Abstract
Verticillins are a group of epipolythiodioxopiperazine alkaloids that have displayed potent cytotoxicity. To evaluate their potential further, a larger supply of these compounds was needed for both in vivo studies and analogue development via semisynthesis. To optimize the biosynthesis of these secondary metabolites, their production was analyzed in two different fungal strains (MSX59553 and MSX79542) under a suite of fermentation conditions. These studies were facilitated by the use of the droplet-liquid microjunction-surface sampling probe (droplet probe), which enables chemical analysis in situ directly from the surface of the cultures. These experiments showed that the production of verticillins was greatly affected by growth conditions; a significantly higher quantity of these alkaloids was noted when the fungal strains were grown on an oatmeal-based medium. Using these technologies to select the best among the tested growth conditions, the production of the verticillin analogues was increased while concomitantly decreasing the time required for fermentations from 5 weeks to about 11 days. Importantly, where we could previously supply 5–10 mg every 6 weeks, we are now able to supply 50–150 mg quantities of key analogues per month via laboratory scale fermentation.
Graphical abstract

Similar content being viewed by others
Introduction
Fungi are a rich resource for biologically active and structurally diverse secondary metabolites [16, 26, 27]. As such, a number of authors have noted the value of investing in fungi as a source of new drug leads, especially to treat cancer and overcome its resistance [6, 44]. Cancer is a worldwide problem, and in the USA alone, over one million new cases were estimated for 2016, resulting in more than half a million cancer-related deaths [70].
As part of ongoing studies to discover new anticancer leads from filamentous fungi [20, 44, 55], our team has isolated hundreds of fungal metabolites over the last decade [16]. Of these, verticillins are one of the most interesting for a variety of reasons [20, 48, 49, 58, 79]. This class of bioactive fungal secondary metabolites (as exemplified by compounds 1–7) is characterized by a dimeric structure [24] and is a key member of the epipolythiodioxopiperazine (ETP) alkaloids. Biologically, these compounds display potent antitumor activity [10, 11, 35, 72, 80], along with antibacterial [36, 53, 81, 82], nematicidal [14] and immune induction properties [18].
The lead compound of this class, verticillin A (1), was first discovered in 1970 [37], and since then, 27 verticillin analogues have been characterized [20]; even 30 years ago, their cytotoxic potential was noted [35]. At the time of discovery, verticillin A (1) showed an ED50 of 0.2 µg/mL (0.27 µM) against HeLa cells [37]. In another study, verticillin A (1), along with the closely related analogues, Sch 52900 (5) and Sch 52901 (2), showed potential in preventing the transcription of c-fos promoter gene, which is a proto-oncogene found overexpressed in a variety of cancers [12]. Two analogues, 11, 11ʹ-dideoxyverticillin A (6) and 11ʹ-deoxyverticillin A (7), demonstrated in vitro cytotoxicity against HCT-116 (human colon carcinoma) cells in the nanomolar range [72]. These later compounds also demonstrated an inhibitory effect of tyrosine kinase on the growth factor receptor, thereby leading to anti-tumor activity [80]. Verticillin A showed potent activity in sensitizing human colorectal cancer cells to apoptosis both in vitro and in vivo, as well as, the ability to induce cell-cycle arrest at the G2 phase [48].
Biologically, interest has been growing in the verticillins, particularly after verticillin A (1) was shown to be a selective histone methyl transferases (HMT) inhibitor [58]. The HMT inhibitory activity resulted in the demethylation of H3K9me3 of the silenced FAS gene [58], thereby allowing its transcription, such that FAS was re-expressed, triggering apoptosis mediated cell death [45]. Moreover, verticillin A (1) induced apoptosis selectively against malignant tumor cells, while minimal effect was exhibited against healthy cells [79]. Verticillin A (1) showed potential to overcome metastatic colon carcinoma resistance to 5-fluorouracil. Mice transplanted with SW620-5FU-R cells were treated with a combination of verticillin A (1) and 5-fluorouracil and displayed significantly smaller tumors compared to the groups receiving only one of these two treatments [58]. In a recent study conducted on pancreatic cancer by Lu et al. [49], verticillin A was given to two groups of mice transplanted with either PANC02-H7 or UN-KC-6141 cells. Verticillin A was administrated alone or in combination with anti PD-L1 (programmed death ligand) treatment to provide immunotherapy. The results demonstrated that verticillin A (1), in conjugation with anti PD-L1 ligand, markedly reduced tumor size and viability. In summary, a suite of pharmacological studies over the last 5 years has increased the interest surrounding this class of compounds.
Despite the promising activities that verticillins have shown in vitro and in vivo, research has been hindered by the lack of material. In previous work, 0.66–1.72 mg of verticillin analogues (1–3) per g of extract could be isolated from large scale fermentations of fungal cultures containing 150 g of rice (with a total isolated of 1.6–5.5 mg) [20]. Similarly, 0.37–0.85 mg of verticillin analogues were isolated per g of extract (with a total isolated of 3.5–8.0 mg) [36]. Based on the literature, the yields of these compounds were not improved greatly when grown in liquid cultures. For example, from a 73 L fermentation, the yield of verticillin G was 0.58 mg per g of extract (with a total isolated of 2.9 mg) [82]. In another study that used 20 L fermentations, compounds 1, 6 and 7 were isolated with a yield of 0.25 mg, 0.61 mg and 1.14 mg per g of extract, respectively (with a total yield of 3.5 mg, 8.5 mg and 16 mg, respectively) [72]. Furthermore, although elegant synthetic approaches have been reported for the preparation of various epipolythiodioxopiperazine alkaloids [8, 42, 43], these processes require numerous reactions steps, making the large-scale production of these compounds challenging. About 0.2 g of (+)-11,11ʹ-dideoxyverticillin A was prepared in 14 steps from ˃ 10 g scale of commercially available amino acids derivatives [42]. In summary, the production of the verticillins in the literature, whether using liquid-based or solid-based fermentations, has been largely on the scale of only a couple mg, and most of the verticillins have never been synthesized. For these reasons, the adequate supply of key analogues must be addressed before the full pharmacological potential of these compounds can be realized.
Many studies have shown the influence of media and environmental factors on fungal biosynthesis [4, 9], where optimal nutrition is required for ideal fungal growth [46, 51]. The OSMAC approach is a prominent strategy to probe the importance of fungal fermentation conditions, potentially generating broader chemical diversity [29, 30] and/or targeting the enhanced production of a distinct class of secondary metabolites. In this study, several fungal strains were selected from the Mycosynthetix library. These cultures were grown on a suite of different solid media, and productivity was assessed by taking advantage of new instrumentation that facilitates the screening of the chemistry of the cultures. The droplet-liquid microjunction-surface sampling probe (droplet probe) permits the in situ analysis of targeted secondary metabolites directly from the surface of cultures [40, 67]. Our goal was to enhance the biosynthesis of verticillins by identifying a strain that produces the highest quantity of the analogues under appropriate fermentation conditions in the shortest period of time.
Materials and methods
Mycology
Fungal cultures were provided by Mycosynthetix, Inc., via an ongoing project to identify diverse natural product study material as a source of anticancer drug leads [44]. Over the last decade, we have analyzed the chemistry and biological activity of thousands of cultures from the Mycosynthetix library, and among these, ten cultures were identified as being able to biosynthesize verticillin A and its analogues (compounds 1–5) based on published dereplication protocols [15, 56]. Accordingly, the following strains were chosen for this study: MSX74391, MSX71844, MSX58124, MSX75296, MSX75281, MSX45374, MSX70777, MSX59553 and MSX79542 (Table 2S). In a previous study, fungal strains MSX59553 and MSX79542 were both identified as Clonostachys rogersoniana [56].
Identification of fungal strains
The different MSX strains utilized in the present study were identified to the genus and/or species level using the fungal ITS region (ITS1–5.8S-ITS2) of the nuclear ribosomal operon, which was amplified with primer combination ITS5 and ITS4 with primers ITS5/ITS1F and ITS4 [23, 77]. Methods for BLAST search in NCBI GenBank and Maximum Likelihood phylogenetic analysis, as utilized to identify fungal strains, have been detailed recently [61]. The BLAST search and phylogenetic analysis using taxon sampling [1] revealed that MSX59553, MSX79542, MSX74391, MSX58124, MSX75296, MSX75281, MSX45374, and MSX70777 belong to the genus Clonostachys (Ascomycota, Hypocreales, Bionectriaceae), while MSX71844 was identified as Purpureocillium lavendulum (Ascomycota, Hypocreales, Ophiocordycipitaceae) (Fig. S1). Phylogenetic analysis using the ITS region also showed that strains MSX59553, MSX45374, MSX74391, MSX79542, MSX70777, and MSX58124 had phylogenetic affinities with C. rogersoniana [64]. Our results were in agreement with a recent genomic study identifying the biosynthetic gene clusters for verticillins in C. rogersoniana [76]. As the ITS region alone is not informative to identify species in certain orders of the Ascomycota, including Hypocreales [33], future taxonomic and molecular phylogenetic studies will incorporate sequence data from protein-coding regions to more precisely identify species names for verticillin-producing strains [54]. The sequence data were deposited in GenBank (accession numbers: KX845687, KX845688, MH421853, MH421854, MH421855, MH421856, MH421857, MH421858, MH421859, and MH421860).
For verticillin producing strains, there has been some confusion on the fungal taxonomy [13]. Earlier studies on the chemistry of verticillin analogues have identified the verticillin-producing fungi as belonging to the genera Gliocladium, Penicillium, and Verticillium [36, 37, 72]. The morphology of the flask shaped phialide in all three genera can be a cause of confusion for non-mycologists. Moreover, fungal taxonomic names have been rapidly changing due to the use of molecular sequence data in the past 20 years [61]. In addition, the name Bionectria (sexual state) has been used previously in the literature for a fungal strain producing verticillin G [82]; while this was accurate, the use of name Bionectria is now phased out due to the adoption of One Fungus = One Name [28, 73], in accordance with the recent changes concerning pleomorphic fungi in the International Code of Nomenclature for algae, fungi, and plants. For pleomorphic names (sexual and asexual) in the family Bionectriaceae, Rossman et al. [63] have proposed the use and protection of the asexual morph (Clonostachys) rather than the sexual morph (Bionectria). Despite the adoption of the name Clonostachys, the subgenus Bionectria is still being used in the literature. Based on our recent identification of fungal strains that make verticillin and its analogues, we have identified two fungal genera, Clonostachys spp. and Purpureocillium lavendulum [59], that biosynthesize the epipolythiodioxopiperazine-type alkaloids.
Media and fermentations
For the analysis of fungal cultures in Petri dishes, seven types of agar media were used, and ingredients to make these were purchased from Difco, ACROS, and Research Products International (RPI): malt extract agar, potato dextrose agar, yeast extract soy peptone dextrose agar, Spezieller Nährstoffarmer agar, Sabouraud dextrose agar, potato dextrose mushroom and oatmeal agar (See Table 1S for compositions). A small piece (~ 0.5 cm2) of fresh culture grown on potato dextrose agar was cut from the leading edge of 2-week-old colony from strains MSX59553 and MSX79542 and was used to inoculate various types of agar media in duplicate, respectively. The plates were then incubated at room temperature until the cultures showed good growth as noted by the expansion of the colony over the surface of the medium (~ 3 to 4 weeks).
Rice fermentations [(1:1 commercial white rice (variety: Calrose Botan; 5 g) and (variety: Sona Masoori; 5 g)] were prepared by adding 10 g of rice to a Petri dish or a 250 mL flask with 25 mL of DI-H2O, followed by autoclaving at 221 °C for 30 min. Oatmeal media (Old fashioned Quaker oats) was prepared using 10 g of rolled oats and ~ 17 mL of DI-H2O, followed by autoclaving at 221 °C for 30 min. The flasks containing the rice or oatmeal media were inoculated with a seed culture grown in 10 mL of YESD broth media (2% soy peptone, 2% dextrose and 1% yeast extract) for about 5 days with agitation (100 rpm) at room temperature. The inoculated cultures were grown at room temperature for ~ 4–5 weeks, except for the time progression studies, where each duplicate of the oatmeal culture was extracted every few days, from days 2 to 35. Strains were scaled-up in a similar way, using 2.8 L Fernbach flasks (Corning, Inc., Corning, NY, USA). The medium was prepared using 150 g of either rice or oatmeal, with 300 mL or 250 mL of DI-H2O added for rice medium and oatmeal medium, respectively, and both sterilized at 221 °C for 30 min. In addition, and as noted in more detail in the supplement (Fig. S7), augmenting the oatmeal media with amino acids was also examined. However, this did not result in significant differences in the biosynthesis of verticillins, and thus, this line of inquiry was not pursued further.
Extraction of fungal cultures
Each culture was chopped with a spatula to ensure thorough extraction. For the cultures grown on Petri dishes, the chopped pieces were transferred into 150 mL flasks. Solutions of 60 mL of 1:1 chloroform:methanol (CHCl3:MeOH) were poured in each flask, followed by overnight shaking (100 rpm) at room temperature. The cultures were filtered by vacuum, and then the filtrates were mixed with 90 mL of CHCl3 and 150 mL of H2O and stirred for 30 min. The mixtures were transferred into a separatory funnel, where the bottom organic layer was evaporated in vacuo and reconstituted with 100 mL of 1:1 acetonitrile:methanol (CH3CN:MeOH) and 100 mL hexanes. Again, the defatted organic layers were collected and evaporated in vacuo.
In situ analysis of fungal metabolite profiles via the droplet probe coupled with UPLC-PDA-HRMSMS/MS
The use of the droplet probe with fungal cultures has been detailed previously [67] and several examples have been published [57, 65, 66, 68, 69]. A CTC/LEAP HTC Pal auto-sampler was converted to a droplet probe with assistance from colleagues at Oak Ridge National Laboratories [38, 39]. Microextractions (3–5 µL) were performed on precise spots directly on fungal cultures using a 1:1 solution of MeOH:H2O. The extractions were injected onto a UPLC, and the chromatographic method followed previous protocols [15] used in the dereplication of fungal extracts. A Waters Acquity UPLC system was used with a BEH C18 column (Waters; 50 mm × 2.1 mm × 1.8 µm) heated to 40 °C and a mobile phase using a gradient from 15 to 100% CH3CN; the other solvent was 0.1% formic acid-H2O. The run utilized a flow rate of 0.3 mL/min for 10 min. UV data were collected from 190 to 500 nm and the eluent was split into a Thermo Fisher Scientific Q Exactive Plus mass spectrometer via electrospray ionization (ESI). MS data were collected from 150 to 2000 m/z at a resolution of 70,000 while alternating between positive and negative modes. MS/MS was performed with an HCD value of 35, and Xcalibur software was used for the primary analysis.
Analysis of fungal metabolite profiles via UPLC-PDA-HRMSMS/MS
Extracts of the plates and flasks were dissolved separately in 1:1 MeOH:dioxane to elaborate concentrations of 2.5 × 10−2, 0.5 or 2.0 mg/mL; 3 μL were injected directly onto the UPLC-PDA-HRMSMS/MS system. In addition to HRMS and MSMS data, retention times and UV data were used as mutually supportive data. The relative peak areas of each targeted secondary metabolite were measured (Fig. 1).
Results and discussion
Different fungal strains biosynthesize distinct patterns of verticillin analogues
The dereplication protocol was used to identify fungal strains that produced verticillins [15] (Table 2S Supporting Information). Extracts of suspected verticillin producing strains from our library were screened using UPLC-HRMS, revealing a higher production of verticillin analogues within the organic extracts of both strains MSX59553 and MSX79542 (Fig. 2). The graph shows the relative percentage of the targeted verticillin analogues as measured in triplicate. To more accurately compare the total production of the secondary metabolites, the calculations were normalized to the largest value and multiplied by the total mass of the defatted organic extract of each strain. Interestingly, some strains produced higher amounts of verticillin D (8) but only low levels of verticillin A (1) and related analogues (and vice versa), suggesting that the biosynthetic pathway for these analogues were somewhat different, even though the fungi were presumably all using non-ribosomal peptide synthesis to produce these metabolites [21].
Relative production of various verticillin analogues grown on rice for 4 weeks across ten different fungal strains as measured by LC-HRMS in three replicates. Strains MSX59553 and MSX79542 demonstrated the highest relative biosynthesis of the targeted verticillin analogues (compounds 1–5). Interestingly, specific strains (such as MSX75296 and MSX75281) showed the biosynthesis of higher amounts of verticillin D (8), corresponding to low to minimal production of the other analogues
Use of the droplet probe for in situ chemical analysis to optimize the production of verticillins
Optimization of fermentation parameters is a well-established approach to enhance the yield of compounds biosynthesized by microorganisms [41, 78]. The alteration of media parameters had an important role in varying the biosynthesis of secondary metabolites in numerous fungal strains [7, 19, 22, 52]. In most cases, investigations of optimal fermentation conditions are targeted for specific active compounds, such as antibiotics [2, 60, 71].
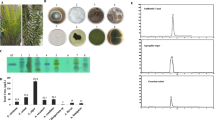
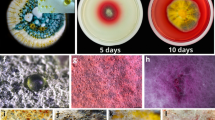
Toward this end, the variation of culture media showed the most immediate results. A visual difference in the growth of the two fungal strains (MSX59553 and MSX79542) was noted (Fig. 3). Guttates, which we and others hypothesize concentrate secondary metabolites [25, 34, 66, 75], were noted when these were grown on Oat-A, YESDA, and PD-mushroom agar (Fig. S6), but not found on the fungi grown on the other nutrient media.
Top: Photos of strains MSX59553 (top) and MSX79542 (bottom). These fungal cultures were grown on SDA, PD-mushroom, Oat-A, SNA and PDA(A)/MEA(B)/YESDA(C) (from left to right, respectively) for 4 weeks. Fungal growths were morphologically distinct based on media, and guttates were observed only on cultures grown on PD-mushroom and Oat-A. Bottom: Relative biosynthesis of verticillin analogues via droplet probe directly from the surface of the strains MSX59553 (bottom left) and MSX79542 (bottom right)
During this study, solid media were chosen over submerged liquid media, despite the advantages of the latter in offering control over the fermentation parameters and the simplification of scale-up for industrial purposes [50]. Pragmatically, we do not have the facilities for large-scale liquid fermentation, and moreover, liquid media for the production of verticillins did not show promise in the literature [12, 72, 82]. Furthermore, the chemistry of the cultures grown on solid media could be studied readily in situ via the droplet probe. In addition, solid fermentation is closer to the natural habitat of fungi, offering the mycelium a support for growth [62] and permitting better oxygen circulation than can be achieved by the high viscosity of liquid fermentation [17]. Previous comparative studies showed a notably higher productivity of pure compounds when microorganisms were grown on solid media, in comparison to liquid cultures [2, 3, 5, 32, 47, 74] with shorter fermentation time and less risk of enzymatic degradation [31].
The in situ microextractions by droplet probe were analyzed by UPLC-HRMS to identify the targeted verticillin analogues in each culture. Previously, a dereplication strategy [67] demonstrated the effectiveness of direct identification of secondary metabolites from cultures grown in Petri dishes. The peak areas of targeted compounds were averaged to build graphs (Fig. 3). For each culture condition, three sampling points were acquired on duplicate plates (n = 6). However, error bars were not calculated for two reasons. First, the three different spots represent different fungal growth stages, essentially oldest to youngest when sampling radially from the center. Also, the fungal growth morphology on each media had different traits, such as guttates (liquid droplets) or mycelium (filamentous hyphal growth). Due to this inherent biological variability, the recovery of the 5 µL droplet during microextraction was not always precisely the same (Fig. S2).
The targeted secondary metabolites (Fig. 1) were eluted within a retention time windows of 5.30–6.30 min on the 10 min gradient method. Parameters such as retention time, UV profile, mass range within 5 ppm, and fragmentation patterns were used to identify the structurally related analogues [56]. The production of these compounds was compared by measuring the percentage of the relative production of each compound normalized to the largest value. The biosynthetic profile varied according to the respective growth media. Both strains showed essentially zero to slight production of verticillin analogues in SDA, SNA, MEA and PDA media. Interestingly, a recent study on the gene clusters of verticillin biosynthesis noted PDA as a condition where the genes for biosynthesis were not expressed [76]; our phenotypic data (i.e., secondary metabolites) were supportive of their genotypic data. Of all the conditions examined, Oat-A was the most productive for verticillin biosynthesis in MSX79542 (Fig. 3). Similar results were observed with Oat-A in strain MSX59553, although two other media also showed promise (YESD and PD-mushroom).
Production of the targeted secondary metabolites verified by quantification via UPLC-HRMS
We next evaluated the production of verticillins on Oat-A vs. readily available substrates, such as breakfast oatmeal and rice (Fig. S3). The UPLC-HRMS analysis of the in situ microextraction on the surface of the three media via the droplet probe were plotted as the relative percentages of the peak normalized according to the compound with highest production (Fig. 4a, b). These results revealed a higher production of the targeted secondary metabolites on oatmeal and Oat-A media compared to the rice. For strain MSX79542, cultures grown on Oat-A showed the highest production, as the fungus expressed guttates (Fig. S6), which have been shown to be concentrated in secondary metabolites [25, 34].
Relative production of verticillin analogues via droplet probe directly from the surface of strains MSX59553 (a) and MSX79542 (b) grown for 4 weeks. Oatmeal agar showed a higher amount of analogues, likely due to the formation of guttates on the surface of the culture. The same cultures grown on the Petri dishes, and used previously for the droplet probe analysis, were fully extracted and then analyzed via UPLC-HRMS to generate graphs (c) and (d), which represent the relative total production of verticillin analogues by strains MSX59553 and MSX79542, respectively. The relative percentages were normalized by multiplying the peak areas by the weight of their corresponding organic extracts (Fig S2 Supplementary information). The results from c and d demonstrate that oatmeal was the best media for total production of verticillin analogues
The cultures on Oat-A did not grow as three-dimensional as those grown on rice or breakfast oatmeal. We hypothesized that the three-dimensional character may be beneficial for total verticillin production and thus, the same in situ microextracted Petri dishes were extracted in their entirety, and oatmeal and rice had the highest extract amounts (Fig. S4). The peak areas of the compounds of interest were calculated in their respective total defatted mass of the extract. Then, those numbers were normalized according to the compound of highest production to give a more accurate comparison of the different profiles of verticillin biosynthesis. Regular oatmeal demonstrated the highest amounts of the targeted secondary metabolites (Fig. 4c, d).
To make a more accurate comparison and quantify the amounts of verticillins from the organic extracts of the two strains MSX59553 and MSX79542, cultures were fermented in three biological replicates using rice and oatmeal for comparative purposes. Oat-A was excluded from this comparison because the agar media forces the fungus to grow mainly on the surface of the culture, and that contributed to a low extract weight compared to the other cultures grown for the same amount of time (Fig. S4).
Verticillin amounts were calculated via UPLC-HRMS calibration curves prepared using purified analogues. To determine the amounts of verticillin analogues produced in mg per flask of growth, the previous numbers were multiplied by the defatted extract mass of each flask. The defatted weights of organic extracts were higher in the strain grown on oatmeal than when it was grown on rice (Fig. S5). Both fungal strains demonstrated the ability to produce higher amounts of verticillin analogues when grown on oatmeal compared to the rice media. Also, using one way ANOVA, strain MSX59553 expressed significantly higher amounts of secondary metabolites when grown on oatmeal than the same strain grown on rice or strain MSX79542 grown on either rice or oatmeal (P < 0.005) (Fig. 5b, c).
The amounts of the secondary metabolites of interest in mg present in each culture of strains MSX59553 and MSX79542 grown on rice or oatmeal media for 4 weeks in three biological replicates (a). The quantity of Sch52901 (2), the verticillin analogue produced in the highest amount by the fungal strains, was calculated per flask. The data plotted are mean ± SD of three biological replicates per medium and per strain. The quantity of 2 produced by strain MSX59553 grown on oatmeal showed a significant difference relative to the same strain grown on rice and also compared to strain MSX79542 grown on oatmeal and rice, respectively (**P < 0.004, ***P < 0.0003) (b). Similarly, the sum of all verticillin analogues showed a significant difference between the oatmeal culture of strain MSX59553 and the other growth conditions (**P < 0.005, ***P < 0.0003) (c)
The extraction and isolation of verticillin analogues from strain MSX59553 grown on a large scale on rice versus that on oatmeal for the same amount of time (5 weeks) was compared. When fermenting on oatmeal, there was a yield of 67.3 mg of verticillins 1–5 per g of extract (with a total weight of pure verticillin analogues isolated being 174.8 mg). Alternatively, when rice was used as a medium, there was 63.3 mg of 1–5 per g of extract (with a total weight of 107.6 mg of purified verticillins). Growth of this fungal culture on oatmeal or rice showed enhanced production of the verticillins, substantially exceeding what had been reported in the literature to date.
Profile of secondary metabolites based on different growth times
Duplicate cultures of MSX59553 grown on oatmeal were sampled over 5 weeks to investigate the effect of incubation time on the production of secondary metabolites. The amount of verticillin A was calculated from a calibration curve after injecting 0.025 mg/mL of the extract of each flask. This preliminary amount was multiplied by the total mass of the defatted organic extract to normalize the results (Fig. 6). The production of verticillins increased up to day 7, after which the production of verticillins plateaued by day 22. Interestingly, a decrease of verticillin A was noticed after day 22 until day 35, which was notable since fungal cultures are routinely grown for approximately 4 weeks in laboratory settings. Thus, our study showed that an average of 11 days was enough time for the biosynthesis, which shortened turnaround time for the scaled-up production of the verticillin analogues.
Amount of verticillin A (mg) produced by MSX59553 per flask (with 10 g of oatmeal) during a period of 35 days. The cultures were extracted and then analyzed using UPLC-HRMS. The fungus showed the most production from the 7th to 22nd days of culture before the start of a decline. These data are mean ± SD of two biological replicates, each analyzed in triplicate (n = 6)
Conclusions
The verticillins are a class of fungal metabolites whose biological activity against a suite of anticancer assays has stimulated recent interest. However, our challenge was to supply them on a large enough scale, so as to facilitate further preclinical studies. This study showed that fermentation on Quaker oatmeal significantly enhanced the biosynthesis of verticillin analogues in strain MSX59553; whereas we once could isolate about 10 mg after about 5 weeks of growth on rice medium, we can now produce about 20 mg in 11 days on oatmeal medium. To examine a suite of fermentation conditions efficiently, we demonstrated the use of the droplet probe to measure the chemistry of fungal cultures grown in Petri dishes in situ. Parallel studies of extracts from those Petri dishes using a more traditional natural products extraction approach followed by quantitative UPLC-HRMS analysis showed the same trend, serving to further validate the reliability of chemical results from droplet probe analyses. For laboratory-scale production, we are confident that a version of the described procedures could be used to supply verticillins on the single to multi-gram scale. However, further research, potentially exploring a suite of options, ranging from liquid fermentations to semi- and/or total synthesis, may be required should the need for kg quantities arise.
References
Abreu LM, Moreira GM, Ferreira D, Rodrigues-Filho E, Pfenning LH (2014) Diversity of Clonostachys species assessed by molecular phylogenetics and MALDI-TOF mass spectrometry. Fungal Biol 118:1004–1012
Barrios-González J, Castillo TE, Mejía A (1993) Development of high penicillin producing strains for solid state fermentation. Biotechnol Adv 11:525–537
Barrios-González J, Rodríguez GM, Tomasini A (1990) Environmental and nutritional factors controlling aflatoxin production in cassava solid state fermentation. J Ferment Bioeng 70:329–333
Bennett JW, Ciegler A (1983) Secondary metabolism and differentiation in fungi. M. Dekker, New York
Bills GF, Dombrowski AW, Goetz MA (2012) The “FERMEX” method for metabolite-enriched fungal extracts. In: Keller NP, Turner G (eds) Fungal secondary metabolism: methods and protocols. Humana Press, New Jersey, pp 79–96
Bills GF, Gloer JB (2016) Biologically active secondary metabolites from the fungi. Microbiol Spectr 4(6). https://doi.org/10.1128/microbiolspec.FUNK-0009-2016
Bode HB, Bethe B, Höfs R, Zeeck A (2002) Big effects from small changes: possible ways to explore nature’s chemical diversity. ChemBioChem 3:619–627
Boyer N, Morrison KC, Kim J, Hergenrother PJ, Movassaghi M (2013) Synthesis and anticancer activity of epipolythiodiketopiperazine alkaloids. Chem Sci 4:1646–1657
Bunch AW, Harris RE (1986) The manipulation of micro-organisms for the production of secondary metabolites. Biotechnol Genet Eng Rev 4:117–144
Chen Y, Guo H, Du Z, Liu XZ, Che Y, Ye X (2009) Ecology-based screen identifies new metabolites from a Cordyceps-colonizing fungus as cancer cell proliferation inhibitors and apoptosis inducers. Cell Prolif 42:838–847
Chen Y, Zhang Y, Li M, Zhao W, Shi Y, Miao Z, Zhang X, Lin L, Ding J (2005) Antiangiogenic activity of 11,11′-dideoxyverticillin, a natural product isolated from the fungus Shiraia bambusicola. Biochem Biophys Res Commun 329:1334–1342
Chu M, Truumees I, Rothofsky ML, Patel MG, Gentile F, Das PR, Puar MS (1995) Inhibition of c-fos proto-oncogene induction by Sch 52900 and Sch 52901, novel diketopiperazines produced by Gliocladium sp. J Antibiot 48:1440–1445
Dirk S, Böttcher C, Lee J, Scheel D (2011) Verticillin A is likely not produced by Verticillium sp. J Antibiot 64:523–524
Dong J, He H, Shen Y, Zhang K (2005) Nematicidal epipolysulfanyldioxopiperazines from Gliocladium roseum. J Nat Prod 68:1510–1513
El-Elimat T, Figueroa M, Ehrmann BM, Cech NB, Pearce CJ, Oberlies NH (2013) High-resolution MS, MS/MS, and UV database of fungal secondary metabolites as a dereplication protocol for bioactive natural products. J Nat Prod 76:1709–1716
El-Elimat T, Zhang X, Jarjoura D, Moy FJ, Orjala J, Kinghorn AD, Pearce CJ, Oberlies NH (2012) Chemical diversity of metabolites from fungi, cyanobacteria, and plants relative to FDA-approved anticancer agents. ACS Med Chem Lett 3:645–649
Elibol M, Mavituna F (1997) Characteristics of antibiotic production in a multiphase system. Process Biochem 32:417–422
Erkel G, Gehrt A, Anke T, Sterner O (2002) Induction of differentiation in acute promyelocytic leukemia cells (HL-60) by the verticillin derivative Sch 52900. Z Naturforsch C 57:759–767
Espeso EA, Tilburn J, Arst H Jr, Penalva M (1993) pH regulation is a major determinant in expression of a fungal penicillin biosynthetic gene. EMBO J 12:3947–3956
Figueroa M, Graf TN, Ayers S, Adcock AF, Kroll DJ, Yang J, Swanson SM, Munoz-Acuna U, Carcache de Blanco EJ, Agrawal R, Wani MC, Darveaux BA, Pearce CJ, Oberlies NH (2012) Cytotoxic epipolythiodioxopiperazine alkaloids from filamentous fungi of the Bionectriaceae. J Antibiot 65:559–564
Fox EM, Howlett BJ (2008) Biosynthetic gene clusters for epipolythiodioxopiperazines in filamentous fungi. Mycol Res 112:162–169
Fuchser J, Zeeck A (1997) Secondary metabolites by chemical screening, 34.–aspinolides and aspinonene/aspyrone co-metabolites, new pentaketides produced by Aspergillus ochraceus. Liebigs Annalen 1997:87–95
Gardes M, White TJ, Fortin JA, Bruns TD, Taylor JW (1991) Identification of indigenous and introduced symbiotic fungi in ectomycorrhizae by amplification of nuclear and mitochondrial ribosomal DNA. Can J Bot 69:180–190
Gardiner DM, Waring P, Howlett BJ (2005) The epipolythiodioxopiperazine (ETP) class of fungal toxins: distribution, mode of action, functions and biosynthesis. Microbiology 151:1021–1032
Gareis M, Gareis E-M (2007) Guttation droplets of Penicillium nordicum and Penicillium verrucosum contain high concentrations of the mycotoxins ochratoxin A and B. Mycopathologia 163:207–214
González-Medina M, Owen JR, El-Elimat T, Pearce CJ, Oberlies NH, Figueroa M, Medina-Franco JL (2017) Scaffold diversity of fungal metabolites. Front Pharmacol 8:180. https://doi.org/10.3389/fphar.2017.00180
González-Medina M, Prieto-Martínez FD, Naveja JJ, Méndez-Lucio O, El-Elimat T, Pearce CJ, Oberlies NH, Figueroa M, Medina-Franco JL (2016) Chemoinformatic expedition of the chemical space of fungal products. Future Med Chem 8:1399–1412
Hawksworth DL, Crous PW, Redhead SA, Reynolds DR, Samson RA, Seifert KA, Taylor JW, Wingfield MJ, Abaci Ö, Aime C, Asan A, Bai F-Y, de Beer ZW, Begerow D, Berikten D, Boekhout T, Buchanan PK, Burgess T, Buzina W, Cai L, Cannon PF, Crane JL, Damm U, Daniel H-M, van Diepeningen AD, Druzhinina I, Dyer PS, Eberhardt U, Fell JW, Frisvad JC, Geiser DM, Geml J, Glienke C, Gräfenhan T, Groenewald JZ, Groenewald M, de Gruyter J, Guého-Kellermann E, Guo L-D, Hibbett DS, Hong S-B, de Hoog GS, Houbraken J, Huhndorf SM, Hyde KD, Ismail A, Johnston PR, Kadaifciler DG, Kirk PM, Kõljalg U, Kurtzman CP, Lagneau P-E, Lévesque CA, Liu X, Lombard L, Meyer W, Miller A, Minter DW, Najafzadeh MJ, Norvell L, Ozerskaya SM, Öziç R, Pennycook SR, Peterson SW, Pettersson OV, Quaedvlieg W, Robert VA, Ruibal C, Schnürer J, Schroers H-J, Shivas R, Slippers B, Spierenburg H, Takashima M, Taşkın E, Thines M, Thrane U, Uztan AH, van Raak M, Varga J, Vasco A, Verkley G, Videira SIR, de Vries RP, Weir BS, Yilmaz N, Yurkov A, Zhang N (2011) The Amsterdam declaration on fungal nomenclature. IMA Fungus 2:105–112
Hemphill CFP, Sureechatchaiyan P, Kassack MU, Orfali RS, Lin W, Daletos G, Proksch P (2017) OSMAC approach leads to new fusarielin metabolites from Fusarium tricinctum. J Antibiot 70:726–732
Hewage RT, Aree T, Mahidol C, Ruchirawat S, Kittakoop P (2014) One strain-many compounds (OSMAC) method for production of polyketides, azaphilones, and an isochromanone using the endophytic fungus Dothideomycete sp. Phytochemistry 108:87–94
Hölker U, Höfer M, Lenz J (2004) Biotechnological advantages of laboratory-scale solid-state fermentation with fungi. Appl Microbiol Biotechnol 64:175–186
Hölker U, Lenz J (2005) Solid-state fermentation—are there any biotechnological advantages? Curr Opin Microbiol 8:301–306
Hongsanan S, Jeewon R, Purahong W, Xie N, Liu J-K, Jayawardena RS, Ekanayaka AH, Dissanayake A, Raspé O, Hyde KD, Stadler M, Peršoh D (2018) Can we use environmental DNA as holotypes? Fungal Divers. https://doi.org/10.1007/s13225-13018-10404-x
Hutwimmer S, Wang H, Strasser H, Burgstaller W (2010) Formation of exudate droplets by Metarhizium anisopliae and the presence of destruxins. Mycologia 102:1–10
Jordan TW, Cordiner SJ (1987) Fungal epipolythiodioxopiperazine toxins have therapeutic potential and roles in disease. Trends Pharmacol Sci 8:144–149
Joshi BK, Gloer JB, Wicklow DT (1999) New verticillin and glisoprenin analogues from Gliocladium catenulatum, a mycoparasite of Aspergillus flavus sclerotia. J Nat Prod 62:730–733
Katagiri K, Sato K, Hayakawa S, Matsushima T, Minato H (1970) Verticillin A, a new antibiotic from Verticillium sp. J Antibiot 23:420–422
Kertesz V, Berkel GJV (2014) Sampling reliability, spatial resolution, spatial precision, and extraction efficiency in droplet-based liquid microjunction surface sampling. Rapid Commun Mass Spectrom 28:1553–1560
Kertesz V, Van Berkel GJ (2010) Liquid microjunction surface sampling coupled with high-pressure liquid chromatography—electrospray ionization-mass spectrometry for analysis of drugs and metabolites in whole-body thin tissue sections. Anal Chem 82:5917–5921
Kertesz V, Weiskittel TM, Van Berkel GJ (2015) An enhanced droplet-based liquid microjunction surface sampling system coupled with HPLC-ESI-MS/MS for spatially resolved analysis. Anal Bioanal Chem 407:2117–2125
Khanna S, Srivastava AK (2005) Statistical media optimization studies for growth and PHB production by Ralstonia eutropha. Process Biochem 40:2173–2182
Kim J, Ashenhurst JA, Movassaghi M (2009) Total synthesis of (+)-11,11′-dideoxyverticillin A. Science 324:238–241
Kim J, Movassaghi M (2015) Biogenetically-inspired total synthesis of epidithiodiketopiperazines and related alkaloids. Acc Chem Res 48:1159–1171
Kinghorn AD, Carcache De Blanco EJ, Lucas DM, Rakotondraibe HL, Orjala J, Soejarto DD, Oberlies NH, Pearce CJ, Wani MC, Stockwell BR, Burdette JE, Swanson SM, Fuchs JR, Phelps MA, Xu L, Zhang X, Shen YY (2016) Discovery of anticancer agents of diverse natural origin. Anticancer Res 36:5623–5637
Koncz G, Hancz A, Chakrabandhu K, Gogolák P, Kerekes K, Rajnavölgyi É, Hueber A-O (2012) Vesicles released by activated T cells induce both Fas-mediated RIP-dependent apoptotic and Fas-independent nonapoptotic cell deaths. J Immunol 189:2815–2823
Lilly VG, Barnett HL (1951) Physiology of the fungi, vol 1. McGraw-Hill Book Company, Inc., New York
Lindenfelser L, Ciegler A (1975) Solid-substrate fermentor for ochratoxin A production. Appl Microbiol 29:323–327
Liu F, Liu Q, Yang D, Bollag WB, Robertson K, Wu P, Liu K (2011) Verticillin A overcomes apoptosis resistance in human colon carcinoma through DNA methylation-dependent upregulation of BNIP3. Cancer Res 71:6807–6816
Lu C, Paschall AV, Shi H, Savage N, Waller JL, Sabbatini ME, Oberlies NH, Pearce C, Liu K (2017) The MLL1-H3K4me3 axis-mediated PD-L1 expression and pancreatic cancer immune evasion. J Natl Cancer Inst 109:djw283
Mascarin GM, Jackson MA, Kobori NN, Behle RW, Dunlap CA, Júnior ÍD (2015) Glucose concentration alters dissolved oxygen levels in liquid cultures of Beauveria bassiana and affects formation and bioefficacy of blastospores. Appl Microbiol Biotechnol 99:6653–6665
Meletiadis J, Meis JF, Mouton JW, Verweij PE (2001) Analysis of growth characteristics of filamentous fungi in different nutrient media. J Clin Microbiol 39:478–484
Miao L, Kwong TF, Qian P-Y (2006) Effect of culture conditions on mycelial growth, antibacterial activity, and metabolite profiles of the marine-derived fungus Arthrinium cf saccharicola. Appl Microbiol Biotechnol 72:1063–1073
Minato H, Matsumoto M, Katayama T (1971) Verticillin A, a new antibiotic from Verticillium sp. J Chem Soc D 0:44–45
Moreira GM, Abreu LM, Carvalho VG, Schroers H-J, Pfenning LH (2016) Multilocus phylogeny of Clonostachys subgenus Bionectria from Brazil and description of Clonostachys chloroleuca sp. nov. Mycol Prog 15:1031–1039
Orjala J, Oberlies NH, Pearce CJ, Swanson SM, Kinghorn AD (2012) Discovery of potential anticancer agents from aquatic cyanobacteria, filamentous fungi, and tropical plants. In: Tringali C (ed) Bioactive compounds from natural sources, 2nd edn. CRC Press, Boca Raton, FL, pp 37–64
Paguigan ND, El-Elimat T, Kao D, Raja HA, Pearce CJ, Oberlies NH (2017) Enhanced dereplication of fungal cultures via use of mass defect filtering. J Antibiot 70:553–561
Paguigan ND, Raja HA, Day CS, Oberlies NH (2016) Acetophenone derivatives from a freshwater fungal isolate of recently described Lindgomyces madisonensis (G416). Phytochemistry 126:59–65
Paschall AV, Yang D, Lu C, Choi J-H, Li X, Liu F, Figueroa M, Oberlies NH, Pearce C, Bollag WB, Nayak-Kapoor A, Liu K (2015) H3K9 trimethylation silences Fas expression to confer colon carcinoma immune escape and 5-fluorouracil chemoresistance. J Immunol 195:1868–1882
Perdomo H, Cano J, Gené J, García D, Hernández M, Guarro J (2013) Polyphasic analysis of Purpureocillium lilacinum isolates from different origins and proposal of the new species Purpureocillium lavendulum. Mycologia 105:151–161
Pfefferle C, Theobald U, Gürtler H, Fiedler H-P (2000) Improved secondary metabolite production in the genus Streptosporangium by optimization of the fermentation conditions. J Biotechnol 80:135–142
Raja HA, Miller AN, Pearce CJ, Oberlies NH (2017) Fungal identification using molecular tools: a primer for the natural products research community. J Nat Prod 80:756–770
Robinson T, Singh D, Nigam P (2001) Solid-state fermentation: a promising microbial technology for secondary metabolite production. Appl Microbiol Biotechnol 55:284–289
Rossman AY, Seifert KA, Samuels GJ, Minnis AM, Schroers H-J, Lombard L, Crous PW, Põldmaa K, Cannon PF, Summerbell RC, Geiser DM, W-y Zhuang, Hirooka Y, Herrera C, Salgado-Salazar C, Chaverri P (2013) Genera in Bionectriaceae, Hypocreaceae, and Nectriaceae (Hypocreales) proposed for acceptance or rejection. IMA Fungus 4:41–51
Schroers H-J (2001) A monograph of Bionectria (Ascomycota, Hypocreales, Bionectriaceae) and its Clonostachys anamorphs. Studies in mycology. Centraalbureau voor Schimmelcultures, Utrecht
Sica VP, El-Elimat T, Oberlies NH (2016) In situ analysis of Asimina triloba (paw paw) plant tissues for acetogenins via the droplet-liquid microjunction-surface sampling probe coupled to UHPLC-PDA-HRMS/MS. Anal Methods 8:6143–6149
Sica VP, Figueroa M, Raja HA, El-Elimat T, Darveaux BA, Pearce CJ, Oberlies NH (2016) Optimizing production and evaluating biosynthesis in situ of a herbicidal compound, mevalocidin, from Coniolariella sp. J Ind Microbiol Biotechnol 43:1149–1157
Sica VP, Raja HA, El-Elimat T, Kertesz V, Van Berkel GJ, Pearce CJ, Oberlies NH (2015) Dereplicating and spatial mapping of secondary metabolites from fungal cultures in situ. J Nat Prod 78:1926–1936
Sica VP, Rees ER, Raja HA, Rivera-Chávez J, Burdette JE, Pearce CJ, Oberlies NH (2017) In situ mass spectrometry monitoring of fungal cultures led to the identification of four peptaibols with a rare threonine residue. Phytochemistry 143:45–53
Sica VP, Rees ER, Tchegnon E, Bardsley RH, Raja HA, Oberlies NH (2016) Spatial and temporal profiling of griseofulvin production in Xylaria cubensis using mass spectrometry mapping. Front Microbiol 7:544. https://doi.org/10.3389/fmicb.2016.00544
Siegel RL, Miller KD, Jemal A (2016) Cancer statistics, 2016. CA Cancer J Clin 66:7–30
Soltero FV, Johnson MJ (1953) The effect of the carbohydrate nutrition on penicillin production by Penicillium chrysogenum Q-176. Appl Microbiol 1:52
Son WB, Jensen RP, Kauffman CA, Fenical W (1999) New cytotoxic epidithiodioxopiperazines related to verticillin A from a marine isolate of the fungus Penicillium. Nat Prod Lett 13:213–222
Taylor JW (2011) One Fungus = One Name: DNA and fungal nomenclature 20 years after PCR. IMA Fungus 2:113–120
VanderMolen KM, Raja HA, El-Elimat T, Oberlies NH (2013) Evaluation of culture media for the production of secondary metabolites in a natural products screening program. AMB Express 3:71
Wang X, Sena Filho JG, Hoover AR, King JB, Ellis TK, Powell DR, Cichewicz RH (2010) Chemical epigenetics alters the secondary metabolite composition of guttate excreted by an atlantic-forest-soil-derived Penicillium citreonigrum. J Nat Prod 73:942–948
Wang Y, Hu P, Pan Y, Zhu Y, Liu X, Che Y, Liu G (2017) Identification and characterization of the verticillin biosynthetic gene cluster in Clonostachys rogersoniana. Fungal Genet Biol 103:25–33
White TJ, Bruns T, Lee SH, Taylor JW (1990) PCR protocols: A guide to methods and applications. Academic Press, San Diego
Yang LH, Miao L, Lee OO, Li X, Xiong H, Pang K, Vrijmoed L, Qian P (2007) Effect of culture conditions on antifouling compound production of a sponge-associated fungus. Appl Microbiol Biotechnol 74:1221–1231
Zewdu A, Lopez G, Braggio D, Kenny C, Constantino D, Bid HK, Batte K, Iwenofu OH, Oberlies NH, Pearce CJ, Strohecker AM, Lev D, Pollock RE (2016) Verticillin A inhibits leiomyosarcoma and malignant peripheral nerve sheath tumor growth via induction of apoptosis. Clin Exp Pharmacol 6:221–234
Zhang Y, Chen Y, Guo X, Zhang X, Zhao W, Zhong L, Zhou J, Xi Y, Lin Z, Ding J (2005) 11,11′-Dideoxy-verticillin: a natural compound possessing growth factor receptor tyrosine kinase-inhibitory effect with anti-tumor activity. Anticancer Drugs 16:515–524
Zheng CJ, Kim CJ, Bae KS, Kim YH, Kim WG (2006) Bionectins A–C, epidithiodioxopiperazines with anti-MRSA activity, from Bionectra byssicola F120. J Nat Prod 69:1816–1819
Zheng CJ, Park SH, Koshino H, Kim YH, Kim WG (2007) Verticillin G, a new antibacterial compound from Bionectria byssicola. J Antibiot 60:61–64
Acknowledgements
This research was supported by Grant P01 CA125066 from the National Cancer Institute/National Institutes of Health, Bethesda, MD, USA. The authors thank Dr. Joanna Burdette of the University of Illinois at Chicago and Dr. James Fuchs of Ohio State University for helpful discussions. We also thank Drs. Vilmos Kertesz and Gary J. Van Berkel (Mass Spectrometry and Laser Spectroscopy Group, Chemical Sciences Division, Oak Ridge National Laboratory) for inspiration and guidance with the droplet probe. The high-resolution mass spectrometry data were collected at the Triad Mass Spectrometry Laboratory at UNCG.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Amrine, C.S.M., Raja, H.A., Darveaux, B.A. et al. Media studies to enhance the production of verticillins facilitated by in situ chemical analysis. J Ind Microbiol Biotechnol 45, 1053–1065 (2018). https://doi.org/10.1007/s10295-018-2083-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10295-018-2083-8