Abstract
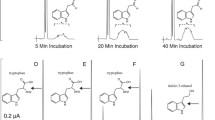
The metabolic degradation of aldehydes is catalyzed by oxidoreductases from which aldehyde dehydrogenases (EC 1.2.1) comprise nonspecific or substrate-specific enzymes. The latter subset is represented, e.g., by NAD+-dependent aminoaldehyde dehydrogenases (AMADHs; EC 1.2.1.19) oxidizing a group of naturally occurring ω-aminoaldehydes including polyamine oxidation products. Recombinant isoenzymes from pea (PsAMADH1 and 2) and tomato (LeAMADH1 and 2) were subjected to kinetic measurements with synthetic aldehydes containing a nitrogenous heterocycle such as pyridinecarbaldehydes and their halogenated derivatives, (pyridinylmethylamino)-aldehydes, pyridinyl propanals and aldehydes derived from purine, 7-deazapurine and pyrimidine to characterize their substrate specificity and significance of the resulting data for in vivo reactions. The enzymatic production of the corresponding carboxylic acids was analyzed by liquid chromatography coupled to electrospray ionization mass spectrometry. Although the studied AMADHs are largely homologous and supposed to have a very similar active site architecture, significant differences were observed. LeAMADH1 displayed the broadest specificity oxidizing almost all compounds followed by PsAMADH2 and 1. In contrast, LeAMADH2 accepted only a few compounds as substrates. Pyridinyl propanals were converted by all isoenzymes, usually better than pyridinecarbaldehydes and aldehydes with fused rings. The K m values for the best substrates were in the range of 10−5–10−4 M. Nevertheless, the catalytic efficiency values (V max/K m) reached only a very small fraction of that with 3-aminopropanal (except for LeAMADH1 activity with two pyridine-derived compounds). Docking experiments using the crystal structure of PsAMADH2 were involved to discuss differences in results with position isomers or alkyl chain homologs.






Similar content being viewed by others
Abbreviations
- AMADH:
-
Aminoaldehyde dehydrogenase
- AEAL:
-
2-Aminoethanal
- ABAL:
-
4-Aminobutanal
- ALDH:
-
Aldehyde dehydrogenase
- APAL:
-
3-Aminopropanal
- BADH:
-
Betaine aldehyde dehydrogenase
- LeAMADH:
-
Tomato (Lycopersicon esculentum) aminoaldehyde dehydrogenase
- LC–MS:
-
Liquid chromatography coupled to mass spectrometry
- PsAMADH:
-
Pea (Pisum sativum) aminoaldehyde dehydrogenase
References
Bradford MM (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 72:248–254. doi:10.1016/0003-2697(76)90527-3
Brocker C, Lassen N, Estey T, Pappa A, Cantore M, Orlova VV, Chavakis T, Kavanagh KL, Oppermann U, Vasiliou V (2010) Aldehyde dehydrogenase 7A1 (ALDH7A1) is a novel enzyme involved in cellular defense against hyperosmotic stress. J Biol Chem 285:18452–18463. doi:10.1074/jbc.M109.077925
Case DA, Cheatham TE, Darden T, Gohlke H, Luo R, Merz KM, Onufriev A, Simmerling C, Wang B, Woods RJ (2005) The amber biomolecular simulation programs. J Comput Chem 26:1668–1688. doi:10.1002/jcc.20290
Chern MK, Pietruszko R (1995) Human aldehyde dehydrogenase E3 isozyme is a betaine aldehyde dehydrogenase. Biochem Biophys Res Commun 213:561–568. doi:10.1006/bbrc.1995.2168
Crews P, Rodríguez J, Jaspars M (2010) Organic structure analysis, 2nd edn. Oxford University Press, New York, p 273
DeLano W (2002) The PyMOL molecular graphics system. DeLano Scientific, Palo Alto. http://www.pymol.org
Farrés J, Wang X, Takahashi K, Cunningham SJ, Wang TT, Weiner H (1994) Effects of changing glutamate 487 to lysine in rat and human liver mitochondrial aldehyde dehydrogenase. A model to study human (oriental type) class 2 aldehyde dehydrogenase. J Biol Chem 269:13854–13860
Frisch MJ, Trucks GW, Schlegel HB, Scuseria GE, Robb MA, Cheeseman JR, Montgomery JA Jr, Vreven T, Kudin KN, Burant JC, Millam JM, Iyengar SS, Tomasi J, Barone V, Mennucci B, Cossi M, Scalmani G, Rega N, Petersson GA, Nakatsuji H, Hada M, Ehara M, Toyota K, Fukuda R, Hasegawa J, Ishida M, Nakajima T, Honda Y, Kitao O, Nakai H, Klene M, Li X, Knox JE, Hratchian HP, Cross JB, Bakken V, Adamo C, Jaramillo J, Gomperts R, Stratmann RE, Yazyev O, Austin AJ, Cammi R, Pomelli C, Ochterski JW, Ayala PY, Morokuma K, Voth GA, Salvador P, Dannenberg JJ, Zakrzewski VG, Dapprich S, Daniels AD, Strain MC, Farkas O, Malick DK, Rabuck AD, Raghavachari K, Foresman JB, Ortiz JV, Cui Q, Baboul AG, Clifford S, Cioslowski J, Stefanov BB, Liu G, Liashenko A, Piskorz P, Komaromi I, Martin RL, Fox DJ, Keith T, Al-Laham MA, Peng CY, Nanayakkara A, Challacombe M, Gill PMW, Johnson B, Chen W, Wong MW, Gonzalez C, Pople JA (2004) Gaussian 03, Revision C.02. Gaussian Inc., Wallingford
Gruez A, Roig-Zamboni V, Grisel S, Salomoni A, Valencia C, Campanacci V, Tegoni M, Cambillau C (2004) Crystal structure and kinetics identify Escherichia coli YdcW gene product as a medium-chain aldehyde dehydrogenase. J Mol Biol 343:29–41. doi:10.1016/j.jmb.2004.08.030
Hill JP, Dickinson FM (1988) Pre-steady-state kinetics of aldehyde oxidation by pig liver cytosolic aldehyde dehydrogenase. Biochem Soc Trans 16:856–857
Ho BK, Gruswitz F (2008) HOLLOW: generating accurate representations of channel and interior surfaces in molecular structures. BMC Struct Biol 8:49. doi:10.1186/1472-6807-8-49
Kaiser JP, Feng Y, Bollag JM (1996) Microbial metabolism of pyridine, quinoline, acridine, and their derivatives under aerobic and anaerobic conditions. Microbiol Rev 60:483–498
Kaplan NO, Goldin A, Humphreys SR, Stolzenbach FE (1957) Pyridine precursors of mouse liver diphosphopyridine nucleotide. J Biol Chem 226:365–371
Kirch HH, Nair A, Bartels D (2001) Novel ABA- and dehydration-inducible aldehyde dehydrogenase genes isolated from the resurrection plant Craterostigma plantagineum and Arabidopsis thaliana. Plant J 28:555–567. doi:10.1046/j.1365-313X.2001.01176.x
Kirch HH, Bartels D, Wei Y, Schnable PS, Wood AJ (2004) The ALDH gene superfamily of Arabidopsis. Trends Plant Sci 9:371–377. doi:10.1007/s11103-004-7796-6
Klyosov AA (1996) Kinetics and specificity of human liver aldehyde dehydrogenase toward aliphatic, aromatic, and fused polycyclic aldehydes. Biochemistry 35:4457–4467. doi:10.1021/bi9521102
Kopečný D, Tylichová M, Snégaroff J, Popelková H, Šebela M (2011) Carboxylate and aromatic active-site residues are determinants of high-affinity binding of ω-aminoaldehydes to plant aminoaldehyde dehydrogenases. FEBS J 278:3130–3139. doi:10.1111/j.1742-4658.2011.08239.x
Mancuso AJ, Huang SL, Swern D (1978) Oxidation of long-chain and related alcohols to carbonyls by dimethyl sulfoxide “activated” by oxalyl chloride. J Org Chem 43:2480–2482. doi:10.1021/jo00406a041
Marchitti SA, Orlicky DJ, Vasiliou V (2007) Expression and initial characterization of human ALDH3B1. Biochem Biophys Res Commun 356:792–798. doi:10.1016/j.bbrc.2007.03.046
Marchitti SA, Brocker C, Stagos D, Vasiliou V (2009) Non-P450 aldehyde oxidizing enzymes: the aldehyde dehydrogenase superfamily. Expert Opin Drug Metab Toxicol 4:697–720. doi:10.1517/17425250802102627
Morris GM, Goodsell DS, Halliday RS, Huey R, Hart WE, Belew RK, Olson AJ (1998) Automated docking using a Lamarckian genetic algorithm and an empirical binding free energy function. J Comput Chem 19:1639–1662. doi:10.1002/(SICI)1096-987X(19981115)19:14<1639:AID-JCC10>3.0.CO;2-B
Panoutsopoulos GI, Kouretas D, Beedham C (2004) Contribution of aldehyde oxidase, xanthine oxidase, and aldehyde dehydrogenase on the oxidation of aromatic aldehydes. Chem Res Toxicol 2004:1368–1376. doi:10.1021/tx030059u
Pearlman DA, Case DA, Caldwell JW, Ross WS, Cheatham TE, Debolt S, Ferguson D, Seibel G, Kollman P (1995) Amber, a package of computer programs for applying molecular mechanics, normal mode analysis, molecular dynamics and free energy calculations to simulate the structural and energetic properties of molecules. Comput Phys Commun 91:1–41. doi:10.1016/0010-4655(95)00041-D
Perozich J, Nicholas H, Wang BC, Lindahl R, Hempel J (1999) Relationships within the aldehyde dehydrogenase extended family. Protein Sci 8:137–146. doi:10.1110/ps.8.1.137
Prokop M, Adam J, Kříž Z, Wimmerová M, Koča J (2008) TRITON: a graphical tool for ligand-binding protein engineering. Bioinformatics 24:1955–1956. doi:10.1093/bioinformatics/btn344
Rhodes D, Hanson AD (1993) Quarternary ammonium and tertiary sulfonium compounds in higher plants. Annu Rev Plant Physiol Plant Mol Biol 44:357–384. doi:10.1146/annurev.pp.44.060193.002041
Sánchez-Sandoval A, Alvarez-Toledano C, Gutiérrez-Pérez Y, Reyes-Ortega Y (2003) A modified procedure for the preparation of linear polyamines. Synth Commun 33:481–492. doi:10.1081/SCC-120015780
Šebela M, Brauner F, Radová A, Jacobsen S, Havliš J, Galuszka P, Peč P (2000) Characterization of a homogenous plant aminoaldehyde dehydrogenase. Biochim Biophys Acta 1480:329–341. doi:10.1016/S0167-4838(00)00086-8
Šebela M, Štosová T, Havliš J, Wielsch N, Thomas H, Zdráhal Z, Shevchenko A (2006) Thermostable trypsin conjugates for high-throughput proteomics: synthesis and performance evaluation. Proteomics 6:2959–2963. doi:10.1002/pmic.200500576
Sophos NA, Vasiliou V (2003) Aldehyde dehydrogenase gene superfamily: the 2002 update. Chem Biol Interact 143–144:5–22. doi:10.1016/S0009-2797(02)00163-1
Sunkar R, Bartels D, Kirch HH (2003) Overexpression of a stress-inducible aldehyde dehydrogenase gene from Arabidopsis thaliana in transgenic plants improves stress tolerance. Plant J 35:452–464. doi:10.1046/j.1365-313X.2003.01819.x
Tylichová M, Briozzo P, Kopečný D, Ferrero J, Moréra S, Joly N, Snégaroff J, Šebela M (2008) Purification, crystallization and preliminary crystallographic study of a recombinant plant aminoaldehyde dehydrogenase from Pisum sativum. Acta Crystallogr Sect F Struct Biol Cryst Commun 64:88–90. doi:10.1107/S1744309107068522
Tylichová M, Kopečný D, Moréra S, Briozzo P, Lenobel R, Snégaroff J, Šebela M (2010) Structural and functional characterization of plant aminoaldehyde dehydrogenase from Pisum sativum with a broad specificity for natural and synthetic aminoaldehydes. J Mol Biol 396:870–882. doi:10.1016/j.jmb.2009.12.015
Vasiliou V, Nebert DW (2005) Analysis and update of the human aldehyde dehydrogenase (ALDH) gene family. Hum Genomics 2:138–143
Vriend G (1990) WHAT IF—a molecular modeling and drug design program. J Mol Graph 8:52–56
Warburg O, Christian W (1943) Isolierung und Kristallisation des Gärungsferments Zymohexase. Biochem Z 314:149–176 (in German)
Weiner SJ, Kollman PA, Case DA, Singh UC, Ghio C, Alagona G, Profeta S, Weiner P (1984) A new force-field for molecular mechanical simulation of nucleic acids and proteins. J Am Chem Soc 106:765–784. doi:10.1021/ja00315a051
Wood AJ, Duff RJ (2009) The aldehyde dehydrogenase (ALDH) gene superfamily of the moss Physcomitrella patens and the algae Chlamydomonas reinhardtii and Ostreococcus tauri. Bryologist 112:1–11. doi:10.1639/0007-2745-112.1.1
Acknowledgments
This work was supported by OP RD&I grant no. ED0007/01/01 (Centre of the Region Haná for Biotechnological and Agricultural Research) and grant no. MSM0021622413 from the Ministry of Education, Youth and Sports, Czech Republic, plus grant no. 522/08/0555 from the Czech Science Foundation. We would also like to thank Hana Moskalíková, a former student of the Department of Biochemistry, Faculty of Science, Palacký University, for her valuable contribution to initial experiments.
Conflict of interest
The authors declare no conflict of interest.
Author information
Authors and Affiliations
Corresponding authors
Additional information
Chemical names of all synthetic aldehyde compounds are abbreviated by acronyms (elucidated directly in the text) ending with a suffix AL.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Frömmel, J., Soural, M., Tylichová, M. et al. Plant aminoaldehyde dehydrogenases oxidize a wide range of nitrogenous heterocyclic aldehydes. Amino Acids 43, 1189–1202 (2012). https://doi.org/10.1007/s00726-011-1174-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00726-011-1174-x