Abstract
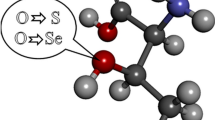
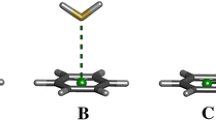
Selenium has been increasingly recognized as an important element in biological systems, which participates in numerous biochemical processes in organisms, notably in enzyme reactions. Selenium can substitute sulfur of cysteine and methionine to form their selenium analogues, selenocysteine (Sec) and selenomethionine (SeM). The nature of amino acid pockets in proteins is dependent on their composition and thus different non-covalent forces determine the interactions between selenium of Sec or SeM and other functional groups, resulting in specific biophysical behavior. The discrimination of selenium toward sulfur has been reported. In order to elucidate the difference between the nature of S-π and Se-π interactions, we performed extensive DFT calculations of dispersive and electrostatic contributions of Se-π interactions in substituted benzenes/hydrogen selenide (H2Se) complexes. The results are compared with our earlier reported S-π calculations, as well as with available experimental data. Our results show a larger contribution of dispersive interactions in Se-π systems than in S-π ones, which mainly originate from the attraction between Se and substituent groups. We found that selenium exhibits a strong interaction with aromatic systems and may thus play a significant role in stabilizing protein folds and protein–inhibitor complexes. Our findings can also provide molecular insights for understanding enzymatic specificity discrimination between single selenium versus a sulfur atom, notwithstanding their very similar chemical properties.











Similar content being viewed by others
Notes
In this text we shall use explicit notation: M(lone pair)····π(aryl), and M–H····π(aryl) (where M stands for a chalcogen atom) whenever we refer to the specific or dominant type of noncovalent interaction studied in this work, and an en-dash or other symbol(s) for interactions of unspecified type, and/or citations from other authors.
References
Hatfield DL, Schweizer U, Tsuji PA, Gladyshev VN (2016) Selenium— its molecular biology and role in human health. Springer, Berlin
Brigelius-Flohe R, Sies H (2016) Diversity of selenium functions in health and disease, oxidative stress and disease, no. 38. CRC, Boca Raton
Arnér ESJ (2010) Selenoproteins — what unique properties can arise with selenocysteine in place of cysteine? Exp Cell Res 316:1296–1303. doi:10.1016/j.yexcr.2010.02.032
Cupp-Sutton KA, Ashby MT (2016) Biological chemistry of hydrogen selenide. Antioxidants 5:42–60. doi:10.3390/antiox5040042
Johansson L, Gafvelin G, Arnér ESJ (2005) Selenocysteine in proteins–properties and biotechnological use. Biochim Biophys Acta 1726:1–13. doi:10.1016/j.bbagen.2005.05.010
Johnson DW, Hof F (2016) Aromatic interactions: frontiers in knowledge and application, monographs in supramolecular chemistry, no. 20. The Royal Society of Chemistry, UK
Waters ML (2004) Aromatic interactions in peptides: impact on structure and function. Biopolymers 76:435–445. doi:10.1002/bip.20144
Hartman I, Raia CA, Zauhar RJ (2006) Evidence for a strong selenium–aromatic interaction derived from crystallographic data and ab initio quantum chemical calculations. Biopolymers 83:595–613. doi:10.1002/bip.20144
Saberinasab M, Salehzadeh S, Maghsoud Y, Bayat M (2016) The significant effect of electron donating and electron withdrawing substituents on nature and strength of an intermolecular Se∙∙∙∙πinteraction. A theoretical study. Comp Theor Chem 1078:9–15. doi:10.1016/j.comptc.2015.12.009
Saberinasab M, Salehzadeh S, Solimannejad M (2016) The effect of a strong cation∙∙∙∙π interaction on a weak selenium∙∙∙∙π interaction: a theoretical study. Comp Theor Chem 1092:41–46. doi:10.1016/j.comptc.2016.07.027
Caracelli I, Haiduc I, Zukerman-Schpector J, Tiekink ERT (2014) Supramolecular architectures based on M(lone pair)···π(arene) interactions for M=Se and Te. In: Patai S, Rappoport Z (eds) Chemistry of organic selenium and tellurium compounds, 4th edn. Wiley, New York, pp 973–988
Caracelli I, Haiduc I, Zukerman-Schpector J, Tiekink ERT (2013) Delocalised antimony(lone pair)- and bismuth-(lone pair)···π (arene) interactions: supramolecular assembly and other considerations. Coord Chem Rev 257:2863–2879. doi:10.1016/j.ccr.2013.05.022
Zukerman-Schpector J, Haiduc I, Tiekink ERT (2012) Supramolecular self-assembly of transition metal carbonyl molecules through M-CO(lone pair)···π (arene) interactions. Adv Phys Org Chem 60:49–92. doi:10.1016/B978-0-12-396970-5.00002-5
Caracelli I, Zukerman-Schpector J, Tiekink ERT (2012) Supramolecular aggregation patterns based on the bio-inspired Se (lone pair) ⋯ π (aryl) synthon. Coord Chem Rev 256:412–438. doi:10.1016/j.ccr.2011.10.021
Caracelli I, Haiduc I, Zukerman-Schpector J, Tiekink ERT (2016) A new non-covalent bonding mode in supramolecular chemistry: main group element lone-pair···π(arene) interactions. In: Johnson DW, Hof F (eds) Aromatic interactions: frontiers in knowledge and application, monographs in supramolecular chemistry, no. 20. The Royal Society of Chemistry, UK, pp 89–124
Salonen LM, Ellermann M, Diederich F (2011) Aromatic rings in chemical and biologicalrecognition: energetics and structures. Angew Chem Int Ed 50:4808–4842. doi:10.1002/anie.201007560
Wheeler SE (2013) Understanding substituent effects in noncovalent interactions involving aromatic rings. Acc Chem Res 46:1029–1038. doi:10.1021/ar300109n
Wheeler SE, Houk KN (2008) Substituent effects in the benzene dimer are due to direct interactions of the substituents with the unsubstituted benzene. J Am Chem Soc 130:10854–10855. doi:10.1021/ja802849j
Wheeler SE (2012) Understanding substituent effects in noncovalent interactions involving aromatic rings. Acc Chem Res 46:1029–1038. doi:10.1021/ar300109n
Senćanski M, Došen-Mićović L, Šukalović V, Kostić-Rajačić S (2015) Theoretical insight into sulfur–aromatic interactions with extension to D2 receptor activation mechanism. Struct Chem 26:1139–1149. doi:10.1007/s11224-015-0574-z
Ringer AL, Senenko A, Sherrill CD (2007) Models of S/π interactions in protein structures: comparison of the H2S–benzene complex with PDB data. Protein Sci 16:2216–2223. doi:10.1110/ps.073002307
Hansch C, Leo A, Taft RW (1991) A survey of Hammett substituent constants and resonance and field parameters. Chem Rev 91:165–195. doi:10.1021/cr00002a004
Cheng Q, Sandalova T, Lindqvist Y, Arnér ESJ (2009) Crystal structure and catalysis of the selenoprotein thioredoxin reductase 1. J Biol Chem 284:3998–4008. doi:10.1074/jbc.M807068200
Hassan AE, Sheng J, Jiang J, Zhang JW, Huang Z (2009) Synthesis and crystallographic analysis of 5-Se-thymidine DNAs. Org Lett 11:2503–2506. doi:10.1021/ol9004867
Schaefer-Ramadan S, Thorpe C, Rozovsky S (2014) Site-specific insertion of selenium into the redoxactive disulfide of the flavoprotein augmenter of liver regeneration. Arch Biochem Biophys 548:60–65. doi:10.1016/j.abb.2014.02.001
Schaefer SA, Dong M, Rubenstein RP, Wilkie WA, Bahnson BJ, Thorpe C, Rozovsky S (2013) (77)Se enrichment of proteins expands the biological NMR toolbox. J Mol Biol 425:222–231. doi:10.1016/j.jmb.2012.11.011
Mander L, Liu HW (2010) Comprehensive natural products II: chemistry and biology, vol. 1. Newnes, Oxford
Daithankar VN, Schaefer SA, Dong M, Bahnson BJ, Thorpe C (2010) Structure of the human sulfhydryl oxidase augmenter of liver regeneration and characterization of a human mutation causing an autosomal recessive myopathy. Biochemistry-US 49:6737–6745. doi:10.1021/bi100912m
Banci L, Bertini I, Calderone V, Cefaro C, Ciofi-Baffoni S, Gallo A, Tokatlidis K (2012) An electrontransfer path through an extended disulfide relay system: the case of the redox protein ALR. J Am Chem Soc 134:1442–1445. doi:10.1021/ja209881f
Dmitrenko O, Thorpe C, Bach RD (2003) Effect of a charge-transfer interaction on the catalytic activity of Acyl-CoA dehydrogenase: a theoretical study of the role of oxidized flavin. J Phys Chem B 107:13229–13236. doi:10.1021/jp0348631
Frisch MJ, Trucks GW, Schlegel HB, Scuseria GE, Robb MA, Cheeseman JR, Scalmani G, Barone V, Mennucci B, Petersson GA, Nakatsuji H, Caricato M, Li X, Hratchian HP, Izmaylov AF, Bloino J, Zheng G, Sonnenberg JL, Hada M, Ehara M, Toyota K, Fukuda R, Hasegawa J, Ishida M, Nakajima T, Honda Y, Kitao O, Nakai H, Vreven T, Montgomery Jr JA, Peralta JE, Ogliaro F, Bearpark M, Heyd JJ, Brothers E, Kudin KN, Staroverov VN, Kobayashi R, Normand J, Raghavachari K, Rendell A, Burant JC, Iyengar SS, Tomasi J, Cossi M, Rega N, Millam JM, Klene M, Knox JE, Cross JB, Bakken V, Adamo C, Jaramillo J, Gomperts R, Stratmann RE, Yazyev O, Austin AJ, Cammi R, Pomelli C, Ochterski JW, Martin RL, Morokuma K, Zakrzewski VG, Voth GA, Salvador P, Dannenberg JJ, Dapprich S, Daniels AD, Farkas O, Foresman JB, Ortiz JV, Cioslowski J, Fox DJ (2009) Gaussian 09, revision E.01. Gaussian Inc., Wallingford
Grimme S (2006) Semiempirical GGA type density functional constructed with a long-range dispersion correction. J Comp Chem 27:1787–1799. doi:10.1002/jcc.20495
Schaefer A, Horn H, Ahlrichs R (1992) Fully optimized contracted Gaussian basis sets for atoms Li to Kr. J Chem Phys 97:2571–2577. doi:10.1063/1.463096
Schaefer A, Huber C, Ahlrichs R (1994) Fully optimized contracted Gaussian basis sets of triple zeta valence quality for atoms Li to Kr. J Chem Phys 100:5829–5835. doi:10.1063/1.467146
Dunning Jr TH, Hay PJ (1977) Gaussian basis sets for molecular calculations. In: Schaefer HF (ed) Modern theoretical chemistry, vol 3. Plenum, New York, pp 1–28
Hay PJ, Wadt WR (1985) Ab initio effective core potentials for molecular calculations. Potentials for the transition metal atoms Sc to Hg. J Chem Phys 82:270–283. doi:10.1063/1.448799
Wadt WR, Hay PJ (1985) Ab initio effective core potentials for molecular calculations. Potentials for main group elements Na to Bi. J Chem Phys 82:284–298. doi:10.1063/1.448800
Hay PJ, Wadt WR (1985) Ab initio effective core potentials for molecular calculations. Potentials for K to Au including the outermost core orbitals. J Chem Phys 82:299–310. doi:10.1063/1.448975
PARADOX cluster at the Scientific Computing Laboratory of the Institute of Physics Belgrade, supported in part by the Serbian Ministry of Education and Science under project No. ON171017, and by the European Commission under FP7 projects HP-SEE, PRACE-1IP, PRACE-2IP, EGI-InSPIRE
Acknowledgments
This work was financially supported by the Serbian Ministry of Education, Science and Technological Development, Republic of Serbia, Contracts No 173001 and 172035, and COST Action CM1405, Molecules in Motion (MOLIM). The authors would like to thank Prof. Ljiljana Došen-Mićović for help with this project, and the anonymous referee for valuable suggestions that improved the clarity of the manuscript. Graphs were rendered in Origin 8, figures were made using Discovery Studio 2016.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
ESM 1
(PDF 335 kb)
Rights and permissions
About this article
Cite this article
Senćanski, M., Djordjević, I. & Grubišić, S. Assessing the dispersive and electrostatic components of the selenium–aromatic interaction energy by DFT. J Mol Model 23, 162 (2017). https://doi.org/10.1007/s00894-017-3330-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00894-017-3330-z