Abstract
Bacterial synthesis of polyhydroxybutyrates (PHBs) is a potential approach for producing biodegradable plastics. This study assessed the ability of Rhodopseudomonas palustris TIE-1 to produce PHBs under various conditions. We focused on photoautotrophy using a poised electrode (photoelectroautotrophy) or ferrous iron (photoferroautotrophy) as electron donors. Growth conditions were tested with either ammonium chloride or dinitrogen gas as the nitrogen source. Although TIE-1’s capacity to produce PHBs varied fairly under different conditions, photoelectroautotrophy and photoferroautotrophy showed the highest PHB electron yield and the highest specific PHB productivity, respectively. Gene expression analysis showed that there was no differential expression in PHB biosynthesis genes. This suggests that the variations in PHB accumulation might be post-transcriptionally regulated. This is the first study to systematically quantify the amount of PHB produced by a microbe via photoelectroautotrophy and photoferroautotrophy. This work could lead to sustainable bioproduction using abundant resources such as light, electricity, iron, and carbon dioxide.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Polyhydroxybutyrates (PHBs) are the most well-studied members of the polyhydroxyalkanoates (PHAs), which is a family of biodegradable intracellular polyesters produced by several bacteria [38, 45, 67, 79, 85]. Due to its thermoresistance, moldability, and biodegradability, PHB is a promising substitute for conventional petroleum-derived plastics [11]. Because of its biocompatibility, PHB is also used in many medical applications such as drug delivery, reconstructive surgery and bone tissue scaffolding [48]. However, its production is currently underexploited due to high feedstock costs [67]. Heterotrophic microbes can be promising PHB producers as they can use low-cost carbon sources including food wastes such as sugar beet, soy, and palm oil molasses [69]. However, the requirement for a continuous supply of food wastes makes them an infeasible source of carbon. Additional challenges of using food wastes are its sorting, transport, and pre-treatment prior to utilization [42, 54]. Lignocellulose from food [54] and forestry industries [41] or glycerol wastes from biofuel production have also been explored in heterotrophic PHB production [4, 75]. Some studies have used pure substrates such as glucose, acetate, and ethanol [79]. However, due to the requirement of arable land, and direct competition with human food consumption, using these substrates for PHB production is not desirable [17]. These potential limitations of using heterotrophs eventually led to the investigation of autotrophs for PHB production [38].
A handful of studies have demonstrated autotrophs as efficient PHB producers over heterotrophs [30]. A chemoautotrophic hydrogen-oxidizing bacterium Ideonella spp. strain O-1 has been shown to produce PHB using industrial exhaust gas containing hydrogen (H2), carbon dioxide (CO2), and carbon monoxide (CO). The exceptional ability of strain O-1 to grow even at CO concentration of 70% (v/v) without suppression of PHB production made it an attractive candidate to produce PHB using industrial exhaust gas (rich in CO) [76]. However, this route of PHB production may not be ideal because of the high cost of operation, and the risk of explosion associated with the use of H2 [14]. A sulfate-reducing bacterium Desulfococcus multivorans has also been shown to produce PHB [26], but its slow growth rate can make this process inefficient.
To make PHB bioproduction more efficient, economically viable and sustainable, research on autotrophic PHB production was further extended to photoautotrophs. The ability of photoautotrophs to use solar energy and CO2 for biosynthesis makes them unique candidates for efficient PHB synthesis [19, 34, 74]. Under photoautotrophic growth conditions, CO2 is fixed using energy harvested from light to generate ATP [9]. The fixed carbon can be used for the biosynthesis of acetyl-CoA, a substrate for PHB synthesis. In addition, photoautotrophs capable of fixing dinitrogen gas (N2) using ATP generated by photosynthesis are even more desirable. Moreover, nitrogen limitation has been reported to increase PHB accumulation [34]. Indeed, a recent study reported the suitability of the photoautotrophic organism Synechocystis sp. PCC 6714 as a potential host strain for PHB production [34]. However, the PHB amount based on cell mass and volumetric productivity was very low [19].
To produce higher PHB with greater efficiency, research on bacterial PHB synthesis was further expanded to microbial electrosynthesis (MES). This approach is based on the ability of some autotrophs (also called electroautotrophs) to acquire electrons from solid-phase conductive substances (SPCSs) such as electrodes using them as electron donors. This process of using SPCSs as electron donors or acceptors is termed “extracellular electron transfer (EET)” [20, 23, 36, 66]. When microbes use SPCSs as electron donors, the form of EET they use is also called microbial electron uptake (EU). This capability of electroautotrophs has been leveraged to produce value-added multi-carbon products via MES by reducing CO2 via either indirect or direct EU. In direct EU, microorganisms attach to the electrode and directly take up electrons from them [3, 6, 25, 52, 53, 65, 70, 78, 87, 88], whereas indirect EU involves transport of electrons by diffusible electron careers such as H2, formate or ammonia (either produced electrochemically or added to the reactors) from the electrode to microbes [13, 18, 28, 33, 39, 43, 58, 62, 68, 72, 80]. Indirect EU has been successfully used for PHB bioproduction by the chemoautotroph Cupriavidus necator (previously named Ralstonia eutropha). Nishio et al. reported that PHB productivity in C. necator was enhanced by EET using a biocompatible mediator (2-poly (2-methacryloyloxyethyl phosphorylcholine-co-vinylferrocene) (PMF) in an electrochemical system with an anode that was poised at + 0.6 V vs. the standard hydrogen electrode (SHE). Here, the anode served as an additional electron acceptor for microbial metabolism, resulting in acceleration of glycolysis and hence PHB synthesis [55]. Indirect MES of PHB by C. necator using formate as an electron carrier has also been reported recently [12]. To enhance CO2 assimilation by C. necator, a formate dehydrogenase (FDH)-assisted MES system was constructed, in which FDH catalyzed the reduction of CO2 to formate in the cathodic chamber. Formate served as the electron carrier to transfer electrons into C. necator generating PHBs [12]. The involvement of mediators in indirect EU lowers the efficiency of product formation. Direct EU in the context of MES is desirable because it omits the extra steps involved in indirect EU [12, 55]. Although a thermodynamic evaluation of bacterial PHB production via MES was proposed nearly a decade ago [64], no pure microbial culture has been known to produce PHBs using MES via direct EU (i.e., without the use of mediators).
Due to the abundance of iron on earth [24], PHB production linked to autotrophy using ferrous iron, Fe(II), as an electron donor could be leveraged for sustainable PHB production. Some Fe(II)-oxidizing chemoautotrophs such as Gallionella ferruginea have been reported to accumulate PHBs intracellularly. However, quantitative measurements on PHB production have not been reported for this organism [27, 46, 83]. The use of oxygen as the terminal electron acceptor by these organisms is a challenge because oxygen reacts readily with Fe(II) to oxidize and precipitates it to Fe(III). Therefore, G. ferruginea can only oxidize iron under low-oxygen concentrations [46]. Photoautotrophs such as purple bacteria and green sulfur bacteria have been shown to oxidize Fe(II) while fixing CO2 using light via a process called photoferroautotrophy [21, 73]. Photoferroautotrophs are more attractive for PHB synthesis because they oxidize Fe(II) in the absence of oxygen. However, thus far PHB accumulation has not been demonstrated during photoferroautotrophic growth.
Here, Rhodopseudomonas palustris TIE-1 was chosen as a platform for PHB bioproduction because it demonstrates extraordinary metabolic versatility [6, 32]. TIE-1 can grow chemoheterotrophically in rich medium as well as photoheterotrophically using various organic carbon sources [32]. It can also use several inorganic electron donors such as H2 and thiosulfate for photoautotrophic growth [32]. More importantly, TIE-1 is the only genetically tractable bacterium that has the ability to perform photoautotrophy using inorganic electron donors such as Fe(II) (photoferroautotrophy) and a poised electrode (photoelectroautotrophy) [6, 32]. TIE-1 performs direct EU to support photoelectrotrophy [6, 65]. These exceptional abilities make TIE-1 a very promising candidate to study photoautotrophic PHB production under different growth conditions. We assessed PHB production quantitatively on several growth conditions and found that TIE-1 can produce PHBs both photoautotrophically and photoheterotrophically. Among the photoautotrophic growth conditions, the highest PHB electron yield [percentage of the electron (mol) from the substrate that was converted into PHBs] was obtained under photoelectroautotrophy, and the highest specific PHB productivity was obtained under photoferroautotrophy. These novel routes of PHB synthesis by TIE-1 can potentially serve as a stepping stone for future bioengineering efforts towards sustainable PHB bioproduction.
Materials and methods
Bacterial strain, media, and growth conditions
Rhodopseudomonas palustris TIE-1 was originally isolated by Jiao et al. and has been used throughout this study [32]. For aerobic chemoheterotrophic growth, TIE-1 was routinely grown in 0.3% yeast extract and 0.3% peptone (YP) medium, with 10 mM MOPS [3-N (morpholino) propanesulfonic acid] at pH 7 in the dark at 30 °C with shaking at 250 rpm. For growth on solid medium, YP medium was solidified with 1.5% agar supplemented with 10 mM MOPS and 10 mM sodium succinate. For anaerobic photoautotrophic growth, TIE-1 was grown in anaerobic bicarbonate-buffered freshwater (FW) medium [21] supplemented with ammonium chloride (NH4Cl) (5.61 mM) or mixed N2/CO2 (80%/20%) gas at a pressure of 34.5 kPa as the sources of nitrogen. For anaerobic photoheterotrophic growth, 10 mL of FW medium was supplemented with anoxic 1 M stocks of sodium succinate, sodium butyrate and sodium 3-hydroxybutyrate to a final concentration of 1 mM in Balch tubes. However, to have higher biomass required for PHB, RNA and protein extraction, substrate concentrations were increased to 10 mM. A pre-grown TIE-1 culture with optical density (OD660) of 1 was inoculated with a final OD660 of 0.01 (100 × dilution) followed by incubation at 30°C in an environmental chamber fitted with infrared LED (880 nm). Time-course cell growth was monitored using Spectronic 200 (Thermo Fisher Scientific, USA). For photoautotrophic growth with H2 and Fe(II), TIE-1 was adapted to photoautotrophic growth using H2 as the sole electron donor as described previously [7]. For growth with Fe(II), 50 mL of FW medium was prepared under the flow of 34.5 kPa N2/CO2 (80%/20%) and dispensed into pre-sterilized serum bottles purged with 34.5 kPa N2/CO2 (80%/20%). The bottles were then sealed using sterile butyl rubber stoppers with aluminum crimp followed by the addition of anoxic sterile stocks of FeCl2 and nitrilotriacetic acid (NTA) to a final concentration of 5 mM and 10 mM, respectively. All sample manipulations were performed inside an anaerobic chamber with 5% H2/75% N2/20% CO2 (Coy laboratory, Grass Lake) [7]. The bacterial generation time was determined as described previously [77]. Lag time (lag) was determined as a period that precedes the exponential phase [47].
Cell enumeration
Samples were fixed with paraformaldehyde (20% v/v), transferred into Amicon centrifuge filters (Amicon Ultracel 100 k, regenerated cellulose membrane, Millipore, Carrigtwohill, CO, Ireland) and centrifuged for 10 min at 1000×g. The pellets were resuspended and washed twice in PBS (phosphate-buffered saline). The cells were recovered by centrifugation at 3000×g for 15 min. After the addition of PicoGreen® (Quant-iT PicoGreen® dsDNA, Life Technologies, Grand Island, NY, USA), the cells were counted in 96-well plates along with 50 μL of Sphero™ AccuCount blank beads (Spherotech, Lake Forest, IL, USA). Cell density was estimated with an LSRII flow cytometer (BD, Sparks, MD, USA) using a 488-nm laser. A calibration curve relating the ratio of cell events to bead events and the cell density was constructed using a serial dilution of a cell sample. Density was then determined by microscopy (Helber Bacteria Cell counting chamber with Thoma ruling, Hawksley, Lancing, Sussex, UK). The OD660 of TIE-1 cells vs. cell numbers were plotted to obtain a standard curve.
Bioelectrochemical setup and growth conditions
All photoelectroautotrophic experiments were performed using a three-electrode configured seal-type bioelectrochemical cell (BEC, C001 Seal Electrolytic cell, Xi’an Yima Opto-electrical Technology Com., Ltd, China). The three electrodes were configured as the working electrode (graphite rod, 3.2 cm2), reference electrode (Ag/AgCl in 3 M KCl) and counter electrode (Pt foil, 5 cm2). 70 mL of FW medium was dispensed into sterile BECs and made completely anaerobic by N2/CO2 (80%/20%) bubbling for 60 min with the final pressure maintained at ~ 50 kPa. 10 mL of TIE-1 cells (OD660 ~ 2.4) pre-grown in FW with H2 was then inoculated with a starting OD660 ~ 0.3 as described previously [65]. The OD660 of the inoculated BECs was monitored with a BugLab Handheld OD Scanner (Applikon Biotechnology, Inc., Foster City, CA). To evaluate the influence of NH4Cl and N2 gas as the nitrogen sources on PHB biosynthesis via photoelectroautotrophy, the BECs were operated simultaneously (c = 3 biological replicates) with NH4Cl and N2 gas as nitrogen sources with negative controls: open-circuit (OC) control (no current) and abiotic controls. The graphite electrode was constantly poised at a potential of + 100 mV vs. standard hydrogen electrode (SHE) for 130 h using a multichannel potentiostat (Interface 1000E, Gamry Multichannel Potentiostat, USA). All photoelectroautotrophic experiments were performed at 26 °C under continuous infrared light (880 nm) unless noted otherwise. At the end of the bioelectrochemical experiment, samples were immediately collected from the BEC reactors for RNA extraction and PHB production analysis as mentioned above.
Analytical procedures
PHB measurement From all the growth conditions tested, 10 mL of bacterial samples at an OD660 0.7 (unless stated otherwise) was pelleted at 8000×g for 10 min and stored at − 80 °C until PHB extraction and analysis were performed. 1 mL of water (LC–MS grade) and 600 µL of methanol (HPLC grade) were added to arrest metabolic activity of TIE-1. 10 mg/mL of poly[(R)-3-hydroxybutyric acid] (Sigma-Aldrich, USA) was used as a PHB standard. Extraction of PHB was followed by its conversion to crotonic acid. The concentration of crotonic acid was measured using an Agilent Technologies 6420 Triple Quad LC/MS as follows: using Hypercarb column, particle 5 µm, 100 × 2.1 mm (Thermo Fisher Scientific, USA) as stationary phase; water with 0.1% (v/v) formic acid as phase A; acetonitrile and 1% (v/v) formic acid as phase B. The injection volume was 5 µL; the flow rate was set at 500 µL min−1; the column temperature was set at 15 °C and the gas temperature was 300 °C [31]. PHB was detected as crotonic acid with mass to charge ratio (m/z) = 87 which was normalized to bacterial cell number. Details on PHB extraction, PHB carbon yield, and PHB electron yield calculations are described in supplemental methods.
H2and CO2measurement Time-course H2 and CO2 from photoautotrophic conditions were analyzed using gas chromatography (Shimadzu BID 2010-plus, equipped with Rt®-Silica BOND PLOT Column: 30 m × 0.32 mm; Restek, USA) with helium as a carrier gas. At each time point, 10 µL of gas was sampled from the headspace of the serum bottles using a Hamilton™ gas-tight syringe and injected into the column. To quantify dissolved CO2, 1 mL of filtered (using 0.22 µm PES membrane filter) aqueous samples from each reactor was collected and injected into helium-evacuated 12-mL septum-capped glass vials (Exetainer, Labco, Houston, TX, USA) containing 1 mL of 85% phosphoric acid. The concentration of the dissolved CO2 was then measured by injecting 10 µL of evolved CO2 in the headspace into the column. The total CO2 in the reactors was calculated as described previously [50].
Organic acid measurement Time-course consumption of organic acids such as sodium succinate, sodium butyrate, and sodium 3-hydroxybutyrate under photoheterotrophic conditions were quantified using an Ion Chromatography Metrohm 881 Compact Pro using a Metrosep organic acid column (250 mm length). 0.5 mM H2SO4 with 15% acetone was used as eluent at a flow rate of 0.4 mL min−1 with suppression (10 mM LiCl regenerant).
Fe(II) measurement Time-course Fe(II) concentration was measured using the Ferrozine Assay as described previously [7].
Total protein measurement Total protein during photoferroautotrophy was measured using trichloroacetic acid (TCA) precipitation as follows: total protein from 2 mL culture (at time point zero and at 192 h for the growth with NH4Cl and 360 h with N2 gas) in microcentrifuge tube was precipitated using 500 µL 100% TCA. This mixture was incubated for 10 min at 4 °C and centrifuged at 18,000×g for 30 min at 4 °C. The pellet was washed with 200 µL cold acetone and centrifuged at 18,000×g for 10 min at 4 °C. The pellet was then dried at 95 °C for 10 min to remove any residual acetone and resuspended in 50 µL HCl buffered with 100 mM Tris–Cl, pH 8.0. The BCA (bicinchoninic acid) Protein Assay Kit was employed using the microtiter plate method for protein estimation as specified by the manufacturer’s protocol following TCA precipitation (Thermo Scientific, Waltham, MA). Total protein was measured at an absorbance of 562 nm using the Biotek Synergy HTXmicrotiter plate reader [7]. For a total protein to OD660 conversion, total protein of known OD660 values of TIE-1 cells was quantified. A standard curve was obtained by plotting OD660 vs. total protein measured.
RNA extraction and sequencing
5 mL of bacterial culture were collected at an OD660 ~ 0.7. The RNA was stabilized using 5 mL RNAlater (Qiagen, USA) (buffer that stabilizes and protects RNA from degradation) and incubated at room temperature for 10 min. Bacterial cells were centrifuged at 5000 × g for 10 min and pellets were stored at − 80 °C until RNA extraction was performed. RNA extraction was performed using the RNeasy Mini Kit (Qiagen, USA) following the manufacturer’s protocol. DNA removal was performed using the Turbo DNA-free Treatment and Removal Kit (Ambion, USA). DNA contamination was tested using PCR using the primers listed in Table S1 as previously described [6, 7]. Illumina unpaired 150-bp libraries were prepared and sequenced at the Genome Technology Access Center, Washington University on an Illumina MiSeq platform (Illumina Inc., San Diego, CA, USA). Trimmomatic (version 0.36) was used to remove Illumina sequencing adapters, quality trim deteriorating bases (threshold = 20), and length filter (min = 60 bp) [5]. Preprocessed RNA-seq reads were mapped to the published R. palustris TIE-1 genome using TopHat2 (version 2.1.1) (https://genomebiology.biomedcentral.com/articles/10.1186/gb-2013-14-4r36) and the gff3 annotation file as a guide for sequence alignment. Bowtie 2 (version 2.3.3.1). (https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3322381/) was used to index the reference genome FASTA file. The number of reads mapping to each feature was counted by HTSeq (version 0.9.1). Differentially expressed genes were predicted in DESEQ 2 (version 1.16.1) using the HTSeq (https://www.ncbi.nlm.nih.gov/pubmed/25260700) read counts and an adjusted p value cutoff of 0.05. Heat maps were drawn in R using ggplot2 [44, 82].
Reverse transcription quantitative PCR analysis (RT-qPCR)
cDNA template was synthesized using the purified RNA samples using the iScript cDNA Synthesis Kit (Biorad, USA). Primers listed in (Table S2) were designed using primer3 software (http://bioinfo.ut.ee/primer3/). RT-qPCR was performed using Biorad CFX connect Real-Time System Model # Optics ModuleA using the following thermal cycling conditions: 1 cycle at 95 °C for 3 min and 30 cycles of 95 °C for 3 s, 60 °C for 3 min, and 65 °C for 5 s according to the manufacturer’s protocol. Fold change comparison and standard deviation calculations were performed as described previously [2].
Identification of PHB cycle genes of TIE-1
The available TIE-1 genome in the JGI Genome Portal (https://genome.jgi.doe.gov/) was used to search for homolog genes involved in the PHB cycle using Blast search.
Scanning transmission electron microscopy–electron energy loss spectroscopy (STEM–EELS)
TIE-1 grown under sodium butyrate, Fe(II)–NTA and poised graphite electrode was used as representative samples for STEM-EELS. Briefly, 5 mL planktonic cell suspensions were centrifuged at 6000×g for 5 min. followed primary fixation by resuspending the cells pellets in 2% formaldehyde and 2.5% glutaraldehyde in 0.05 M sodium cacodylate buffer (pH 7.2) for ~ 45 min at room temperature. After agar encapsulation followed by primary fixation for ~ 20 min, agar cubes were subjected to secondary fixation for ~ 5 h followed by acetone dehydration and resin infiltration. Ultrathin sections (~ 50–60 nm) were obtained using Reichert Ultracut UCT ultramicrotome (Donald Danforth Plant Science Center, Saint Louis, MO), then mounted directly on amorphous-carbon film-coated TEM Cu-grids. Intracellularly localized PHB granules were characterized using a JEOL JEM-2100F field emission scanning transmission electron microscopy (FE-STEM) with an accelerating voltage of 200 keV (Institute of Material Science and Engineering, WUSTL); the microscope is attached with a Gatan 805 BF/DF detector, Gatan 806 HAADF detector and Gatan 863 Tridiem imaging filter (GIF) system. Images were obtained in STEM mode using HAADF detector and BF detector. EELS spectral images were acquired through working HAADF and GIF jointly. Carbon-K edge and nitrogen-K edge elemental maps were retrieved from STEM–EELS spectral images.
Statistical analysis
The P values were determined by one-way ANOVA followed by a pairwise test with Bonferroni adjustment. For the pairwise test with Bonferroni adjustment, the cutoff P value is equal to 0.025
Nucleotide sequence accession numbers
All RNAseq datasets have been deposited in NCBI under BioProject accession number PRJNA417278.
Results and discussion
Rhodopseudomonas palustris TIE-1 possesses putative PHB cycle genes
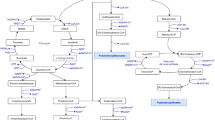
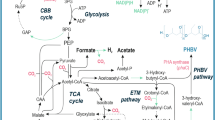
PHB production has been previously reported by several R. palustris strains [15, 51, 84]. However, the PHB cycle genes have not been explored thus far. This lack of information critically limits the potential prospects for future bioengineering efforts for PHB bioproduction. Availability of the TIE-1 genome allowed us to identify the genes that are homologous to the PHB cycle genes of C. necator [59, 60, 81], an H2-oxidizing betaproteobacterium that is known to produce and sequester PHAs intracellularly [8]. Reconstruction of the PHB cycle in E. coli using PHB genes from C. necator has been previously used to elucidate the biochemical pathway of PHB production and its subsequent metabolism [81]. Briefly, the pathway starts with the condensation of two acetyl-CoAs into acetoacetyl-CoA, a reaction driven by the β-ketothiolase enzyme, PhaA [81] (Fig. 1a). Acetoacetyl-CoA gets reduced to (R)-3-hydroxybutyryl-CoA by the enzyme acetoacetyl-CoA reductase, PhaB. Eventually, PHB polymerization is achieved by the PHB polymerase, PhaC (Fig. 1a). Interestingly, PHB can also serve as an important source of carbon and energy during environmental stress, where PHB molecules are catabolized by the PHB depolymerase, PhaZ (Fig. 1a) [81]. In Bradyrhizobium diazoefficiens, a nitrogen-fixing symbiont closely related to TIE-1, PhaR represses the expression of phaC1 and phaC2. In addition, PhaR regulates PhaP, the phasin protein that binds to and controls the number and size of the PHB granules. PhaR also binds to PHB granules and dissociates from it as the granule size grows [63]. TIE-1 encodes one phaR gene (Rpal_0531); one phaZ gene (Rpal_0578); multiple copies of phaA and phaB; two copies of phaC (Rpal_2780 and Rpal_4722); and three genes for phaP2 (Rpal_4291, 4616 and 4617) (Fig. 1a, b). A phaB gene (Rpal_0533) is located next to the phaA gene (Rpal_0532) (Fig. 1b) forming a putative operon (Fig. 1b). The gene for phaR (Rpal_0531) is positioned next to the phaA (Rpal_0532) gene in this operon but expressed from the opposite strand. This shows that TIE-1 possesses all the necessary genes for both PHB biosynthesis, polymerization and depolymerization.
The PHB cycle and its putative genes in TIE-1. a Putative PHB cycle of TIE-1. Acetyl-CoA is produced using multiple carbon sources such as organic carbon or carbon dioxide (CO2) followed by the condensation of two acetyl-CoAs into acetoacetyl-CoA. Acetoacetyl-CoA gets reduced to (R)-3-hydroxybutyryl-CoA by the enzyme acetoacetyl-CoA reductase, PhaB. (R)-3-Hydroxybutyryl-CoA is eventually polymerized into PHB granules by PhaC1 and/or PhaC2. During carbon storage mobilization, PHB is degraded back to (R)-3-hydroxybutyryl-CoA, and then to acetyl-CoA by multiple enzymatic reactions (shown by the double white lines). b Organization of the genes involved in the putative PHB cycle in TIE-1. Adapted from [81]
TIE-1 produces PHB under photoautotrophic conditions using different electron donors
Photoautotrophic PHB production using H2 as an electron donor
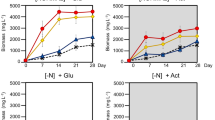
TIE-1 was grown with H2 as the sole electron donor with N2 and NH4Cl as fixed nitrogen sources. We call the N2 fixing conditions as the electron donor–N2 system and the NH4Cl conditions as the electron donor–NH4Cl system throughout. TIE-1 showed a higher maximum OD660 of 1.16 (P ≤ 0.001, Table 1) with NH4Cl (H2–NH4Cl system) compared to OD660 of 0.54 with N2 (H2–N2 system). This growth defect observed in H2–N2 was also reflected in the longer generation time of 41 h in the H2–N2 system compared to 34 h in the H2–NH4Cl system (P =0.005, Table 1). Slow growth under N2 fixing conditions was previously observed in R. palustris strain 42 OL [15] and is likely due to the high-energy requirement of this process. TIE-1 showed a lower PHB carbon yield [percentage of carbon (mol) from the substrate that was converted into PHBs] of 2.55% in the H2–N2 system compared to the ~ 3 times higher yield of 7.23% in the H2–NH4Cl system. In contrast, no significant difference was observed in PHB electron yield [percentage of the electron (mol) from the substrate that was converted into PHBs] between the H2–N2 system and the H2–NH4Cl system (Fig. 2a, Table 2). Interestingly, the specific PHB productivity almost doubled in the H2–N2 system compared to the H2–NH4Cl system (from 1.30 × 10−14 to 3.08 × 10−14 mg/L/Cell/h) (Table 2). The higher specific productivity under N2 fixing conditions and the growth defect (Table 1) indicate a direct impact of stress caused by the high-energy-consuming N2 fixation process. This stress might have induced TIE-1 to accumulate intracellular PHBs as a carbon and energy reserve. Similar observations have been reported for other purple non-sulfur bacteria [29] grown under nitrogen-limited conditions, including R. palustris strains grown in nitrogen-deprived conditions [51]. Increased accumulation of PHB to over 30% of the dry cell weight was observed previously when R. palustris CGA009 cells were N2 starved [51]. Because H2 is known to be produced during N2 deprivation by CGA009 [51], this H2 could serve as an additional electron donor for photoautotrophy, accounting for additional PHB accumulation. Further studies will be required to test whether this is happening in CGA009 and TIE-1. Although the growth with NH4Cl had N2 gas in the headspace under most conditions in our experiment, the presence of NH4Cl has been known to inhibit nitrogenase gene expression, preventing N2 fixation from occurring in the presence of NH4Cl [40].
PHB carbon yield by TIE-1 grown in freshwater (FW) medium with ammonium chloride (NH4Cl) or under N2 fixing conditions (N2 gas). a Photoautotrophic conditions with H2, Fe(II) (photoferroautotrophy) or poised electrodes (photoelectroautotrophy) as electron donors; b photoheterotrophic conditions with succinate, butyrate or 3-hydroxybutyrate as electron donors. Error bars are from the standard deviations calculated using 2–3 biological replicates as specified in Table 2. The P values were determined by one-way ANOVA followed by a pairwise test with Bonferroni adjustment; ns not significant. For a pairwise test, the cutoff P value is 0.025. P values are indicated in Tables 2, S5
Photoautotrophic PHB production using a poised electrode as an electron donor
To evaluate PHB production under photoelectroautotrophy using poised electrodes as the sole electron donor, graphite electrodes were poised at a potential + 100 mV vs. SHE to mimic the Fe(OH)3/Fe2+ redox couple. Although the electrode–NH4Cl system resulted in higher maximum OD660 compared to the electrode–N2 system (0.73 vs. 0.61, respectively) (Table 1), there was no significant difference in the generation time of TIE-1 between these two conditions (Fig. 3a; Table 1). Nonetheless, after 96 h of incubation, EU by TIE-1 in the electrode–NH4Cl system was 1.92 µA/cm2, about double of that obtained in the electrode–N2 with an EU of 0.93 µA/cm2 (Fig. 3b, Table S6, P ≤ 0.01). Despite the difference in the total EU, no notable change was observed in the PHB carbon yield, PHB electron yield and specific PHB productivity between the electrode–NH4Cl system and the electrode–N2 system (Fig. 2a; Table 2). This result could be due to the continuous supply of electrons from a poised electrode, which did not seem to directly impact PHB biosynthesis. As expected, no cell growth was observed in the biotic reactors with unpoised electrodes (open circuit, OC) with both NH4Cl and N2 gas (Fig. 3a). Abiotic controls did not show any EU (Fig. 3b). Bioelectrosynthesis of PHB has been recently reported via a method using enzymatic and electrochemical approaches. A modified electrode poised at − 386 mV vs. SHE was used to synthesize NADH in the presence of enzymes of the PHB cycle to convert acetate to PHB. The amount of PHB produced was 0.3 mg/L (under a maximum current density (Jmax) of 27.9 ± 1.3 μA cm−2) [1]. In another study, 226 ± 6 mg/L of PHB was produced via indirect EU using formate as a mediator by C. necator (Ralstonia eutropha) at − 395 mV vs. SHE (~ Jmax 213 μA cm−2) [12]. In addition, overexpression of the ruBisCO (Ribulose-1,5-bisphosphate carboxylase/oxygenase) gene was performed to increase the CO2 fixation [12]. As a result, PHB production was enhanced to 485 mg/L [12]. However, both of these approaches involve multiple steps including enzyme purification. They also operate at a higher reduction potential than what we use to grow TIE-1 for PHB production (+ 100 mV vs. SHE, Jmax of only<2 μA cm−2). Although our results show a low (~ 5–6 mg/L) amount of PHB production under photoelectroautotrophy via direct MES, our approach offers numerous advantages: (1) using direct EU, we minimize the complexity of the MES system; (2) TIE-1 grows at a lower reduction potential than that used in the studies above. This lower reduction potential ultimately saves electrical energy; and (3) TIE-1 is a photoautotroph and, therefore, can use the energy of light to make excess ATP for biosynthesis. The photoautotrophic ability of TIE-1 makes is especially attractive for sustainable PHB bioproduction because light is an abundant resource. The major hurdle for using TIE-1 for bioproduction is the low electron uptake it demonstrates from graphite electrodes. This is reflected in the lower maximum current density (Jmax) values that were observed here in our study. Improving electron uptake would increase Jmax values, which would also increase bioproduction [65]. A previous study from our laboratory has shown that inexpensive electrode modifications such as coating the electrodes with Prussian Blue can enhance electron uptake in the absence of a mediator via direct EU [65]. We are pursuing reactor design and electrode modifications further to enhance direct EU by TIE-1 because that will ultimately improve product formation.
Photoautotrophic PHB production using ferrous iron as an electron donor
Specific PHB productivity was higher in the H2–N2 compared to the H2–NH4Cl systems (Table 2) but no difference was observed under photoelectroautotrophy. However, under photoferroautotrophy, the specific PHB productivity was higher in the Fe(II)–NH4Cl system compared to the Fe(II)–N2 (Table 2). In addition, PHB carbon yield decreased in the Fe(II)–N2 system compared to the Fe(II)–NH4Cl system (Fig. 2a; Table 2). By carefully examining Fe(II) oxidation in Fe(II)–NH4Cl system, a significant drop in Fe(II) concentration via microbial Fe(II) oxidation was observed during the first 96 h, and Fe(II) was completely oxidized by 384 h (Fig. 4a). In contrast, the Fe(II)–N2 system showed a very slow Fe(II) oxidation where a significant drop in Fe(II) concentration was observed only after 360 h (Fig. 4b). In addition, there was a decrease in the maximum OD660 in the Fe(II)–N2 system compared to the Fe(II)–NH4Cl system (Table 1). Additionally, total protein concentration with the Fe(II)–NH4Cl nearly doubled after 192 h whereas, with the Fe(II)–N2, the total protein concentration doubled only after 360 h (Fig. 4a, b). Based on these results, it is plausible that the growth defect in the Fe(II)–N2 system is a consequence of the high-energy demand during N2 fixation (Table 1; Fig. 4).
a TIE-1 growth during photoelectroautotrophy. OD660 values of TIE-1 grown under photoelectroautotrophy in freshwater medium (FW) with NH4Cl and under N2 fixing conditions with their respective control with unpoised electrodes (open circuit, OC). Error bars are from the standard deviations calculated using three biological replicates. b Current density during photoelectroautotrophy. Current density (µA/cm2) from TIE-1 grown with freshwater medium with NH4Cl as a nitrogen source or with N2 gas as a nitrogen source using a poised electrode at a potential of + 100 mV vs. Standard hydrogen electrode (SHE) and the associated abiotic control. The negative sign on the Y-axis indicates current uptake. Error bars are the standard deviations calculated using two biological replicates. The P values were determined by one-way ANOVA followed by a pairwise test with Bonferroni adjustment; ns not significant. For a pairwise test, the cutoff P value is 0.025. P values are indicated in Table S5
Growth of TIE-1 under photoferroautotrophic conditions. Fe(II) oxidation and protein concentration measured during photoferroautotrophy a with NH4Cl as a nitrogen source, and b with N2 gas as a nitrogen source. Error bars are from the standard deviations calculated using three biological replicates. The P values were determined by one-way ANOVA followed by a pairwise test with Bonferroni adjustment; ns not significant. For a pairwise test, the cutoff P value is 0.025). P values are indicated in Table S5
Comparison of PHB production under photoautotrophic conditions
Among the three electron donors tested, photoautotrophic growth with H2 showed the highest maximum OD660 when grown with NH4Cl (Table 1, P ≤ 0.001). Although the Fe(II)–NH4Cl and the Fe(II)–N2 conditions resulted in the lowest maximum OD660 (Table 1, P ≤ 0.001), the PHB carbon yield under the Fe(II)–NH4Cl is comparable to that obtained from photoautotrophy in the H2–NH4Cl system (Table 2, P =0.0613) and higher than under the electrode–NH4Cl (Table 2, P = 0.028). Moreover, the specific PHB productivity obtained in the Fe(II)–NH4Cl is the highest amongst all the photoautotrophic conditions with NH4Cl (Table 2, P ≤ 0.001). The electrode–N2 system showed the longest generation times amongst all the photoautotrophic growth conditions (Table 1, P ≤ 0.001). Photoelectroautotrophy showed the lowest PHB carbon yield both with NH4Cl and N2 (Fig. 2a, P = 0.0045). Nevertheless, TIE-1 showed the highest efficiency in converting electrons to PHB in the electrode–NH4Cl system (4.39% PHB electron yield) and in the electrode–N2 system (7.34% PHB electron yield) (Table 2, P ≤ 0.001). These results indicate that although the growth of TIE-1 during photoelectroautotrophy was slow, this condition was the most efficient at converting electrons obtained from a poised electrode into PHB. A previous comparative growth study of TIE-1 using H2 and soluble Fe(II) as electron donors revealed H2 as a preferred electron donor over Fe(II) [32]. This preference was reflected in the lower PHB carbon yield during photoferroautotrophy vs. photoautotrophy with H2 (Fig. 2; Table 2).
Effect of N2 fixation on photoautotrophic PHB production
The effect of N2 fixation was clearly observed in both H2 and Fe(II) systems, where cell growth and the PHB carbon yield was significantly reduced under N2 fixing conditions (Tables 1, 2). Based on previous studies, it is likely that the stress caused by the high-energy N2 fixation process led to the accumulation of PHB by TIE-1 [29, 51]. However, N2 fixation did not have a significant impact on the growth of TIE-1, the specific PHB productivity and the PHB carbon yield under photoelectroautotrophy (Table 2). A continuous electron supply might have allowed TIE-1 to fix N2 to ammonium without affecting the supply of electrons to produce PHBs. Interestingly, PHB biosynthesis was proposed to be a potential electron sink when Rhodopseudomonas palustris CGA009 was incubated in the presence of argon (nitrogen deprived) [51]. However, our results show that only a small percentage of electrons go to PHB biosynthesis under all N2 fixing conditions (Table 2). In the electrode–N2 system, where the maximum PHB electron yield was obtained, only 7.34% of the available electrons contributed to PHB biosynthesis.
TIE-1 produces PHB under chemoheterotrophic and photoheterotrophic conditions
Chemoheterotrophic PHB production
To further investigate the effect of different media on PHB production, TIE-1 was grown chemoheterotrophically (aerobic) with rich media containing yeast extract and peptone (YP). Aerobic growth of TIE-1 on YP resulted in the longest generation time (g = 11.18 h) compared to the generation time observed from all the other photoheterotrophic conditions (Table 1, P ≤ 0.001). YP-grown cells produced the highest specific PHB productivity of 24.60 × 10−14 mg/L/Cell/h compared to all the conditions tested (Table 2, P ≤ 0.001). It is likely that the amino acids provided by peptone contributed to the increased PHB production by TIE-1 under this condition. The availability of amino acids precludes their de novo biosynthesis, and peptone has been previously reported to increase PHB production in Azotobacter vinelandii [56].
Photoheterotrophic PHB production with succinate
To test the effect of different carbon sources with different oxidation/reduction values (Supplemental Table S4) on PHB production, TIE-1 was further grown photoheterotrophically (anaerobic) using three different substrates: succinate, butyrate, and 3-hydroxybutyrate with NH4Cl and N2 gas. Similar to photoautotrophy, use of N2 gas as the source of nitrogen resulted in longer lag time, longer generation time and a longer time to reach maximum OD660 compared to NH4Cl as the nitrogen source (Table 1). However, there was no significant difference in specific PHB productivity between the succinate–N2 system and the succinate–NH4Cl system (Table 2). Rather there was an increase in the PHB carbon yield and PHB electron yield in the succinate–N2 system compared to the succinate–NH4Cl system (Fig. 2b; Table 2) indicating that there was no significant effect of N2 fixation on PHB productivity (Table 2).
Photoheterotrophic PHB production with butyrate and 3-hydroxybutyrate
Interestingly, TIE-1 grown photoheterotrophically with the less oxidized substrate, butyrate, showed a decrease in maximum OD660 (0.42) in the butyrate–NH4Cl compared to 0.69 with the butyrate–N2 systems (Table 1). An increase of about ninefold in PHB carbon yield was obtained in the butyrate–N2 system compared to the butyrate–NH4Cl system (Fig. 2b; Table 2). Similarly, specific PHB productivity in the butyrate–N2 system was more than three times higher compared to the butyrate–NH4Cl system (Table 2). The highest PHB production (~ 17.1 mg/L), as well as the highest PHB carbon yield, was obtained in the butyrate–NH4Cl system (Table 2). Although this PHB production is lower than the PHB production previously reported by another photosynthetic purple bacterium, Rhodobacter sphaeroides (60 mg/L) grown in olive mill wastewater under N2-limited conditions [22], TIE-1’s ability to produce PHB under various conditions such as photoautotrophy offers an obvious advantage when considering mixotrophic (photoheterotrophy and photoautotrophy) growth conditions for PHB production. Butyrate has been previously reported to be a preferred substrate over acetate in a PHB-producing mixed culture dominated by Plasticicumulans acidivorans due to the lower ATP need for PHB production using butyrate [49]. In addition, a study performed on C. necator has shown that butyrate is metabolized into 3-hydroxybutyryl-CoA via the beta-oxidation pathway. 3-Hydroxybutyryl-CoA is a direct precursor for PHB biosynthesis [16]. This shorter pathway might explain the higher PHB production along with faster generation time and a higher maximum OD660 obtained from butyrate compared to succinate and 3-hydroxybutyrate (Table 1; Table 2). Moreover, the higher numbers of electrons in butyrate compared to succinate and 3-hydroxybutyrate could contribute to higher PHB production in TIE-1 [29, 51]. A previous study by Shi et al. also reported similar results using metabolic flux balance analysis of PHB biosynthesis by C. necator under nitrogen-limited conditions using butyrate [71].
When TIE-1 was grown photoheterotrophically in 3-hydroxybutyrate, shorter lag time was observed in the 3-hydroxybutyrate–NH4Cl compared to the 3-hydroxybutyrate–N2 (Table 1). Although the maximum OD660 values were similar, N2 fixing conditions significantly increased the time to reach the maximum OD660 values (Table 1). A decrease in PHB electron yield was observed when TIE-1 was grown in the 3-hydroxybutyrate–NH4Cl system compared to the 3-hydroxybutyrate–N2 (Table 2). However, no significant differences were observed in the PHB carbon yield and specific productivity under these conditions (Table 2). These results indicate that N2 fixation slowed the growth of TIE-1 in 3-hydroxybutyrate but increased the PHB electron yield similar to the results obtained under other heterotrophic growth conditions (Table 1; Table 2).
Effect of N2 fixation on photoheterotrophic PHB production
Overall, N2 fixing conditions during photoheterotrophy delayed cell growth and resulted in a longer lag time as well as a longer time to achieve maximum OD660 (two times or longer) (Table 1). In contrast, PHB carbon yield under photoheterotrophy under N2 fixing conditions was higher than with NH4Cl (Table 2). This increase in PHB production under N2 fixing or N2 deprivation conditions is consistent with previous findings [29, 51]. This trend was not observed under photoautotrophic conditions. Interestingly, our results show that only a small percentage of carbon from the different substrates contributes to PHB synthesis by TIE-1. For example, the maximum PHB carbon yield during photoheteroautotrophic growth with butyrate was 8.81%. This indicates that the remaining carbon is likely used for biomass production. A previous report has shown that R. palustris grown on acetate converts 93% of its carbon into biomass [50].
STEM–EELS confirms intracellular accumulation of PHB granules in TIE-1
Because the LC–MS method used for PHB quantification involves digestion of the PHB polymer into crotonic acid [35], it was necessary to confirm the presence of intracellular PHB granules in TIE-1 using an additional technique. Nile red staining has previously been used to screen for PHB and fatty acid esters in bacteria [61]. However, this staining technique was ineffective in showing any intracellular inclusion bodies in TIE-1 possibly due to its small size. Hence, scanning transmission electron microscopy-electron energy loss spectroscopy (STEM–EELS) under conditions that produced the highest accumulation of PHB (mg/L/Cell) was performed. All the conditions imaged contained NH4Cl. The intracellular localization of PHB granules was confirmed by the carbon and nitrogen maps (Fig. 5). The nitrogen signal is likely from the phasin protein that is known to bind PHB granules [63]. PHB mostly aggregated as small multiple granules under photoelectroautotrophy compared to larger granules under photoferroautotrophy and photoheterotrophy with butyrate (Fig. 5). A change in the number and morphology of PHB granules was also observed previously in an anoxygenic phototrophic bacteria Dinoroseobacter sp. JL1447 when it was grown with different carbon sources such as sodium acetate, glucose, sodium glutamate, sodium pyruvate, and trisodium citrate [85]. Moreover, in a study on a purple non-sulfur bacterium Rhodovulum visakhapatnamense, a change in size and an increase in the number of PHB granules were also observed under nitrogen stress [29].
STEM–EELS images of TIE-1 grown photoheterotrophically with butyrate, photoferroautotrophically with Fe(II) and photoelectroautotrophically with a poised electrode. From top to bottom panel: bright-field image, carbon, and nitrogen map, and the composite images. Bright areas represent the dominance of the corresponding element (carbon, and nitrogen). The red background in the composite images is due to carbon signals from Spurr’s resin used for embedding the cells during sample preparation. The scale bars are 0.2 μm
PHB cycle genes are not differentially expressed
Our data show that there is significant variation in PHB production by TIE-1 under different growth conditions (Fig. 2; Table 2). To determine if genes involved in PHB production are transcriptionally regulated, transcript levels of the genes involved in the PHB cycle was assessed using RNA-Seq and reverse transcription quantitative PCR (RT-qPCR). PHB cycle genes were chosen and are summarized in Fig. 1a. RNA-Seq analysis showed that the PHB cycle genes were not differentially expressed with respect to growth conditions or levels of PHB (values having p > 0.05 are statistically not significant) (Fig. 6, Tables S7–S10). RT-QPCR was performed to corroborate these data. For this analysis, we chose one phaA isozyme (Rpal_0532) of many because it showed the highest expression in the RNAseq data. This gene exists in an operon with a phaB homolog (Rpal_0533) (Fig. 1b), which was the only phaB homolog that was chosen for RT-qPCR analysis. The other PHB cycle genes only had 1–2 representatives in the TIE-1 chromosome and these were all analyzed using RT-qPCR. RNAseq analysis combined with RT-qPCR analysis of these phaAB homologs (Table S11) revealed no significant changes in gene expression (Figure S1; Tables S8, S9). Moreover, the analysis of phaC1 and phaC2 polymerase genes did not show any significant differential expression (Fig. 6, Tables S7, S10). McKinlay et al. reported that R. palustris CG009 when incubated photoheterotrophically with acetate, under N2 deprivation (with argon), accumulated PHBs with no change in transcript levels of genes known to be involved in PHB biosynthesis [51].
Heat map showing log2 fold change in the expression of PHB genes from RNA sequencing analysis (RNASeq). Results are from TIE-1 grown in freshwater (FW) medium photoheterotrophically with butyrate, photoautotrophically with H2, photoferroautotrophically with Fe(II), and photoelectroautotrophically using a poised electrode. Growth with NH4Cl is indicated by NH4Cl whereas growth under N2 fixing conditions is indicated by N2. Error bars are the standard deviations calculated using three biological replicates. The P values were determined by one-way ANOVA followed by a pairwise test with Bonferroni adjustment; ns not significant. For a pairwise test, the cutoff P value is 0.025). P values are indicated in Table S7–9
Phasin proteins have been reported to play a significant role in the PHB cycle [86]. Deletion of the phaP gene reduced PHB production significantly in C. necator [86]. Here, no significant upregulation of phaP2 genes was observed using RNAseq even during photoheterotrophic growth with the butyrate–N2 system, where the highest PHB production in mg/L/cell was obtained (Supplemental Table S7). Although phaP2a appears to show a slight upregulation of 2.33-fold change under photoelectroautotrophy with N2 fixing conditions, the P value was more than 0.05, rendering it not significant (Fig. 6, Table S7). Gene expression analysis results during photoautotrophy and photoheterotrophy using both RNAseq and RT-qPCR show that there is no differential expression in the genes involved in the PHB cycle. This could suggest that in TIE-1 (and perhaps even CGA009), the variation in PHB accumulation might be regulated post-transcriptionally. These findings are useful for the further optimization of PHB production using TIE-1 [45].
Conclusions and future perspectives
Our study demonstrates the ability of a metabolically versatile photoautotroph Rhodopseudomonas palustris TIE-1 to produce PHB intracellularly under various growth conditions using different electron donors. The novel photoautotrophic metabolism using a poised electrode as the source of electrons produced the highest PHB electron yield. Another key discovery of this study is the ability of TIE-1 to yield the highest specific PHB productivity using Fe(II) as an electron donor for photoautotrophy. In summary, these newly described routes can serve as potential substitutes for PHB bioproduction. The application of these novel approaches can be especially important in areas where organic carbon sources are limited while resources such as light, CO2 [37], iron, and electricity [57] are abundant. TIE-1’s ability to fix N2 gas photoautotrophically makes it a more attractive biocatalyst for many applications including PHB biosynthesis. The extreme metabolic versatility of TIE-1 can also be considered for waste management efforts combined with MES (for example for PHB biosynthesis). This approach can be further scaled up using underwater tubular photobioreactors that have been used previously to investigate the photosynthetic efficiency of R. palustris 42OL [10]. The future of using biocatalysts like TIE-1 via direct EU for bioproduction needs further consideration. We are pursuing modifications of electrodes and changes in reactor design to improve direct EU by TIE-1 as this represents the first major hurdle in the application of such microbes for bioproduction [65]. Using TIE-1 in the context of photoferroautotrophy also needs further investigation as our data support the idea that biomolecule production can be linked to this process.
References
Alkotaini B, Abdellaoui S, Hasan K, Grattieri M, Quah T, Cai R, Yuan M, Minteer SD (2018) Sustainable bioelectrosynthesis of the bioplastic polyhydroxybutyrate: overcoming substrate requirement for NADH regeneration. ACS Sustain Chem Eng 6:4909–4915
Applied Biosystem (2008) Guide to performing relative quantitation of gene expression using real-time quantitative PCR. Available via DIALOG. https://assets.thermofisher.com/TFS-Assets/LSG/manuals/cms_042380.pdf. Accessed 28 Mar 2019
Aryal N, Tremblay PL, Lizak DM, Zhang T (2017) Performance of different Sporomusa species for the microbial electrosynthesis of acetate from carbon dioxide. Bioresour Technol 233:184–190
Bhattacharya S, Dubey S, Singh P, Shrivastava A, Mishra S (2016) Biodegradable polymeric substances produced by a marine bacterium from a surplus stream of the biodiesel industry. Bioengineering 3:34–44
Bolger A, Lohse MM, Usadel B (2014) Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30:2114–2120
Bose A, Gardel EJ, Vidoudez C, Parra E, Girguis PR (2014) Electron uptake by iron-oxidizing phototrophic bacteria. Nat Commun 5:3391–3397
Bose A, Newman DK (2011) Regulation of the phototrophic iron oxidation (pio) genes in Rhodopseudomonas palustris TIE-1 is mediated by the global regulator, FixK. Mol Microbiol 79:63–75
Bowien B, Kusian B (2002) Genetics and control of CO2 assimilation in the chemoautotroph Ralstonia eutropha. Arch Microbiol 178:85–93
Calvin M, Benson AA (1948) The path of carbon in photosynthesis. Science 107:476–480
Carlozzi P, Pushparaj B, Degl’Innocenti A, Capperucci A (2006) Growth characteristics of Rhodopseudomonas palustris cultured outdoors, in an underwater tubular photobioreactor, and investigation on photosynthetic efficiency. Appl Microbiol Biotechnol 73:789–795
Chen GQ (2010) Plastics completely synthesized by bacteria: polyhydroxyalkanoates. Plastics from bacteria. Springer, Berlin, Heidelberg, pp 17–37
Chen X, Cao Y, Li F, Tian Y, Song H (2018) Enzyme-assisted microbial electrosynthesis of poly (3-hydroxybutyrate) via CO2 bioreduction by engineered Ralstonia eutropha. ACS Catal 8:4429–4437
Clauwaert P, Toledo R, Van der Ha D, Crab R, Verstraete W, Hu H, Udert K, Rabaey K (2008) Combining biocatalyzed electrolysis with anaerobic digestion. Water Sci Technol 57:575–579
Darani KK, Vasheghani-Farahani E, Tanaka K (2006) Hydrogen oxidizing bacteria as poly(hydroxybutyrate) producers. Iran J Biotechnol 4:193–196
De Philippis RA, Ena A, Guastiini M, Sili C, Vincenzini M (1992) Factors affecting poly-β-hydroxybutyrate accumulation in cyanobacteria and in purple non-sulfur bacteria. FEMS Microbiol Lett 103:187–194
Doi Y, Tamaki A, Kunioka M, Soga K (1988) Production of copolyesters of 3-hydroxybutyrate and 3-hydroxyvalerate by Alcaligenes eutrophus from butyric and pentanoic acids. Appl Microbiol Biotechnol. 28:330–334
Dornburg V, Lewandowski I, Patel M (2003) Comparing the land requirements, energy savings, and greenhouse gas emissions reduction of biobased polymers and bioenergy: An analysis and system extension of life-cycle assessment studies. J Ind Ecol 7:93–116
Doud DFR, Angenent LT (2014) Toward electrosynthesis with uncoupled extracellular electron uptake and metabolic growth: enhancing current uptake with Rhodopseudomonas palustris. Environ Sci Technol Lett 1:351–355
Drosg B, Fritz I, Gattermayr F, Silvestrini L (2015) Photo-autotrophic production of poly (hydroxyalkanoates) in cyanobacteria. Chem Biochem Eng Q 29:145–156
Eddie BJ, Wang Z, Hervey WJ, Leary DH, Malanoski AP, Tender LM, Lin B, Strycharz-Glaven SM (2017) Metatranscriptomics supports the mechanism for biocathode electroautotrophy by “Candidatus Tenderia electrophaga”. MSystems 2:e00002–e00017
Ehrenreich A, Widdel F (1994) Anaerobic oxidation of Fe(II) by purple bacteria, a new type of phototrophic metabolism. Appl Environ Microbiol 60:4517–4526
Eroğlu E, Gündüz U, Yücel M, Türker L, Eroğlu İ (2004) Photobiological hydrogen production by using olive mill wastewater as a sole substrate source. Int. J Hydrog Energy 29:163–171
Gralnick JA, Newman DK (2007) Extracellular respiration. Mol Microbiol 65:1–11
Greenwood NN, Earnshaw A (2012) Chemistry of the elements. Elsevier, Leeds
Ha PT, Lindemann SR, Shi L, Dohnalkova AC, Fredrickson JK, Madigan MT, Beyenal H (2017) Syntrophic anaerobic photosynthesis via direct interspecies electron transfer. Nat Commun 8:13924–13930
Hai T, Lange D, Rabus R, Steinbüchel A (2004) Polyhydroxyalkanoate (PHA) accumulation in sulfate-reducing bacteria and identification of a class III PHA synthase (PhaEC) in Desulfococcus multivorans. Appl Environ Microbiol 70:4440–4448
Hallbeck L, Ståhl F, Pedersen K (1993) Phytogeny and phenotypic characterization of the stalk-forming and iron-oxidizing bacterium Gallionella ferruginea. J Gen Microbiol 139:1531–1535
Harrington TD, Tran VN, Mohamed A, Renslow R, Biria S, Orfe L, Call DR, Beyenal H (2015) The mechanism of neutral red-mediated microbial electrosynthesis in Escherichia coli: menaquinone reduction. Bioresour Technol 192:689–695
Higuchi-Takeuchi Morisaki MK, Toyooka K, Numata K (2016) Synthesis of high-molecular-weight polyhydroxyalkanoates by marine photosynthetic purple bacteria. PLoS One 11:e0160981–e0160997
Ishizaki A, Tanaka K (1991) Production of poly-β-hydroxybutyric acid from carbon dioxide by Alcaligenes eutrophus ATCC 17697T. J Ferment Bioeng 71:254–257
Jaitz L, Mueller B, Koellensperger G, Huber D, Oburger E, Puschenreiter M, Hann S (2011) LC–MS analysis of low molecular weight organic acids derived from root exudation. Anal Bioanal Chem 400:2587–2596
Jiao Y, Kappler A, Croal LR, Newman DK (2005) Isolation and characterization of a genetically tractable photoautotrophic Fe(II)-oxidizing bacterium, Rhodopseudomonas palustris strain TIE-1. Appl Environ Microbiol 71:4487–4496
Jourdin L, Lu Y, Flexer V, Keller J, Freguia S (2016) Biologically induced hydrogen production drives high rate/high efficiency microbial electrosynthesis of acetate from carbon dioxide. ChemElectroChem 3:581–591
Kamravamanesh D, Pflügl S, Nischkauer W, Limbeck A, Lackner M, Herwig C (2017) Photosynthetic poly-β-hydroxybutyrate accumulation in unicellular cyanobacterium Synechocystis sp. PCC 6714. AMB Express 7:143–154
Karr B, Waters JK, Emerich DW (1983) Analysis of poly-3-hydroxybutyrate in Rhizobium japonicum bacteroids by ion-exclusion high-pressure liquid chromatography and UV detection. Appl Environ Microbiol 46:1339–1344
Kato S (2015) Biotechnological aspects of microbial extracellular electron transfer. Microbes Environ 30:133–139
Kheshgi HS, Prince RC (2005) Sequestration of fermentation CO2 from ethanol production. Energy 30:1865–1871
Khosravi-Darani K, Mokhtari ZB, Amai T, Tanaka K (2013) Microbial production of poly (hydroxybutyrate) from C1 carbon sources. Appl Microbiol Biotechnol 97:1407–1424
Khunjar WO, Sahin A, West AC, Chandran K, Banta S (2012) Biomass production from electricity using ammonia as an electron carrier in a reverse microbial fuel cell. PLoS One 7:e44846–e44853
Klugkist J, Haaker H (1984) Inhibition of nitrogenase activity by ammonium chloride in Azotobacter vinelandii. J Bacteriol 157:148–151
Kucera D, Benesova P, Ladicky P, Pekar M, Sedlacek P, Obruca S (2017) Production of polyhydroxyalkanoates using hydrolyzates of spruce sawdust: comparison of hydrolyzates detoxification by application of overliming, active carbon, and lignite. Bioengineering 4:53–61
Kwan TH, Hu Y, Lin CSK (2018) Techno-economic analysis of a food waste valorization process for lactic acid, lactide and poly (lactic acid) production. J Clean Prod 181:72–87
Li H, Opgenorth PH, Wernick DG, Rogers S, Wu TY, Higashide W, Malati P, Huo YX, Cho KM, Liao JC (2012) Integrated electromicrobial conversion of CO2 to higher alcohols. Science 335:1596–1597
Love MI, Huber W, Anders S (2014) Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol 15:550–570
Luengo J, Garcia B, Sandoval A, Naharroy G, Olivera E (2003) Bioplastics from microorganisms. Curr Opin Microbiol 6:251–260
Lütters-Czekalla S (1990) Lithoautotrophic growth of the iron bacterium Gallionella ferruginea with thiosulfate or sulfide as energy source. Arch Microbiol 154:417–421
Madigan MT, Martinko JM, Parker J (1996) Nutrition and metabolism. Brock biology of microorganisms, 8th edn. Prentice-Hall, New Jersey, pp 109–177
Manavitehrani I, Fathi A, Badr H, Daly S, Negahi Shirazi A, Dehghani F (2016) Biomedical applications of biodegradable polyesters. Polymers 8:51–82
Marang L, Jiang Y, Van Loosdrecht MCM, Kleerebezem R (2013) Butyrate as a preferred substrate for polyhydroxybutyrate production. Bioresour Technol 142:232–239
McKinlay JB, Harwood CS (2010) Carbon dioxide fixation as a central redox cofactor recycling mechanism in bacteria. Proc Natl Acad Sci 107:11669–11675
McKinlay JB, Oda Y, Rühl M, Posto AL, Sauer U, Harwood CS (2014) Non-growing Rhodopseudomonas palustris increases the hydrogen gas yield from acetate by shifting from the glyoxylate shunt to the tricarboxylic acid cycle. J Biol Chem 289:1960–1970
Nevin KP, Hensley SA, Franks AE, Summers ZM, Ou J, Woodard TL, Snoeyenbos-West OL, Lovley DR (2011) Electrosynthesis of organic compounds from carbon dioxide is catalyzed by a diversity of acetogenic microorganisms. Appl Environ Microbiol 77:2882–2886
Nevin KP, Woodard TL, Franks AE, Summers ZM, Lovley DR (2010) Microbial electrosynthesis: feeding microbes electricity to convert carbon dioxide and water to multicarbon extracellular organic compounds. MBio 1:e00103–e00110
Nielsen C, Rahman A, Rehman AU, Walsh MK, Miller CD (2017) Food waste conversion to microbial polyhydroxyalkanoates. Microb Biotech 10:1338–1352
Nishio K, Kimoto Y, Song J, Konno T, Ishihara K, Kato S, Hashimoto K, Nakanishi S (2013) Extracellular electron transfer enhances polyhydroxybutyrate productivity in Ralstonia eutropha. Environ Sci Tech Let 1:40–43
Page WJ (1992) Production of poly-β-hydroxybutyrate by Azotobacter vinelandii UWD in media containing sugars and co containing sugars and complex nitrogen sources. Appl Microbiol Biotechnol 38:117–121
Parida B, Iniyan S, Goic R (2011) A review of solar photovoltaic technologies. Renew Sust Energ Rev 15:1625–1636
Peguin S, Soucaille P (1996) Modulation of metabolism of Clostridium acetobutylicum grown in chemostat culture in a three-electrode potentiostatic system with methyl viologen as electron carrier. Biotechnol Bioeng 51:342–348
Peoples OP, Sinskey AJ (1989) Poly-beta-hydroxybutyrate (PHB) biosynthesis in Alcaligenes eutrophus H16. Identification and characterization of the PHB polymerase gene (phbC). J Biol Chem 264:15298–15303
Peoples OP, Sinskey AJ (1989) Poly-beta-hydroxybutyrate biosynthesis in Alcaligenes eutrophus H16. Characterization of the genes encoding beta-ketothiolase and acetoacetyl-CoA reductase. J Biol Chem 264:15293–15297
Pinzon NM, Aukema KG, Gralnick JA, Wackett LP (2011) Nile red detection of bacterial hydrocarbons and ketones in a high-throughput format. MBio 2:e00109–e00111
Pozo G, Jourdin L, Lu Y, Keller J, Ledezma P, Freguia S (2016) Cathodic biofilm activates electrode surface and achieves efficient autotrophic sulfate reduction. Electrochim Acta 213:66–74
Quelas J, Mesa S, Mongiardini Jendrossek D, Lodeiro A (2016) Regulation of polyhydroxybutyrate synthesis in the soil bacterium Bradyrhizobium diazoefficiens. Appl Environ Microbiol 82:4299–4308
Rabaey K, Rozendal RA (2010) Microbial electrosynthesis—revisiting the electrical route for microbial production. Nat Rev Microbiol 8:706–716
Rengasamy K, Ranaivoarisoa T, Singh R, Bose A (2018) An insoluble iron complex coated cathode enhances direct electron uptake by Rhodopseudomonas palustris TIE-1. Bioelectrochem 122:164–173
Richter K, Schicklberger M, Gescher J (2012) Dissimilatory reduction of extracellular electron acceptors in anaerobic respiration. Appl Environ Microbiol 78:913–921
Sabbagh F, Muhamad II (2017) Production of poly-hydroxyalkanoate as secondary metabolite with main focus on sustainable energy. Renew Sust Energ Rev 72:95–104
Sakakibara Y, Kuroda M (1993) Electric prompting and control of denitrification. Biotechnol Bioeng 42:535–537
Salgaonkar BB, Bragança JM (2017) Utilization of sugarcane bagasse by Halogeometricum borinquense strain E3 for biosynthesis of poly (3-hydroxybutyrate-co-3-hydroxyvalerate). Bioengineering 4:50–67
Schlager S, Haberbauer M, Fuchsbauer A, Hemmelmair C, Dumitru LM, Hinterberger G, Neugebauer H, Sariciftci NS (2017) Bioelectrocatalytic application of microorganisms for carbon dioxide reduction to methane. Chemsuschem 10:226–233
Shi H, Shiraishi M, Shimizu K (1997) Metabolic flux analysis for biosynthesis of poly (β-hydroxybutyric acid) in Alcaligenes eutrophus from various carbon sources. J Ferment Bioeng 84:579–587
Steinbusch KJJ, Hamelers HVM, Schaap JD, Kampman C, Buisman CJN (2010) Bioelectrochemical ethanol production through mediated acetate reduction by mixed cultures. Environ Sci Technol 44:513–517
Straub KL, Rainey FA, Widdel F (1999) Rhodovulum iodosum sp. nov. and Rhodovulum robiginosum sp. nov., two new marine phototrophic ferrous-iron-oxidizing purple bacteria. Int J Syst Bacteriol 49:729–735
Taepucharoen K, Tarawat S, Puangcharoen M, Incharoensakdi A, Monshupanee T (2017) Production of poly (3-hydroxybutyrate-co-3-hydroxyvalerate) under photoautotrophy and heterotrophy by non-heterocystous N2-fixing cyanobacterium. Bioresour Technol 239:523–527
Takahashi RYU, Castilho NAS, Silva MAC, Miotto MC, Lima AOS (2017) Prospecting for marine bacteria for polyhydroxyalkanoate production on low-cost substrates. Bioengineering 4:60–72
Tanaka K, Miyawaki K, Yamaguchi A, Khosravi-Darani K, Matsusaki H (2011) Cell growth and P (3HB) accumulation from CO2 of a carbon monoxide-tolerant hydrogen-oxidizing bacterium, Ideonella sp. O-1. Appl Microbiol Biotechnol 92:1161–1169
Todar K (2002) Growth of bacterial populations. In: Todar’s online textbook of bacteriology. Available via DIALOG. http://textbookofbacteriology.net/growth.html. Accessed 28 Mar 2019
Tremblay P-L, Zhang T (2015) Electrifying microbes for the production of chemicals. Front Microbiol 6:201–210
Verlinden RA, Hill DJ, Kenward M, Williams CD, Radecka I (2007) Bacterial synthesis of biodegradable polyhydroxyalkanoates. J Appl Microbiol 102:1437–1449
Villano M, Aulenta F, Ciucci C, Ferri T, Giuliano A, Majone M (2010) Bioelectrochemical reduction of CO2 to CH4 via direct and indirect extracellular electron transfer by a hydrogenotrophic methanogenic culture. Bioresource Technol 101:3085–3090
Wang Q, Yu H, Xia Y, Kang Z, Qi Q (2009) Complete PHB mobilization in Escherichia coli enhances the stress tolerance: a potential biotechnological application. Microb Cell Fact 8:47–55
Wickham H (2016) ggplot2: elegant graphics for data analysis. Springer, Texas
Widdel F, Schnell S, Heising S, Ehrenreich A, Assmus B, Schink B (1993) Ferrous iron oxidation by anoxygenic phototrophic bacteria. Nature 362:834–836
Wu SC, Liou SZ, Lee CM (2012) Correlation between bio-hydrogen production and polyhydroxybutyrate (PHB) synthesis by Rhodopseudomonas palustris WP3-5. Bioresour Technol 113:44–50
Xiao N, Jiao N (2011) Formation of polyhydroxyalkanoate in aerobic anoxygenic phototrophic bacteria and its relationship to carbon source and light availability. Appl Environ Microbiol 77:7445–7450
York G, Stubbe J, Sinskey A (2001) New insight into the role of the PhaP phasin of Ralstonia eutropha in promoting synthesis of polyhydroxybutyrate. J Bact 183:2394–2397
Yu L, Yuan Y, Tang J, Zhou S (2017) Thermophilic Moorella thermoautotrophica-immobilized cathode enhanced microbial electrosynthesis of acetate and formate from CO2. Bioelectrochemistry 117:23–28
Zhang T, Nie HR, Bain TS, Lu HY, Cui MM, Snoeyenbos-West OL, Franks AE, Nevin KP, Russell TP, Lovley DR (2013) Improved cathode materials for microbial electrosynthesis. Energ Environ Sci 6:217–224
Acknowledgements
We are grateful to Dr. Joshua Blodgett and Yifei Hu for their support during PHB analysis using LC–MS. We appreciate funding provided by The David and Lucile Foundation (201563111) as well as the Department of Energy (Grant number DESC0014613), and the U.S. Department of Defense, Army Research Office (Grant number W911NF-18-1-0037). M.S.G. was supported by NIH Initiative for Maximizing Student Development (IMSD) training Grant R25-GM103757.
Author information
Authors and Affiliations
Contributions
TOR performed all the experiments. RK assisted with the bioelectrochemical experiments. MSG assisted with RNASeq data analysis. RS assisted with STEM-EELS experiments. TOR, RS, and AB wrote the manuscript with input from all authors.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Figure S1.
Heat map showing log2 fold change in the expression of PHB genes from reverse transcription quantitative PCR analysis (RT qPCR). Results are from TIE-1 grown in Fresh Water (FW) medium photoheterotrophically with butyrate, photoautotrophically with H2, photoferroautotrophically with Fe(II), and photoelectroautotrophically with a poised electrode. Growth with NH4Cl is indicated by NH4Cl whereas growth under N2 fixing conditions is indicated by N2. Error bars are the standard deviations calculated using 3 biological replicates. The P values were determined by one-way ANOVA followed by a pairwise test with Bonferroni adjustment; ns: not significant. For a pairwise test, the cutoff P value is 0.025). P values are indicated in Table S10. (TIFF 88863 kb)
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Ranaivoarisoa, T.O., Singh, R., Rengasamy, K. et al. Towards sustainable bioplastic production using the photoautotrophic bacterium Rhodopseudomonas palustris TIE-1. J Ind Microbiol Biotechnol 46, 1401–1417 (2019). https://doi.org/10.1007/s10295-019-02165-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10295-019-02165-7