplementary fodder is offered at any time of the year in any of the study areas.
In Germany, we collected samples from red deer, roe deer, and wild boar between May and October 2017 along line transects that were established as part of the biodiversity monitoring in the Bavarian Forest National Park (Bässler et al. 2009). The park protects mixed mountain forests consisting mainly of Norway spruce, European beech (Fagus sylvatica), and silver fir (Abies alba).
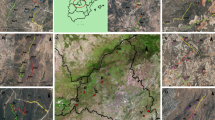
In Poland, we also collected samples between May and October 2017 from moose, European bison, red deer, roe deer, and wild boar along four transects (each 1 km) placed inside the Białowieża National Park and four in the adjacent forest managed by the State Forestry. Ungulate pellets were collected on six plots of 100 m2 that were equally spaced along the transect. Transects were randomly placed at different distances from the forest edge to cover the variation in forest types present in the area and hence provide a representative sample of the vegetation present in the area. The Białowieża Forest contains the last remnants of Europe’s temperate primeval forest, which is largely comprised of lime (Tilia sp.), hornbeam (Carpinus betulus), and oak (Quercus sp.) trees. Small-scale agriculture occurs at the forest edges.
Species identification in the field and molecular validation
Putative ungulate species were assigned to pellet groups by observers in the field based on visual inspection and their knowledge about ungulate communities in the respective study areas. To validate the field identifications of species, we used a universal primer pair for DNA metabarcoding of mammals (Giguet-Covex et al. 2014), which amplifies a section of the mitochondrial 16s gene. DNA processing, filtering, and taxonomic annotation are described in Appendix A (Online Resource 1). Samples with ambiguous taxonomic annotation, e.g., containing sequences from two ungulate species at nearly equal proportions, were removed from further analyses (see Appendix A/Online Resource 1 for details). We report species misidentification rates as the discrepancies between field identification of pellets by observers versus the corresponding DNA results.
Morphometric measurements
Morphometric measurements were recorded for moose, red deer, fallow deer, reindeer, and roe deer pellets collected at the two Swedish study areas between March and June 2017. To increase the likelihood of sampling different individuals, we placed at least 200 m between samples of the same putative species. This distance corresponds to the spacing of pellet count plots in the Swedish ungulate monitoring system and represents a compromise between avoiding pseudoreplication and finding enough dung in areas with low ungulate population densities. For each fresh pellet group, we first counted the number of pellets and recorded the species based on visual inspection. In cases of suspected fallow deer or roe deer, we noted both the species identification based on a pellet-number threshold (fallow deer > 45 pellets; roe deer ≤ 45 pellets) as suggested by Edenius (2012) and the observer’s opinion, if it differed. We then randomly selected five pellets (Haesler and Senn 2012), measured length and width (to the nearest 0.1 mm), and averaged the values for each pellet group (Woodruff et al. 2016). Initially, we used precision calipers for the measurements but this proved to be impractical due to the softness of pellets, which made it difficult to measure dimensions precisely and consistently. These difficulties were exacerbated during rainy weather. We therefore tested the alternative and faster approach of photographing pellets on laminated grid paper and then later measuring digitally using the free software Digimizer (version 4.6.1, available at www.digimizer.com). Upon finding no significant differences between the two methods, based on measuring 50 pellets, in either length (t(98) = 0.36, p = 0.72) or width (t(98) = 0.64, p = 0.53), we switched to photographs only. To measure pellet shape, we used the length/width ratio. We excluded reindeer from the analyses due to the small sample size (n = 3). To test for interspecific differences of the morphometric measurements, we used non-parametric Kruskall-Wallis followed by Dunn’s post hoc tests with Benjamin-Hochberg corrections. Additionally, we used discriminant analysis with “leave-one-out” (jackknifed) cross-validation (R-package MASS (Venables and Ripley 2002)) on the morphometric variables (log10-transformed to improve homogeneity of variance) to assign pellet groups to species.
Effects of observer experience and season on species identification success
To investigate the effect of observer experience on species identification, we used a subset of the Swedish data consisting of 386 moose, 122 roe deer, 228 red deer, and 311 fallow deer samples. We spatially restricted the dataset to the Öster-Malma study area where moose, red deer, fallow deer, and roe deer occur sympatrically, and further confined the sampling period to March and April. This corresponds to the time when dung count surveys are usually carried out in the area by management. To avoid species-specific collection bias, we only included observers who had collected samples from all four deer species. Observers were classified as either “experienced” (i.e., five field technicians with at least 3 years of experience in carrying-out dung pellet counts) or “novice” (i.e., three first-time participants in dung pellet counts). To these data, we fitted a logistic regression model using species identification success (true/false [1/0], based on field vs. DNA identification) as the binary response and “species” and “experience,” as well as their interaction, as categorical predictor variables. For post hoc pairwise comparisons, we used Tukey-adjusted least-squares means as offered in the lsmeans R package (Lenth 2016).
To assess the effect of season on species identification success, we used another subset of the Swedish data consisting of monthly collections at both study sites (Öster-Malma and Nordmaling) by two similarly experienced observers (359 moose, 163 roe deer, 359 red deer, and 227 fallow deer samples). Seasons were assigned by phenology rather than by calendar (spring: April–May, summer: June–August, autumn: September–October, winter: November–March).
Results
A total of 3889 samples (87%) were successfully amplified and taxonomically assigned based on DNA markers as described in Appendix A. Table 1 provides an overview of sample numbers and misidentification rates by species and location. Misidentification of the species ranked (from highest to lowest): roe deer > fallow deer > reindeer > cattle > wild boar > red deer > bison > moose > horse. In cases of misidentification in the field, the correct species generally corresponded to the one most similar in body size and pellet shape (see Fig. 2). For example, false identifications of bison turned out to be mostly cattle or in the case of moose, red deer.
Overview of the most common identification mistakes. Species names correspond to visual classification of fecal pellets by observers in the field. The size of the gray circles denotes body mass. Arrows point to the correct species as identified by DNA testing. Arrow thickness represents the percentage of misidentification while the color indicates the study area (GER—Germany, NL—Netherlands, POL—Poland, SE—Sweden). For example, incorrectly observer-identified roe deer samples correspond mostly to red deer in Germany and Poland, but fallow deer in Sweden and the Netherlands
Morphometric measurements
In total, 196 (78%) of the measured pellet groups passed the DNA identification quality threshold (77 moose, 38 roe deer, 51 red deer, 27 fallow deer, and 3 reindeer). Species had a significant effect on all measurements (Kruskal-Wallis tests, p < 0.05), with the pellet size measurements following the gradient in body size from largest (moose) to smallest (roe deer, Fig. 3). Larger ungulates produced not only larger pellets but also larger pellet groups. In the post hoc between-species comparisons, moose and red deer differed significantly from each other in all morphometric measurements. They also differed from fallow deer and roe deer, except in pellet shape, where only red deer feces were significantly different (slightly rounder) from the other species. Fallow deer and roe deer strongly overlapped for all measurements except pellet group size. The commonly used threshold of 45 pellets separated the means for both species. However, 30% of the fallow deer samples fell below that threshold, and 34% of the roe deer samples above. Although moose generally separated best in the morphometric measurements from the other ungulates, we still found substantial overlap with red deer. Separation of moose from roe deer was complete in pellet length and width, albeit only by very small margins (0.02 mm and 0.22 mm respectively).
Comparison of morphometric measurements on 193 DNA-verified fecal pellet groups from four ungulate species. Mean values are denoted by red diamonds and statistically significant differences are indicated by different letters. The dotted blue line indicates a commonly used threshold for distinguishing between fallow and roe deer. Although moose and red deer pellets differ significantly in length, width, and pellet group size from each other as well as from the two smaller ungulates, there is too much overlap between species to allow for clear discrimination
Discriminant analysis based on the morphometric data assigned the correct species to 84% of the samples but performed significantly better than visual identification by field observers only in case of fallow deer (Fig. 4). Of the four ungulate species, only moose pellets were sufficiently well identified (> 95% correct) by either method. While discriminant analysis performed better than observers on fallow deer (χ2 = 6.48, df = 1, p = 0.01), it still only assigned 60% of pellet groups correctly.
Proportions of correctly identified ungulate pellets by observers in the field (dark gray bars) versus correctly identified pellets by discriminant analysis on morphometric measurements of DNA-verified fecal samples (DA, light gray bars). There are almost no differences between the two methods, except for fallow deer where DA performed significantly better
Effects of observer experience and season on species identification success
We found that species, experience, and their interaction had significant effects on species identification success. The effect was strongest for species (χ2 = 145.5, df = 3, p < 0.001), followed by experience (χ2 = 55.1, df = 1, p < 0.001) and the species*experience interaction (χ2 = 13.1, df = 3, p = 0.005).
Correct identification rates of red deer and fallow deer depended on observer experience (p-values < 0.0001, Table 2) and was higher for experienced observers. Moose was successfully distinguished from the smaller deer species, but these smaller species could not be separated well by field observers. The proportions of correct species identification for novice/experienced observer pairs were moose (98:97), roe deer (55:69), red deer (19:74), and fallow deer (48:83). The overall identification success (averaged across moose, red deer, fallow deer and roe deer) declined from spring (90%) to summer (73%) and autumn (74%) before increasing again in winter (83%) (Fig. 5).
Overall species identification success by season. The dots represent the mean and whiskers the standard error. The proportions of correctly identified pellets are averaged across the four species (moose, red deer, fallow deer, and roe deer) for each season. Identification success is high in winter and spring and declines during summer and autumn
Discussion
The results from this study highlight three key findings. First, the levels of accuracy obtained from dung morphometry raise concern about reliable discrimination between species in multi-species ungulate systems. Second, misidentification rates are highest between ungulates that are from the same family and of similar body size. Third, observer experience can improve identification success but cannot compensate for the fundamental problem of interspecific overlap in dung morphometry, especially between similarly sized species. These findings have a bearing on ecological studies as well as conservation and management practices that rely on visual identification of ungulate feces.
Species identification in the field and molecular validation
As expected, we found that in instances of misclassification, the error usually corresponded to the species most similar in size and physiology, e.g., bison to cattle or roe to fallow deer. The overall misclassification rate was highest for roe deer (41%), the smallest ungulate in our study. At the country level, roe deer misidentification was highest at the Kraansvlak study site in the Netherlands (78%) where it occurs in sympatry with the similarly sized fallow deer. At the German and Polish sites, where roe deer occur together with the substantially larger red deer, misidentifications were lower, 57% in Germany and 58% in Poland. These misidentification rates are almost identical despite differences in sample size of almost an order of magnitude between the two countries (see Table 1). Misidentification of roe deer was even lower in Sweden (29%) with the most common source of error being the confusion caused by fallow deer. We suspect that the lower error rate compared to Kraansvlak in the Netherlands stems from a possible smaller spatial overlap of the two species in Sweden. Nevertheless, the generally high misidentification rates for roe deer are likely a result of the large overlap in morphometrics between roe deer pellets and those of other deer species as illustrated in Fig. 3. At the Dutch Maashorst site, where roe deer represent the only cervid species in the ungulate assembly, their misidentification rate was zero. Similarly, bison and cattle were separated by fences in the area and thus accurately identified, much in contrast to the Kraansvlak site, where they occured sympatrically and were often mistaken for each other.
Red deer was generally identified correctly in the field with misidentification rates of zero in Poland and only 1% in Germany. In Sweden, where red deer occurs sympatrically not only with roe deer but several other deer species, i.e., moose, fallow deer, and reindeer, misidentification was higher at 16%. However, even in Poland and Germany, observers collected putative roe deer samples, which turned out to be red deer. This has some noteworthy implications. In case of pellet counts, it would cause an underestimation of red deer and an overestimation of roe deer numbers. This reflects concerns raised by Theuerkauf et al. (2008), who studied detectability of roe and red deer pellets in the Polish Bialowieza Forest, where some of the samples for our study were collected as well. Without the possibility of molecular verification, the authors tried to quantify their misidentification rates based on species sightings along pellet count transects. They concluded that misidentification bias would probably be small in case of the abundant red deer but possibly much more pronounced for the less common roe deer. They also thought it to be more likely that small red deer pellets would be misidentified as roe deer than vice versa. Our data corroborate these assumptions. If the objective of a study was species-specific, e.g., diet, samples collected as red deer at the Polish and German sites would be mostly correct while samples presumed to be roe deer would likely be mixed with more than 50% red deer, essentially rendering results meaningless. However, even in the case of red deer, the characteristics of the diet corresponding to the smaller pellets would be missed. Horse dung, with its almost unmistakable shape (“horse apples”), and moose, whose pellets resemble that of other deer but are markedly larger, had the lowest misidentification rates. The confusion between wild boar and deer dung, particularly red deer, appears to be not well documented in the literature and might be explained by seasonal diet choices. Especially during spring and early summer, when deer forage on soft, wet vegetation, their dung loses the distinct pellet shape and becomes clumped and amorphous. Alvarez (1994) reported season to have the strongest effect on changes in red deer and fallow deer dung morphology, but we could only find an Internet source (www.pestdetective.org.nz/culprits/red-deer, retrieved 2018 Nov 09) and an old German hunting textbook (Gesellschaft Forstmänner und Jäger 1796), which specifically addressed the similarities between red deer dung and that of wild boar during the vegetative season.
Occasionally, the data showed unexpected results. For example, two field-identified roe deer samples from the Netherlands turned out to be bison and cattle in the DNA results. Due to the very different dung morphologies between cervids and bovids, a true identification mistake is almost impossible. This leaves recording errors during the field sampling (i.e., the bovine dung samples were accidently entered as roe deer into the field protocol) or DNA cross-contamination as alternative explanations. In such cases, careful examination of the molecular data is necessary. In both instances, no other ungulate sequences than the assigned bovine species were detected in the sample, strongly indicating a recording error during field sampling. However, the risk of contamination is a serious threat due to the sensitivity of the PCR reaction that amplifies the DNA. Especially when working with degraded environmental DNA (eDNA), the problem can be exacerbated if the contaminant DNA is less degraded than the DNA in the target sample and thus amplifying better (Taberlet et al. 2018). We strongly suspected this to be the case with hare (Lepus sp.) DNA as detailed in Appendix A (Online Resource 1). As a consequence, we could unfortunately not quantify possible misidentification rates between hare and the smaller ungulates. We also detected moose in one German sample, which had been collected as wild boar. Moose occasionally occur in the area (Schönfeld 2009) and during the summer, when the sample in question was collected, frequently produce large, amorphous dung piles that could be mistaken for wild boar by observers not expecting the presence of moose. However, we also detected low levels of wild boar sequences (approximately 40x less abundant than moose) in the sample so some uncertainty remains. These examples serve to highlight the fact that DNA results are not free from artifacts and should be treated with appropriate caution. However, due to the strict removal of ambiguous samples, we are confident that the DNA results presented here are trustworthy and possible remaining annotation errors negligibly small.
Morphometric measurements
We found significant differences between four species in Sweden, but at the same time also too much overlap in the measurements to allow for sufficient discrimination. These results resemble the findings of Bowkett et al. (2013) who studied the dung morphometry of five sympatric species of differently sized forest antelopes in Tanzania. Roe deer and fallow deer differed significantly only in the number of pellets per group. Pellet shape (length/width) was the least informative of all morphometric variables.
Discriminant analysis and observer identification success in the field yielded very similar results, except in the case of fallow deer where discriminant analysis performed significantly better than observers. For red deer and fallow deer, the proportions of correct species assignment by discriminant analysis (80% and 60%) are comparable to values reported by Alvarez (1994) (81% and 78%), who used the same technique. Similarly, the range of correct species assignment we found (99% for moose to 60% for fallow deer) resembles the results of Costa et al. (2017) (92–56%) in a study of six sympatric neotropical deer species. The results from the discriminant analyses suggest that even an observer with perfect assessment of pellet morphometrics would still be prone to high rates of misidentification, especially for the smaller ungulate species.
As one would expect, distinguishing between the two smallest ungulates, fallow deer and roe deer, proved to be most difficult. While we were able to confirm that the commonly used partition threshold of 45 pellets per group is not unreasonable as it separated the means of both species, we nevertheless considered it to be insufficient for reliable separation of the species because approximately one third of samples for either species fell to the wrong side of the threshold. This mirrors the results of Yamashiro et al. (2013) who, using DNA analysis, was able to show that 36% of Japanese serow fell below the presumed to be reliable threshold of 200 pellets/group. Although the number of reindeer samples (n = 3) in the dung morphometry part of the study was too small to be included in the statistical analyses, their measurements (available as Online Resource 2) and the misidentification rate for all reindeer samples (29%, see Table 1) strongly indicate that in areas of overlap with red deer and roe deer, correct identification might be complicated further. We therefore concur with the conclusions reached by other authors who have studied similar multi-species ungulate systems (Costa et al. 2017; Bowkett et al. 2013; Ramón-Laca et al. 2014), in that dung morphometry is an inaccurate tool for distinguishing ungulate species of similar size.
Effects of observer experience and season on species identification success
Observer experience did not significantly affect identification success in moose and roe deer, but successful identification of red deer and fallow deer did depend on observer experience. Experienced observers identified red deer and fallow deer noticeably better than novices, 74% versus 19% and 83% versus 48% respectively. We could not, however, determine whether the more accurate assignation of pellets by experienced observers resulted from better interpretation of the dung morphometry, from greater familiarity with the local distribution of species or a combination thereof. It is conceivable, for example, that experienced observers were more certain about identifying dung based on their knowledge about local abundances of red and fallow deer. In case of the latter, they might therefore have less strictly adhered to the 45 pellets/group threshold. In addition, our observer experience results are mostly observational and lack some of the characteristics of strict experimental testing. We did not have a control group nor was the collection effort standardized per observer. For example, although we only compared observers who had collected all four species during roughly the same period in the same study area, there may still have been differences among observers in the habitat types searched as well as in ungulate densities and community composition in the transects they visited. Moreover, the novices received their initial training from different combinations of the experienced personnel. Bearing these caveats in mind, our results nevertheless suggest that observer experience can reduce misidentification errors, but they also indicate that even among experienced observers, misidentification rates for red deer and fallow deer are still high at a mean of 22%. We suggest to view these findings as exploratory and caution against generalization.
Across the seasons, we noticed a decline in the proportion of correctly identified pellets from spring (just after snowmelt) to summer and autumn. We attribute this largely to the circumstance that fecal pellets become more liquid and indistinct in shape during the growing season, owing to a less fibrous diet of fresh vegetation. These observations are in accordance with Alvarez (1994) who found seasonal differences to be the main source of variability in ungulate pellets. In addition, the influx of calves into the population during late spring and early summer further complicates species identification because their small pellets increase the interspecific overlap in dung morphometry.
Conclusion and recommendations
We found high rates of species misidentification in all of the four studied European multi-species ungulate systems. Errors were largest between ungulates of similar body size and physiology. Morphometric measurements of feces could not satisfactorily discriminate at the interspecific level but there were noteworthy species-specific differences. In Sweden, 95% of moose pellets were correctly identified and separated well from roe deer. For the time being, dung pellet counts aimed at monitoring moose likely continue to produce credible results. Within a multi-species management framework, the method may still perform well in areas where moose and roe deer co-occur but appears to be more limited when fallow and/or red deer are also present. In Germany and Poland, red deer were mostly identified correctly, but small red deer pellets were frequently mistaken as roe deer, which could bias population estimates or other species-specific studies using fecal samples. Horses, bison, and cattle could generally be well distinguished from the cervids but misidentification between wild boar and red deer occured in Poland and Germany during the growing season. If wild boar in Sweden continue expanding (Malmsten et al. 2017) into the range of brown bear, misidentification errors between the feces of these two omnivores might occur. For any study that relies on dung morphometry for the identification of ungulate species, sample collections should ideally be undertaken during early spring or winter when misidentification rates are lowest. Observer training and the use of ancillary local knowledge can improve results, but even experts may be subject to high error rates because they often lack verification of their pellet identifications in the field. For a comprehensive study of the efficacy of citizen scientists in dung counts, we recommend Buesching et al. (2014).
While our results highlight that reliance on dung morphometry for species identification can be problematic among similarly sized ungulates, they also indicate that misidentification errors are dependent on community composition and do not affect all species equally. The practical implications of our findings therefore depend largely on the objectives of researchers and managers. Pellet counts are a fast, simple, and inexpensive way of monitoring ungulates, especially on large scales. Our results suggest that they would still be reliable if focused on species with clearly distinguishable dung morphology (e.g., moose) or in areas where species with clearly distinct dung morphologies coexist (e.g., moose and roe deer). We hope that our results provide guidance on what levels of misidentification can be expected in European multi-species ungulate systems. This will aid researchers and managers in critically evaluating practices that rely on dung morphometry for species identification. If the dung composition is the main interest, as for example in diet studies, DNA testing should be employed to verify the species. If the monitoring of abundance or population dynamics is the objective, camera trapping could be considered as an alternative to pellet counts, particularly for smaller species like roe deer and fallow deer, which experience the highest rates of misidentification. While misidentification can also be a problem with camera trapping, the challenge typically lies in recognizing individuals rather than species. For example, Pfeffer et al. (2017), who studied an ungulate community of four deer species in Sweden using camera traps, could not identify the species in only 8% of the capture events.
References
Alvarez G (1994) Morphological variability and identification of deer pellets in Central Spain. Folia Zool 43:25–37
Apollonio M, Andersen R, Putman R (2010) Present status and future challenges for European ungulate management. In: Apollonio M, Andersen R, Putman R (eds) European ungulates and their management in the 21st century, vol 75. Cambridge University Press, Cambridge, p 474. https://doi.org/10.1016/S1616-5047(10)00101-1
Apollonio M, Belkin V, Borkowski J, Borodin O, Borowik T, Cagnacci F, Danilkin A, Danilov P, Faybich A, Ferretti F, Gaillard J, Hayward M, Heshtaut P, Heurich M, Hurynovich A, Kashtalyan A, Kerley G, Kjellander P, Kowalczyk R, Kozorez A, Matveytchuk S, Milner J, Mysterud A, Ozoliņš J, Panchenko D, Peters W, Podgórski T, Pokorny B, Rolandsen C, Ruusila V, Schmidt K, Sipko T, Veeroja R, Velihurau P, Yanuta G (2017) Challenges and science-based implications for modern management and conservation of European ungulate populations. Mammal Res 62:209–217. https://doi.org/10.1007/s13364-017-0321-5
Ball M (2010) Faecal pellet size can be used to differentiate age-classes in caribou: implications for non-invasive genetic studies. Conserv Genet Resour 2:239–241. https://doi.org/10.1007/s12686-010-9252-z
Bässler C, Förster B, Moning C, Müller J (2009) The BIOKLIM project: biodiversity research between climate change and wilding in a temperate montane forest—the conceptual framework. Waldökol Online 7:21–34
Bowkett A, Jones T, Laizzer R, Plowman A, Stevens J (2013) Can molecular data validate morphometric identification of faecal pellets in Tanzanian forest antelope species? Conserv Genet Resour 5:1095–1100. https://doi.org/10.1007/s12686-013-9935-3
Buesching CD, Newman C, Macdonald DW (2014) How dear are deer volunteers: the efficiency of monitoring deer using teams of volunteers to conduct pellet group counts. Oryx 48:593–601. https://doi.org/10.1017/S0030605313000227
Campbell D, Swanson G, Sales J (2004) Comparing the precision and cost-effectiveness of faecal pellet group count methods. J Appl Ecol 41:1185–1196
Chame M (2003) Terrestrial mammal feces: a morphometric summary and description. Mem Inst Oswaldo Cruz 98:71–94. https://doi.org/10.1590/S0074-02762003000900014
Costa EBV, De Oliveira ML, Peres PHDF, Grotta-Neto F, Vogliotti A, Piovezan U, Duarte JMB (2017) Low accuracy of identifying Neotropical deer species by scat morphology. Stud Neotrop Environ 52:37–42. https://doi.org/10.1080/01650521.2016.1263418
Cromsigt J, Rensburg S, Etienne R, Olff H (2009) Monitoring large herbivore diversity at different scales: comparing direct and indirect methods. Biodivers Conserv 18:1219–1231. https://doi.org/10.1007/s10531-008-9506-1
DeMay SM, Becker PA, Eidson CA, Rachlow JL, Johnson TR, Waits LP (2013) Evaluating DNA degradation rates in faecal pellets of the endangered pygmy rabbit. Mol Ecol Resour 13:654–662. https://doi.org/10.1111/1755-0998.12104
Edenius L (2012) Referensområden för klövviltförvaltning i södra Sverige: Ett projekt inom programområde Skog. Vilt, fisk & miljö, SLU, Umeå
Elliott HW, Longhurst WM (1984) Fecal pH of axis, fallow, and black-tailed deer. J Wildl Manag 48:659–662. https://doi.org/10.2307/3801214
Gesellschaft Forstmänner und Jäger (1796) Handbuch fuer die praktische Forst- und Jagdkunde. Schwickertscher Verlag, Leipzig
Giguet-Covex C, Pansu J, Arnaud F, Rey P-J, Griggo C, Gielly L, Domaizon I, Coissac E, David F, Choler P, Poulenard J, Taberlet P (2014) Long livestock farming history and human landscape shaping revealed by lake sediment DNA. Nat Commun 5. https://doi.org/10.1038/ncomms4211
Haesler H, Senn J (2012) Ungulate browsing on European silver fir Abies alba: the role of occasions, food shortage and diet preferences. Wildl Biol 18:67–74. https://doi.org/10.2981/09-013
Hemami M, Dolman P (2005) The disappearance of muntjac ( Muntiacus reevesi ) and roe deer ( Capreolus capreolus ) pellet groups in a pine forest of lowland England. Eur J Wildl Res 51:19–24. https://doi.org/10.1007/s10344-004-0067-7
Kohn MH, Wayne RK (1997) Facts from feces revisited. Trends Ecol Evol 12:223–227. https://doi.org/10.1016/S0169-5347(97)01050-1
Lenth RV (2016) Least-squares means: the R package lsmeans. J Stat Softw 69:1–33. https://doi.org/10.18637/jss.v069.i01
Lioy S, Braghiroli S, Dematteis A, Meneguz PG, Tizzani P (2015) Faecal pellet count method: some evaluations of dropping detectability for Capreolus capreolus Linnaeus, 1758 (Mammalia: Cervidae), Cervus elaphus Linnaeus, 1758 (Mammalia: Cervidae) and Lepus europaeus Pallas, 1778 (Mammalia: Leporidae). Ital J Zool 82:1–7. https://doi.org/10.1080/11250003.2014.963178
Lonsinger RC, Gese EM, Waits LP (2015) Evaluating the reliability of field identification and morphometric classifications for carnivore scats confirmed with genetic analysis. Wildl Soc Bull 39:593–602. https://doi.org/10.1002/wsb.549
MacCracken JG, Van Ballenberge V (1987) Age- and sex-related differences in fecal pellet dimensions of moose. J Wildl Manag 51:360–364. https://doi.org/10.2307/3801017
Malmsten A, Jansson G, Lundeheim N, Dalin A-M (2017) The reproductive pattern and potential of free ranging female wild boars (Sus scrofa) in Sweden. Acta Vet Scand 59:1–7. https://doi.org/10.1186/s13028-017-0321-0
Marques FFC, Buckland ST, Goffin D, Dixon CE, Borchers DL, Mayle BA, Peace AJ (2001) Estimating deer abundance from line transect surveys of dung: sika deer in southern Scotland. J Appl Ecol 38:349–363. https://doi.org/10.1046/j.1365-2664.2001.00584.x
Pfeffer SE, Spitzer R, Allen AM, Hofmeester TR, Ericsson G, Widemo F, Singh NJ, Cromsigt JPGM (2017) Pictures or pellets? Comparing camera trapping and dung counts as methods for estimating population densities of ungulates. Remote Sens Ecol Conserv 4:173–183. https://doi.org/10.1002/rse2.67
Plhal R, Kamler J, Drimaj M, Homolka J (2014) An assessment of the applicability of dung count to estimate the wild boar population density in a forest environment. J For Sci 60:174–180
Prokesova J, Barancekova M, Homolka M (2006) Density of red and roe deer and their distribution in relation to different habitat characteristics in a floodplain forest. Folia Zool 55:1–14
Putman RJ (1984) Facts from faeces. Mammal Rev 14:79–97
R Core Team (2017) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna
Ramón-Laca A, Gleeson D, Yockney I, Perry M, Nugent G, Forsyth DM (2014) Reliable discrimination of 10 ungulate species using high resolution melting analysis of faecal DNA. PLoS One 9:e92043. https://doi.org/10.1371/journal.pone.0092043
Rönnegård L, Sand H, Andrén H, Månsson J, Pehrson Å (2008) Evaluation of four methods used to estimate population density of moose Alces alces. Wildl Biol 14:358–371. https://doi.org/10.2981/0909-6396(2008)14[358:EOFMUT]2.0.CO;2
Rouco C, Delibes-Mateos M, Moreno S (2009) Evidence against the use of fecal pellet size for age determination in European wild rabbits. Acta Oecol 35:668–670. https://doi.org/10.1016/j.actao.2009.06.006
Schönfeld F (2009) Presence of moose ( Alces alces ) in southeastern Germany. Eur J Wildl Res 55:449–453. https://doi.org/10.1007/s10344-009-0272-5
Seton ET (1925) On the study of scatology. J Mammal 6:47–49. https://doi.org/10.2307/1373469
Taberlet P, Bonin A, Zinger L, Coissac E (2018) Environmental DNA: for biodiversity research and monitoring. Oxford University Press, Oxford
Theuerkauf J, Rouys S, Jedrzejewski W (2008) Detectability and disappearance of ungulate and hare faeces in a European temperate forest. Ann Zool Fenn 45:73–80
Tolleson DR, Randel RD, Stuth JW, Neuendorff DA (2005) Determination of sex and species in red and fallow deer by near infrared reflectance spectroscopy of the faeces. Small Rumin Res 57:141–150. https://doi.org/10.1016/j.smallrumres.2004.06.020
Tottewitz F, Stubbe C, Ahrens M, Dobiáš K, Goretzki J, Paustian K (1996) Counting droppings as a method of estimating the population of ruminant game. Z Jagdwiss 42:111–122. https://doi.org/10.1007/BF02240506
Venables WN, Ripley BD (2002) Modern applied statistics with S, 4th edn. Springer, New York
Woodruff SP, Johnson TR, Waits LP (2016) Examining the use of fecal pellet morphometry to differentiate age classes in Sonoran pronghorn. Wildl Biol 22:217–227. https://doi.org/10.2981/wlb.00209
Yamashiro A, Kamada M, Yamashiro T (2013) A comparative study of the fecal characters of Japanese serow (Capricornis crispus) and sika deer (Cervus nippon). Mamm Study 38:117–122. https://doi.org/10.3106/041.038.0207
Acknowledgements
We would like to thank all staff, students, and volunteers at the Swedish University of Agricultural Sciences (SLU), the Kraansvlak and Maashorst Nature Reserves (Netherlands), the Bavarian Forest National Park (Germany) and the Polish Mammal Research Institute in Bialowieza, who participated in the collection of fecal samples. The zoo in Lycksele (Västerbotten county, Sweden) kindly provided species-verified ungulate fecal pellets, and Jimmy Pettersson contributed ungulate tissue samples as positive controls. We are indebted to Delphine Rioux and Ludovic Gielly at the Laboratoire d’Ecologie Alpine (LECA) in Grenoble (France) for their assistance with PCR and purification. Eric Coissac and Frédéric Boyer (LECA, CNRS & University Grenoble-Alpes) helped with data processing and bioinformatics. We thank the two anonymous reviewers whose suggestions helped to improve and clarify this manuscript.
Funding
This study formed part of the research program “Beyond Moose – ecology and management of multispecies ungulate systems” and was financially supported by the Swedish Environmental Protection Agency (Naturvårdsverket, NV-01337-15), Kempestiftelserna (JCK-1514), the Swedish Association for Hunting and Wildlife Management and Västerbotten County’s Älgvårdsfonden (no. 218-9314-15). The work of Marcin Churski and part of the laboratory analysis of ungulate pellets was financed by National Science Center grant nr 2018/02/X/NZ8/01672.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Online Resource 1
(Appendix A): Molecular analyses, bioinformatics, and taxonomic annotation (DOCX 20 kb)
Online Resource 2
Summary statistics of dung morphometry measurements (XLSX 12 kb)
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Spitzer, R., Churski, M., Felton, A. et al. Doubting dung: eDNA reveals high rates of misidentification in diverse European ungulate communities. Eur J Wildl Res 65, 28 (2019). https://doi.org/10.1007/s10344-019-1264-8
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10344-019-1264-8