Abstract
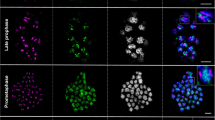
The karyotype of diploid Aster iinumae is morphologically similar to that of diploid Aster ageratoides var. ageratoides, however, its chromosome size is apparently smaller (S-type chromosomes versus L-type chromosomes, respectively). The hybrid origin of tetraploid Aster microcephalus var. ovatus (LS-type chromosomes) has previously been suggested by cytogenetics and chloroplast DNA (cp DNA) data. The cp DNA phylogeny also implies that the S-type chromosome is apomorphic, which means that genome size reduction occurred on the evolutionary way to A. iinumae. In this study, we have demonstrated that the chromosome size difference does not depend on the intensity of chromosome condensation but on the DNA content. The simultaneous genomic in situ hybridization (GISH) results show the similarity between S-type chromosomes of A. iinumae and A. microcephalus var. ovatus, and between L-type chromosomes of A. ageratoides and A. microcephalus var. ovatus, which provide additional evidence for A. microcephalus var. ovatus being a tetraploid amphidiploid produced by hybridization between S-type chromosomes and L-type chromosomes. The distribution patterns of Ty1-copia-like retrotransposons were similar in L- and S-type chromosomes. The copies of this retrotransposon dispersed uniformly on all chromosomes, and it is not yet apparent how the Ty1-copia-like retrotransposon affects the size difference between them.





Similar content being viewed by others
References
Ali HBM, Lysak MA, Schubert I (2004) Genomic in situ hybridization in plants with small genomes is feasible and elucidates the chromosomal parentage in interspecific Arabidopsis hybrids. Genome 40:954–960
Arnold ML (1997) Natural hybridization and evolution. Oxford University, Oxford
Bennetzen JL (2000) Transposable element contributions to plant gene and genome evolution. Plant Mol Biol 42:251–269
Bremer K (1994) Asteraceae. Cladistics & classification. Timber, Portland
Brysting AK, Holst-Jensen A, Leitch I (2000) Genomic organization of the hybrid Poa jemtlandica (Poaceae) verified by genomic in situ hybridization and chloroplast DNA sequences. Ann Bot 85:439–445
Desel C, Jansen R, Dedong G, Schmidt T (2002) Painting of parental chromatin in beta hybrids by multi-color fluorescent in situ hybridization. Ann Bot 89:171–181
Doležel J, Bartoš J (2005) Plant DNA flow cytometry and estimation of nuclear genome size. Ann Bot 95:99–110
Flavell RB (1986) Repetitive DNA and chromosome evolution in plants. Phil Trans R Soc Lond B Biol Sci 312:227–242
Flavell AJ, Pearce SR, Heslop-Harrison JS, Kumar A (1997) The evolution of Ty1-copia group retrotransposons in eukaryote genomes. Genetica 100:185–195
Galasso I, Harrison GE, Pignone D, Brandes A, Heslop-Harrison JS (1997) The distribution and organization of Ty1-copia-like retrotransposable elements in the genome of Vigna unguiculata (L.) Walp. (Cowpea) and its relatives. Ann Bot 80:327–333
Gatt M, Hammett K, Murray B (1999) Confirmation of ancient polyploidy in Dahlia (Asteraceae) species using genomic in situ hybridization. Ann Bot 84:39–48
Gu HY (1989) On chromosome numbers of Kalimeris (Astereae, Asteraceae) and some related taxa. Cathaya 1:1–16
Hanson L, Brown RL, Boyd A, Johnson MAT, Bennett MD (2003) First nuclear DNA c-values for 28 angiosperm genera. Ann Bot 91:31–38
Heslop-Harrison JS, Brandes A, Taketa S, Schmidt T, Vershinin AV, Alkhimova EG, Kamm A, Doudrick RL, Schwarzacher T, Katsiotis A, Kubis S, Kumar A, Pearce SR, Flavell AJ, Harrison GE (1997) The chromosomal distribution of Ty1-copia group retrotransposable elements in higher plants and their implications for genome evolution. Genetica 100:197–204
Huziwara Y (1957a) Karyotype analysis in some genera of Compositae. II. The karyotype of Japanese Aster species. Cytologia 22:96–112
Huziwara Y (1957b) Karyotype analysis in some genera of Compositae. III. The karyotype of the Aster ageratoides group. Am J Bot 44:783–790
Huziwara Y (1958) Karyotype analysis in some genera of Compositae. IV. The karyotypes within the genera Gymnaster, Kalimeris and Heteropappus. Cytologia 23:33–45
Huziwara Y (1967) Chromosomal evolution in Aster and related genera. Taxon 16:303–304
Ito M, Soejima A, Hasebe M, Watanabe K (1995) A chloroplast-DNA phylogeny of Kalimeris and Aster, with reference to generic circumscription. J Plant Res 108:93–96
Ito M, Soejima A, Watanabe K (1998) Phylogenetic relationships of Japanese Aster (Asteraceae, Astereae) sensu lato based on chloroplast–DNA restriction site mutations. J Plant Res 111:217–223
Johnston JS, Pepper AE, Hall AE, Chen ZJ, Hodnett G, Drabek J, Lopez R, Price HJ (2005) Evolution of genome size in Brassicaceae. Ann Bot 95:229–235
Kamm A, Doudrick RL, Heslop-Harrison JS, Schmidt T (1996) The genomic and physical organization of Ty1-copia-like sequences as a component of large genomes in Pinus elliottii var. elliottii and other gymnosperms. Proc Natl Acad Sci USA 93:2708–2713
Kitamura S (1937) Compositae Japonicae. I. Mem Coll Sci Kyoto Imp Univ Ser B 13:337–357
Leitch IJ, Heslop-Harrison JS (1992) Physical mapping of the 18S–5.8S–26S rRNA genes in barley by in situ hybridization. Genome 35:1013–1018
Leitch IJ, Schwarzacher T, Jackson D, Leitch IJ (1994) In situ hybridization: a practical guide. BIOs Scientific, UK, pp 50–108
Li CB, Zhang DM, Ge S, Lu BR, Hong DY (2001) Differentiation and inter genomic relationships among C, E and D genomes in the Oryza officinalis complex (Poaceae) as revealed by multicolor genomic in situ hybridization. Theor Appl Genet 103:197–203
Maluszynska J, Hasterok R (2005) Identification of individual chromosomes and parental genomes in Brassica juncea using GISH and FISH. Cytogenet Genome Res 109:310–314
Marasek A, Hasterok R, Wiejacha K, Orlikowska T (2004) Determination by GISH and FISH of hybrid status in Lilium. Hereditas 140:1–7
Matoba H, Soejima A, Hoshi Y, Kondo K (2005) Molecular cytogenetic organization of 5S and 18S rDNA loci in Aster ageratoides var. ageratoides, A. iinumae (=Kalimeris pinnatifida) and A. microcephalus var. ovatus in Japan. Cytologia 70:323–330
Meinkoth J, Wahl G (1984) Hybridization of nuclei acids immobilized on solid supports. Anal Biochem 138:267–284
Miller JT, Dong F, Jackson SA, Song J, Jiang J (1998) Retrotransposon related DNA sequences in the centromeres of grass chromosomes. Genetics 150:1615–1623
Norrmann N, Hanson L, Renvoize S, Leitch IJ (2004) Genomic relationships among diploid and hexaploid species of Andropogon (Poaceae). Genome 47:1220–1224
Pearce SR, Harrison G, Li D, Heslop-Harrison JS, Kumar A, Flavell AJ (1996) The Ty1-copia group retrotransposons in Vicia species: copy number, sequence heterogeneity and chromosomal location. Mol Gen Genet 250:305–315
Price HJ, Dillon SL, Hodnett G, Rooney W, Ross L, Johnston JS (2005) Genome evolution in the genus Sorghum (Poaceae). Ann Bot 95:219–227
Refoufi A, Jahier J, Esnault MA (2001) Genomic analysis of Elytrigia pycnantha and Thinopyrum junceiforme and of their putative natural hybrid using the GISH technique. Genome 44:708–715
Richard D, Rieseberg LH (1999) ITS sequence data support a single origin for North American Astereae (Asteraceae) and reflect deep geographic divisions in Aster s. l. Am J Bot 83:398–412
Rieseberg LH (1997) Hybrid origins of plant species. Annu Rev Ecol Syst 28:359–389
Shaw CH (ed.) (1988) Plant molecular biology. A practical approach. Oxford University Press, Oxford
Shindo K (1967) Cytological, morphological and geographical studies on the differentiation of species in section Asteromoea of Kalimeris in Japan. J Sci Hiroshima Univ, Ser B Div 2 11:127–199
Soltis PS, Soltis DE (1995) The role of genetic and genomic attributes in the success of polyploids. Proc Natl Acad Sci U S A 97:7051–7057
Soltis PS, Doyle JJ, Soltis DE (1992) Molecular data and polyploid evolution in plants. In: Soltis PS, Soltis DE, Doyle JJ (eds) Molecular systematics of plants, 177–201. Chapman and Hall, New York, p. 434
Tara M (1977) Cytologenetic studies on natural intergeneric hybridization on Aster alliances. IV. Experimental confirmation of the hybrid origin of Aster ageratoides subsp. ovatus. Bot Mag Tokyo 90:253–258
Testolin R, Cipriani G (1997) Paternal inheritance of chloroplast DNA and maternal inheritance of mitochondrial DNA in the genus Actinidia. Theor Appl Genet 94:897–903
Vicient CM, Suoniemi A, Anamthawat-Jónsson K, Tanskanen J, Beharav A, Nevo E, Schulman AH (1999) Retrotransposon BARE-1 and its role in genome evolution in the genus Hordeum. Plant Cell 11:1769–1784
Wendel JF, Cronn RC, Johnston JS, Price HJ (2002) Feast and famine in plant genomes. Genetica 115:37–47
Acknowledgments
This study was financially supported by the Ministry of Education, Science, Sports and Culture, Grant-in-Aid for Scientific Research (C), 2006, 18570094.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Matoba, H., Soejima, A. & Hoshi, Y. Identification of parental genomes and genomic organization in Aster microcephalus var. ovatus . J Plant Res 120, 585–593 (2007). https://doi.org/10.1007/s10265-007-0101-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10265-007-0101-4