Abstract
We use a genetic algorithm to explore the subspace of combination and parametrization patterns spanned by a set of popular exchange and correlation functional approximations. Using the well-balanced GMTKN30 benchmark database to guide the evolutionary process, we find that the genetic algorithm is able to recover variants of several popular generalized gradient approximation functionals and hybrid functionals. For the latter class, the algorithm is able to identify a reparametrized version of the three-parameter hybrid B3PW91, which shows significantly improved performance compared to conventional versions of B3PW91. Furthermore, the possible application of this algorithm to automatically construct so-called “niche”-functionals—specially tailored to specific applications—is demonstrated.
Graphical abstract

Similar content being viewed by others
Introduction
Kohn–Sham density functional theory (DFT) [1] ranks amongst the most popular and most frequently employed methods in computational chemistry [2, 3]. Due to its favorable ratio of computational efficiency to accuracy, it can be used routinely in the quantum mechanical treatment of large molecular systems and the ground state dynamical simulation of long timescales.
Yet, although an “exact” functional providing a perfect description of the electronic ground state of a system does exist in principle, little is known of its actual form or how to derive it systematically. To ameliorate this drawback, empirical approximations have to be introduced to render DFT feasible for practical applications. These approximations are usually designed based on physical intuition and/or fitting to reference data sets, introducing several empirical parameters in the process. DFT owes a significant portion of its widespread success to the emergence of various highly successful empirical functional approximations, which can in turn be combined and reparametrized to create new functionals [4]. Accordingly, the search and parametrization strategies to identify promising new functionals in the vast functional space are almost as diverse as the approximations themselves, ranging from highly systematic brute-force approaches [5, 6] to procedures leveraging, e.g., insights from game theory [7].
Another class of algorithms that shows excellent performance for similar high-dimensional search spaces are genetic algorithms [8]. This type of optimization algorithm is based on the concept of Darwinian evolution [9] and has been applied to a diverse range of problems in computational chemistry, with geometry optimization, docking studies, and catalyst design only being some of the most prominent examples [10].
In the present work, we investigate whether genetic algorithms can also be used to automate the search for promising density functional combinations and parametrization patterns. To facilitate an assessment of the algorithms performance, we restrict the optimization problem to a subregion of the functional space spanned by several popular exchange and correlation functional approximations. To this end, we employ a specially developed genetic algorithm to search for generalized gradient approximation (GGA) and hybrid functionals using the GMTKN30 benchmark database [11] as guidance. Based on the evolved functionals, their performance, emerging trends in functional composition and parametrization patterns, as well as their relationships to conventional functionals are analyzed.
One particularly promising feature of the genetic algorithm presented here is the ability to construct functionals tailored to specific needs and systems. The resulting “niche” functionals can be obtained in a completely automated fashion and exhibit excellent performance for their intended task. We demonstrate the potential of this approach by evolving a functional focused exclusively on describing long-range dispersion interactions.
Genetic algorithm
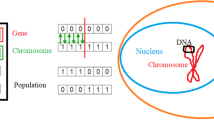
Genetic algorithms are heuristic optimization algorithms designed according to the principles of Darwinian evolution [8]. Potential solutions to the optimization problem, so-called individuals, compete with each other in a survival of the fittest scenario in order to evolve solutions of increasing quality.
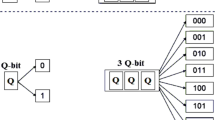
In genetic algorithms, every individual is represented by a genome, which encodes a particular solution in discrete form, e.g., a list of binary numbers. The genetic algorithm is initiated by creating a set of individuals (a population) with randomly generated genomes. In the next step, a fitness score is assigned to each individual by a fitness function. This fitness score is a measure of the quality of the current individual (e.g., how good the solution is). Afterwards, genetic operations in the form of crossover and mutation events (see below) are applied to the current population to generate a set of children. These children are once again evaluated using the fitness function. To ensure the continuous improvement of the solutions present in a population, selection pressure is applied during the crossover procedure. This pressure is exerted by introducing a selection scheme based on the fitness score, where individuals possessing a better score and thus representing better solutions are selected more frequently for crossovers. Finally, a new population is created from the members of the parent population and the children. Similar to the crossover selection, individuals compete with each other at this point. The fitter individuals are more likely to be chosen for the next population while the most unfit individuals are exterminated. This process of evaluation, selection and reproduction is iterated until a solution of sufficient quality is obtained.
While the general motifs outlined above are common to all genetic algorithms, real-world applications typically vary in the details of implementation—such as the encoding of the genome, the fitness function used and the form of the genetic and selection operators—as they are adapted to best suit the optimization problem at hand. The same holds true for the current work. To make genetic algorithms suitable for the evolution of density functional methods, several adaptions are required, which will be discussed below.
Steady-state genetic algorithms
Instead of the standard genetic algorithm, where new populations are determined repeatedly, a steady-state genetic algorithm is used in this work. Here, only one population is maintained for the whole run and individuals are continuously generated, evaluated and replaced. The steady-state genetic algorithm is advantageous for parallel computing, as the fitness function of each individual is calculated on a separate computer core. Since, in a standard genetic algorithm, the next population is only generated once the fitness assessment of all individuals has finished (different individuals may need different computational time), many of the cores might run idle. This problem is avoided in a steady-state genetic algorithm.
Selection
Two different modes of selection are used in the steady-state genetic algorithm. Parent genomes for crossover operations are determined using tournament selection [12]. In tournament selection, N individuals are chosen from the population at random and their fitness scores are compared. The fittest individual from this subgroup is then selected. The size of the tournament N can be used to control the selection pressure.
The individuals to be deleted from the population are chosen via elitist random selection, i.e., an individual is chosen from the population at random. If it does not belong to the M fittest members of the population, it is removed and replaced by one of the children.
Fitness function
The fitness score in the present work is obtained as the weighted total mean absolute deviation (WTMAD) computed for the GMTKN30 (general main group thermochemistry, kinetics, and noncovalent interactions) reference database as defined by Grimme [11]. The GMTKN30 database encompasses 30 subsets addressing different problems, such as basic properties, reaction energies, and long-range interactions and has been used extensively to study the quality of density functionals. To obtain one single WTMAD score for a functional, a total of 1218 single point computations are performed. The weighting scheme was introduced to account for the varying sizes of the different reference subsets and the relative difficulty for DFT methods to describe these subsets. A more detailed description of the WTMAD score and the subsets used in the GMTKN30 database can be found in Ref. [11]. Since the WTMAD is an error, lower scores correspond to fitter individuals. A potential alternative to GMTKN30 is the recently published GMTKN55 [13] database, which features an improved selection of reference systems. Since the main focus of this study is the evaluation of the GA approach in general, the smaller GMTKN30 database was chosen over GMTKN55 for two reasons: (1) due to the smaller size of the former, fitness evaluations can be carried out faster while still retaining a well-balanced set of systems and (2) GMTKN30 facilitates a comparison to conventional functionals, as more in-depth studies are available due to its earlier publication date.
The density functional genome
In the present work, two different versions of genomes are required since two genera of functionals are investigated using the genetic algorithm: GGA functionals, which depend only on the gradient of the electron density, and hybrid functionals, which also incorporate part of the exact Hartree–Fock exchange.
Within the framework of Kohn–Sham DFT, the exchange correlation energy \( E_{GGA}^{XC} \) of a typical GGA functional is composed of several terms: the local Slater exchange \( E_{LSDA}^{X} \), the gradient correction to the exchange \( E_{GGA}^{X} \), the local part of the correlation energy \( E_{LSDA}^{C} \), and the gradient correction to the correlation energy \( E_{GGA}^{C} \). These different energy contributions are in turn modeled by individual functionals. While an analytical expression can be derived for the functional component \( F_{LSDA}^{X} \) describing the energy contribution \( E_{LSDA}^{X} \) based on the uniform electron gas [14, 15], only empirical approximations exist for \( F_{GGA}^{X} \), \( F_{LSDA}^{C} \), and \( F_{GGA}^{C} \) [4]. These three latter functional approximations are the building blocks that differ among the GGA functionals, so that the GGA genome therefore takes the form shown in the upper half of Fig. 1. Here, every GGA functional is encoded by three string entries—one for every functional component.
Basic genomes used for GGA and hybrid functionals. The GGA (e.g, BP86) is represented by three entries specifying the approximate functionals used in the description of \( E_{GGA}^{X} \), \( E_{LSDA}^{C} \), and \( E_{GGA}^{C} \). The genome of the hybrid functional (e.g, B3PW91) has a similar structure but is extended by a set of real numbers controlling the admixture of exact exchange (a) and the scaling of the gradient corrections to the exchange and correlation energy (b, c)
The exchange–correlation energy \( E_{\text{hybrid}}^{XC} \) of hybrid functionals is typically modeled according to
where \( E_{HF}^{X} \) is the exact Hartree–Fock exchange, a is a parameter controlling the admixture of exact exchange and b and c are scaling factors to adjust the gradient corrections to the exchange and correlation energies [4]. The genetic representation used for hybrid functionals is based on the GGA genome but also includes the scaling factors, which are encoded as real numbers (bottom half of Fig. 1).
In addition to the general composition of the functional, the influence of dispersion corrections is studied in this work. For this purpose, the basic functionals are augmented by an empirical dispersion correction during the genetic algorithm optimization procedure, using either the atom-pairwise D3 dispersion correction with the Becke–Johnson damping scheme (D3(BJ)) by Grimme or the density-dependent non-local (NL) part of the VV10 functional of Vydrov and Van Voorhis [16, 18]. Similar to the switch from GGA genomes to hybrid genomes, the basic genome is expanded by appending the fitting parameters of the dispersion correction. In the case of D3(BJ), all 4 parameters (s6, s8, a1, a2; for nomenclature and explanation of the parameters, see Refs. [16], [17]) are included, while for NL only the short-range attenuation parameter (bNL; see Ref. [18]) is optimized.
In the present work, the allowed approximations for the functional specification in the genome were B88 [19], PW91(X) [20], mPW(X) [21], PBE(X) [22], RPBE(X) [23], OPTX [24], X [25], TPSS(X) [26], B97-D(X) [27], and B97(X) [28] for F X GGA , VWN-III [29], VWN-V [29], and PW91(LDA) [20] for the F C LSDA term, and P86 [30], PW91(C) [20], PBE(C) [22], LYP [31], TPSS(C) [26], B97-D(C) [27], and B97(C) [28] for F C GGA .
It should be noted at this point, that in general, arbitrary combinations of functional components could be used to form new expressions (e.g., a combination of three different \( F_{GGA}^{X} \) and only one \( F_{LSDA}^{C} \) using different coefficients). Here, we restrict ourselves to the above functional forms as conventional DFT code generally does not allow more complex combinations. The few codes which do, proved to be too unstable with respect to the convergence of the self-consistent field procedure to be of any practical use, as the genetic algorithm requires a certain level of robustness.
Genetic operations
The search space is explored by the genetic algorithm using crossover and mutation operations.
During crossover, the genomes of two parent individuals are recombined to yield two children. This is done by applying crossover operations to the individual entries of the aligned parent genomes with the crossover probability Pc. Since the density functional genomes can contain discrete strings as well as real numbers, two different basic crossover operations have to be introduced (Fig. 2).
Crossover between two sample genomes for hybrid functionals, where the first sites \( \left( {F_{GGA}^{X} } \right) \) and the fifth sites (b) have been marked for a crossover event. Entries corresponding to functional approximations are simply swapped between genomes, while a weighted average is formed for the real-valued parts of the genome (e.g, ω = 0.75). Changes in the genomes are highlighted in red (color figure online)
If the affected entries specify the type of functional approximation, they are simply swapped between the genomes. If the entries are real numbers, such as used in the hybrid genome and for the dispersion correction, weighted averages are formed. The new value for the first child is obtained by the relation \( w \times x_{\text{parent1}} + (1 - w) \times x_{{{\text{parent}}2}} \), where the x are the respective values in the parent genomes and w is a random number drawn from the standard uniform distribution. The entry for the second child is calculated by swapping the order of the parent genomes in the weighted average. To avoid the generation of clones, at least one crossover event is enforced.
Mutation introduces random permutations to a single genome. Similar to crossover, individual entries of a genome are mutated with a certain probability, the mutation probability Pm. If a mutation occurs in a functional component, the original functional approximation is substituted by a randomly chosen approximation from the same class (e.g., only from the \( F_{GGA}^{C} \) approximations). If the entry corresponds to a real parameter, it is perturbed by Gaussian noise. An example for the two different cases is shown in Fig. 3.
Mutation of a hybrid functional genome. The \( F_{LSDA}^{C} \) entry is mutated by replacing the functional approximation by a randomly chosen approximation of the same type, while the real parameter a is mutated by adding Gaussian noise. Changes in the genomes are highlighted in red (color figure online)
At least one mutation event is enforced to ensure the genetic diversity of the population.
Results and discussion
GGAs
Three separate genetic algorithm optimization runs were carried out for GGA type functionals, using either no dispersion correction in the genome, the D3(BJ) atom-pairwise potential by Grimme (D3), or the density-dependent NL correction by Vydrov and Van Voorhis. The compositions and parameters obtained for the top performing Evolutionary GGA (EG) functionals in each species—labeled EG, EG-D3, and EG-NL—are given in Table 1. The WTMAD scores of the evolved functionals computed with the quadruple-ζ basis set are shown in Fig. 4. This figure also includes WTMAD scores computed with the same basis set of the closely related PBE and B97-D3 GGA functionals as computed in this work.
Comparing the functionals yielded by the genetic algorithm amongst each other, the following trend can be observed. The functionals which incorporate dispersion correction parameters in their genome during the genetic algorithm optimization process (EG-D3 and EG-NL) exhibit significantly lower WTMAD scores (18.41 kJ/mol for both functionals) than the functional evolved without dispersion correction (EG with 24.27 kJ/mol). This behavior is hardly surprising. The inability of standard DFT to properly describe dispersion type interactions has been the subject of several recent studies and different empirical corrections have been developed to counteract this shortcoming, such as the D3(BJ) correction and the NL correction. Since the GMTKN30 reference database used in this work contains many test systems where dispersion effects are important, it can be expected that evolved functionals with dispersion corrections perform better. No difference in accuracy is observed between the D3(BJ) correction and the NL correction for the GMTKN30 database.
Pertaining to the composition of the various GGA functionals, we find that the genetic algorithm is able to recover several conventional functionals that are known to show excellent performance for the GMTKN30 database. The EG functional without dispersion correction essentially recovers the popular PBE functional [22] (see Table 1). Although EG uses the PW91(C) correlation functional, this correlation functional is almost identical to PBE(C) with exception of one additional term and is expected to exhibit almost exactly the same performance. This is indeed the case in the present work, as can be seen when comparing the WTMADs of both functionals (24.27 kJ/mol for EG versus 24.69 kJ/mol for standard PBE). Similar results are found for the functionals using dispersion correction (EG-D3 and EG-NL). In both cases, Becke’s B97-D functional [27] is recovered, one time using D3 dispersion correction and the other time using the NL correction, showing the same performance of the original B97-D3 functional (18.41 kJ/mol in all cases). Since B97-D and its variants are amongst the most reliable GGA functionals incorporating long-range dispersion interactions, it is of little surprise that the genetic algorithm correctly identifies them as one of the top performers on the GMTKN30 database. While the above findings highlight the saliency of the genetic algorithm approach, the search for GGAs should only be regarded as a proof of principle study. Since the valid GGA genomes were only drawn from ten GGA exchange functionals (including one meta-GGA), three local correlation functionals, and nine GGA correlation functionals (including one meta-GGA), the total number of possible combinations is 210 functionals, which could also be explored in a more systematic manner. In the case of the GGAs incorporating dispersion, the genetic algorithm explores a slightly larger search space, since the long-range parameters are determined at the same time as the functional composition. This approach is unusual, as typically dispersion corrections are parametrized in an a posteriori manner. However, no difference in the performance is observed in the present work (see, e.g., EG-D3 and B97-D3).
Hybrids
Similar to the GGA functionals reported above, three genetic algorithm evolution runs have been carried out to identify well-performing Evolutionary Hybrid (EH) functionals, using once again genomes with no dispersion correction, as well as the D3(BJ) and NL corrections (a complete genealogy of the EH-NL functionals can be found in the supporting information, which illustrates the work of the genetic algorithm). The WTMAD scores of the best resulting functionals—termed EH, EH-D3, and EH-NL—can be found in Fig. 5. Their compositions as well as hybrid and dispersion parameters are given in Table 1.

taken from Ref. [32] (color figure online)
WTMAD scores (in kJ/mol) of the evolved hybrid functionals EH, EH-D3 and EH-NL (shown in orange) and several common hybrid functionals (blue) for comparison. WTMAD values for B97, B3PW91-D3(BJ), and B3PW91 were computed in this work, while the WTMAD of ωB97X-D3 was
As expected, all evolved hybrid functionals yield lower WTMAD scores than their GGA counterparts. The benefit of including a dispersion correction on the overall WTMAD scores is also observed. However, in contrast to the GGA functionals, where the D3(BJ) and NL dispersion corrections perform equally well, NL outperforms D3(BJ) in the case of hybrid functionals. The trend obtained for the evolved GGAs suggests that this difference is mainly due to the larger parameter space of D3(BJ) (four additional parameters) compared to NL (one additional parameter). The already enlarged genome of hybrid functionals in combination with the additional parameters introduced by the D3(BJ) correction complicates the search for the global optimum and, as a result, the genetic algorithm is terminated before complete convergence is reached. Further iterations until convergence were not carried out to ensure comparability with the other genetic algorithm runs (EH-NL, EH).
The hybrid functional EH without explicit dispersion correction is closely related to the one parameter hybrid B97 of Becke [33]. The main differences between both hybrids are the change of the B97(X) exchange term for B88(X) and the introduction of two additional parameters present in the standard three parameter hybrid form employed above (b and c). Compared to the standard B97 functional (WTMAD of 20.50 kJ/mol), the variant EH (WTMAD of 17.99 kJ/mol) shows better overall performance on the GMTKN30 database. The reason for this behavior is a combination of two effects: First, the functional form of EH is more flexible due to the two additional parameters. Second, EH is directly optimized on GMTKN30, while B97 was parametrized on a different set of molecules. The primary difference between both sets is the presence of a wide range of non-covalent interaction benchmarks in GMTKN30, which contribute to the overall WTMAD with a high weight. Based on the information contained in these benchmarks, the genetic algorithm utilizes the additional flexibility of EH to introduce a dispersion-like behavior (see Fig. 6). Consequently, the WTMAD of EH associated with non-covalent interactions (10.04 kJ/mol) is much lower than in B97 (16.74 kJ/mol), leading to the improved WTMAD score. Considering only basic properties and reaction energies, both hybrids exhibit a much closer performance, with B97 achieving slightly better accuracy for basic properties (19.66 kJ/mol vs. 20.50 kJ/mol) and EH for the reaction test set (24.27 kJ/mol vs. 22.18 kJ/mol). This trend is to be expected, as B97 was parametrized systematically in order to provide a good performance over a wide range of model chemistries. At the same time, this finding demonstrates the power of the genetic algorithm search procedure, as it is able to utilize information in the reference data in a manner contrasting to conventional parametrization strategies. Whether this use of information is physically founded or not remains to be addressed in future investigations. The composition of EH-D3 is similar to EH, but includes explicit dispersion correction. A direct comparison of this functional to B97 with D3(BJ) dispersion correction is not possible, as no D3(BJ) parameters have been reported for the B97 hybrid. However, due to the above trends, it is expected that both functionals would exhibit a similar performance, as non-covalent interactions are now accounted for in an explicit manner in both cases. An important observation related to EH-D3 is that while the genetic algorithm is able to identify a sufficiently good solution, other conventional functionals with lower WTMADs are in principle accessible but not found during optimization (e.g., B3PW91-D3, see Fig. 6). This failure to identify the minimum corresponding to B3PW91-D3 can be once again attributed to the expanded parameter space introduced by the D3(BJ) correction, which slows down the convergence of the genetic algorithm (see above). The final hybrid functional EH-NL is indeed a reparametrized version of B3PW91 [34] using the NL dispersion correction. However, in this case, the genetic algorithm is able to identify an improved set of parameters and EH-NL shows a significantly lower WTMAD than both its D3 and NL counterparts (11.30 kJ/mol vs. 12.55 kJ/mol and 14.23 kJ/mol, respectively). Unlike in the case of EH, this gain in performance is not achieved using the extra flexibility of the hybrid to introduce artificial dispersion behavior. Instead, ED-NL primarily improves upon the other B3PW91 versions in the basic properties (19.66 kJ/mol vs. 20.50 kJ/mol for B3PW91-D3 and 23.85 kJ/mol for B3PW91-NL) and reaction energies benchmarks (8.79 kJ/mol vs. 11.30 kJ/mol and 12.55 kJ/mol), while exhibiting nearly the same performance for non-covalent interactions (3.35 kJ/mol vs. 3.78 kJ/mol and 3.35 kJ/mol). Especially the accuracy obtained for reaction energies is remarkable. On the whole, EH-NL shows excellent performance across the entire GMTKN30 dataset even when compared to such successful functionals as ωB97X-D [35] (WTMAD of 11.72 kJ/mol). The close relation between the hybrid functionals found in this work to their conventional counterparts also serves as a general demonstration for the excellent performance of the latter for a wide range of chemical systems.
Potential energy curves for the benzene dimer as computed for the functionals EH-NL, EH, EH-ED with the def2-QZVP basis set. The CCSD(T) curve (shown in black) is taken from Ref. [40]. In addition, the curve computed for the B97 hybrid functional is shown in gray for comparison
An ongoing discussion in the DFT community concerns the amount of exact exchange (referred to as a in this work) used in hybrid functionals. Typical values range from 10% Hartree–Fock exchange for TPSSh [36] up to 54% for M06-2X [37], with the majority of standard hybrid functionals clustered around 25%, which is also the optimal value of exchange admixture suggested by theory [38]. Analyzing the top 100 evolved hybrid functionals of the three different species with respect to the amount of exact exchange, average values of 25, 24, and 23% are obtained if no dispersion correction, the D3(BJ) correction or the NL correction is used, respectively. This finding supports the consensus that for three parameter hybrids an admixture Hartree–Fock exchange close to 25% is the best compromise if good performance over a variety of different systems is desired [38].
Basis-set dependence
Methods that were optimized using a certain basis set can show erratic behavior when paired with other basis sets (i.e., reduced accuracy, even when a larger basis sets than the original is used). Consequently, parametrization is often carried out with a basis set of similar quality as the one intended for the accuracy assessment or subsequent practical applications. In this work, the WTMAD computed with a quadruple-ζ basis set (def2-QZVP) is used in the final comparison of the different functionals. However, carrying out all 2,582,000 single point calculations (as comprised in the GMTKN30 database) required over the course of a single genetic algorithm run would be too demanding with a basis set of this size. Hence, a smaller double-ζ basis set (def2-SVP) was chosen for the fitness evaluation of functionals, in order to render the computations required for the genetic algorithm tractable. To assess whether the use of this protocol is justified, the WTMADs of several evolved functionals are compared using basis sets of double-ζ, triple-ζ, and quadruple-ζ quality. The obtained values are reported in Table 2. Apart from the six functionals described above, another hybrid functional with NL dispersion correction, EH2-NL, was included.
Since the WTMAD scores of all functionals under investigation improve systematically with basis set size, no erratic basis set dependence seems to be present. Trends between functionals of different species (e.g., no dispersion, D3(BJ) correction) are also captured well, independent of the quality of the basis set. Potential problems can occur for functionals of the same species, which show only small differences in their WTMAD at double-ζ level, as it is possible for the relative trend between these functionals to invert upon increase of the basis set size. An example are the functionals EH-NL and EH2-NL. For the double-basis, the WTMAD of EH2-NL is lower than the one of EH-NL, although the difference is only small (0.25 kJ/mol). This ordering is inverted if a triple-ζ or larger basis set is used, since EH-NL now lies 1.80 kJ/mol below EH2-NL, with the difference being even more pronounced for the quadruple-ζ basis.
However, since this phenomenon occurs only rarely and only for very small differences in the WTMAD score, the computations at double-ζ level still provide a good general guideline for the genetic algorithm. A simple countermeasure is to select not only the best functional yielded by the genetic algorithm for evaluation at quadruple-ζ level, but also those functionals whose WTMAD scores lie within a certain limit to the best score. An advantage of this approach is that the genetic algorithm run with a double-ζ basis set and a subsequent re-evaluation of only a handful of functionals with the quadruple-ζ basis set is still far more efficient than a whole genetic algorithm run carried out with the larger basis set. All functionals reported above were identified in this manner.
Influence of RI and RIJCOSX approximations
To reduce the required computational time, use was made of the resolution of identity (RI) approximation for GGAs and the resolution of identity chain of spheres (RIJCOSX) approximation for hybrid functionals. These approximations allow for an efficient computation of Coulomb and Hartree–Fock exchange terms, respectively, and can lead to speed-ups of one order of magnitude. RI and RIJCOSX, however, introduce small errors in the electronic energies. To study the impact of these errors on the WTMAD scores, reference computations for the evolved GGA and hybrid functionals were carried out with and without RI and RIJCOSX. It was found that upon use of the RI and RIJCOSX approximations, the WTMADs increase by an average of 0.04 kJ/mol for GGA functionals and by 0.21 kJ/mol for hybrid functionals. The semi-numerical RIJCOSX yields slightly larger deviations than standard RI, but regarding the energies computed here, both approximations are safe to use and do not influence the quality of the results significantly. Hence, all WTMAD scores reported here for the evolved functionals were computed with these approximations.
Evolution for specific applications
So far, all evolved functionals presented here were obtained using the WTMAD computed over the GMTKN30 database as a performance measure. This protocol was chosen in order to study general functional patterns emerging for a chemically balanced set of problems. However, one particular strength of genetic algorithms is their flexibility with respect to the criteria to be optimized, since only the fitness function has to be adapted accordingly and no gradients of any form are required.
To test this versatility of genetic algorithms in the context of density functionals, a hybrid functional was optimized targeting only the mean absolute error on those subsets of the GMTKN30 database that focus on non-covalent interactions. Moreover, no dispersion correction was included in the functional genome. The resulting functional (see Table 1) was then applied to compute the potential energy curve for the benzene dimer, a well-known example for a Van-der-Waals bound system, where standard DFT fails (see, e.g., Ref. [39]). The potential energies obtained for the functional with adapted fitness function, called EH-ED (ED stands for evolved dispersion), the functionals EH and EH-NL, as well as CCSD(T) reference values taken from Ref. [40] are shown in Fig. 6.
While EH-NL shows only minor deviations from the CCSD(T) reference, the minimum is completely absent in the case of EH and the dimers are unbound. This result is typical for functionals not augmented by dispersion corrections, as they lack the physical capability to describe long-range interactions of the Van-der-Waals type. Yet, compared to EH, the potential curve of EH-ED exhibits a qualitatively correct behavior, possessing a distinct minimum at slightly larger distances than the CCSD(T) reference, despite the explicit absence of a dispersion correction.
This result is an excellent demonstration for the performance of the genetic algorithm in optimizing a specially designed objective. Moreover, it opens up the possibility to use the genetic algorithm to automatically create “niche”-functionals tailored to specific needs. While the increased accuracy for the target properties comes at the cost of generality (e.g., reduced performance for other properties, EH-ED exhibits a WTMAD of 48.53 kJ/mol), the associated trade-off should be acceptable compared to the gains, provided the reference data and fitness function are chosen carefully.
At the same time, this finding also illustrates one of the potential drawbacks of DFT. As was shown above, even the standard three-parameter hybrid functional form is extremely flexible with respect to the combination of different functional approximations and parametrization. With enough patience and creativity, almost any desired result can be obtained. Hence, this high intrinsic flexibility of density functional approaches should always be considered carefully when searching for a universally valid functional.
Conclusion
A genetic algorithm was applied to the exploration of the generalized gradient approximation (GGA) and hybrid density functional subspace spanned by several popular functional components. The genetic optimization of individual functionals was guided by their performance on the GMTKN30 reference database, which was also used as a measure for accuracy when comparing different functionals. For both types of functionals—GGAs and hybrids—the genetic algorithm is able to identify variants of popular density functionals, which show good performance on the GMTKN30 benchmark. These results demonstrate the ability of the genetic algorithm to efficiently explore the possible combinations of exchange and correlation functionals as well as different parametrization patterns. Several interesting effects are observed for the hybrid functionals in particular. Monitoring the admixture of exact exchange for the top performing members of each population, it is found to converge towards a numerical value close to 25%, which is commonly accepted to offer the best accuracy for a wide range of chemical systems [38]. In addition, the genetic algorithm is able to identify a reparametrized variant of B3PW91, which shows excellent performance over the whole GMTKN30 benchmark, not only outperforming conventional versions of B3PW91, but also coming close to top performing functionals, such as ωB97X-D.
An important feature of the genetic algorithm is its ability to automatically construct functionals tailored to specific problems or molecules by employing different reference data sets to guide the evolution process. The potential utility of this feature is demonstrated by introducing dispersion-like behavior in a functional that possesses no inherent dispersion correction. This automated construction of “niche”-functionals with improved accuracy for certain systems or properties will prove useful in situations, where fast and accurate computations are required and a sufficient amount of reference data is available (e.g., in ab initio molecular dynamics).
Other potential future applications for the genetic algorithm are the automated determination of parametrization patterns required for newly developed functional components, as well as the re-parametrization of existing functionals. Moreover, the use of genetic algorithms in the field of density functional theory also offers the tantalizing possibility not only to optimize parametrization schemes for functionals, but also to evolve better approximations to the exact exchange–correlation functionals via genetic programming [41].
Computational details
All computations were performed with Orca [42]. Calculations of the WTMAD required for the fitness assessment during the genetic algorithm run were carried out with the def2-SVP basis set, while the best evolved functionals were re-evaluated at the def2-QZVP level [43]. In case of the G21EA and WATER27 subsets of the GMTKN30 database, the standard basis set was augmented by diffuse functions from the aug-cc-pVDZ and aug-cc-pVQZ basis sets, respectively [44]. Scalar relativistic effects in the HEAVY28 and RG6 datasets were accounted for using the appropriate Stuttgart–Dresden effective core potentials as implemented in Orca [45, 46]. To speed up the evaluation process, the resolution of identity (RI) approximation was employed for GGA functionals and the RI-chain-of-spheres exchange (RIJCOSX) approximation for hybrid functionals [47,48,49]. Open-shell systems present in the reference data were described within the unrestricted Kohn–Sham framework.
A population of 100 individuals was used for all genetic algorithm runs. Crossover and mutation probabilities were set to Pc = 0.6 and Pm = 0.1, respectively. A total of 20 children were generated during crossover events and the parents were selected using tournaments of size 2. To ensure a diverse gene-pool, 5 completely new individuals were generated at the same time and added to the children. During the random replacement selection, the 5 best individuals were preserved. For every genetic algorithm run, a total of 2000 fitness evaluations were carried out.
Supplementary Information
ORCA inputs for all functionals, as well as a genealogy of the EH-NL species can be found in the supplementary information.
References
Kohn W, Sham LJ (1965) Phys Rev 140:A1133
Burke K (2012) J Chem Phys 136:150901
Kohn W, Becke AD, Parr RG (1996) J Phys Chem 100:12974
Jensen F (2006) Introduction to computational chemistry, 2nd edn. Wiley-Blackwell, Hoboken
Mardirossian N, Head-Gordon M (2015) J Chem Phys 142:074111
Mardirossian N, Head-Gordon M (2014) Phys Chem Chem Phys 16:9904
McAnanama-Brereton S, Waller MP (2018) J Chem Inf Model 58:61
Holland JH (1975) Adaptation in natural and artificial systems: An introductory analysis with applications to biology, control, and artificial intelligence. University of Michigan Press, Ann Arbor
Darwin C (1859) On the origin of species by means of natural selection. Murray, London
Leardi R (2001) J Chemom 15:559
Goerigk L, Grimme S (2011) J Chem Theory Comput 7:291
Goldberg DE (1989) Genetic algorithms in search, optimization, and machine learning. Addison-Wesley, Reading
Goerigk L, Hansen A, Bauer C, Ehrlich S, Najibi A, Grimme S (2017) Phys Chem Chem Phys 19:32184
Dirac PAM (1929) Proc R Soc A 123:714
Slater JC (1951) Phys Rev 81:385
Grimme S, Antony J, Ehrlich S, Krieg H (2010) J Chem Phys 132:154104
Grimme S, Ehrlich S, Goerigk L (2011) J Comput Chem 32:1456
Vydrov OA, Van Voorhis T (2010) J Chem Phys 133:244103
Becke AD (1988) Phys Rev A 38:3098
Perdew JP, Ziesche P, Eschrig H (1991) Electronic structure of solids’91. Akademie Verlag, Berlin
Adamo C, Barone V (1998) J Chem Phys 108:664
Perdew JP, Burke K, Ernzerhof M (1996) Phys Rev Lett 77:3865
Hammer B, Hansen LB, Nørskov JK (1999) Phys Rev B 59:7413
Molawi K, Cohen AJ, Handy NC (2002) Int J Quantum Chem 89:86
Xu X, Goddard WA (2004) Proc Natl Acad Sci USA 101:2673
Tao J, Perdew JP, Staroverov VN, Scuseria GE (2003) Phys Rev Lett 91:146401
Grimme S (2006) J Comput Chem 27:1787
Becke AD (1997) J Chem Phys 107:8554
Vosko SH, Wilk L, Nusair M (1980) Can J Phys 58:1200
Perdew JP (1986) Phys Rev B 33:8822
Lee C, Yang W, Parr RG (1988) Phys Rev B 37:785
Goerigk L, Grimmer S (2011) Phys Chem Chem Phys 13:6670
Becke AD (1997) J Chem Phys 107:8554
Becke AD (1993) J Chem Phys 98:5648
Chai J-D, Head-Gordon M (2008) Phys Chem Chem Phys 10:6615
Staroverov VN, Scuseria GE, Tao J, Perdew JP (2003) J Chem Phys 119:12129
Zhao Y, Truhlar DG (2007) Theor Chem Acc 120:215
Ernzerhof M (1998) In: Joubert D (ed), Density functionals: theory and applications. Proceedings of the Tenth Chris Engelbrecht Summer School in Theoretical Physics, Meerensee near Cape Town South Africa, 19–29 January 1997. Springer, Berlin, p 60
Grimme S (2011) WIREs Comput Mol Sci 1:211
Sherrill CD, Takatani T, Hohenstein EG (2009) J Phys Chem A 113:10146
Koza JR (1992) Genetic programming: On the programming of computers by means of natural selection. MIT Press, Cambridge
Neese F (2012) WIREs Comput Mol Sci 2:73
Weigend F, Ahlrichs R (2005) Phys Chem Chem Phys 7:3297
Kendall RA, Dunning TH, Harrison RJ (1992) J Chem Phys 96:6796
Metz B, Stoll H, Dolg M (2000) J Chem Phys 113:2563
Peterson KA, Figgen D, Goll E, Stoll H, Dolg M (2003) J Chem Phys 119:11113
Eichkorn K, Treutler O, Öhm H, Häser M, Ahlrichs R (1995) Chem Phys Lett 240:283
Neese F, Wennmohs F, Hansen A, Becker U (2009) Chem Phys 356:98
Vahtras O, Almlöf J, Feyereisen MW (1993) Chem Phys Lett 213:514
Acknowledgements
Open access funding provided by University of Vienna. Allocation of computer time at the Vienna Scientific Cluster (VSC) is gratefully acknowledged. M.G. thanks the Monatshefte für Chemie for financial support via the MoChem scholarship. M.G. thanks the mysterious referee of March 2016 on a previous version of this work for his invaluable feedback.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Gastegger, M., González, L. & Marquetand, P. Exploring density functional subspaces with genetic algorithms. Monatsh Chem 150, 173–182 (2019). https://doi.org/10.1007/s00706-018-2335-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00706-018-2335-3