Abstract
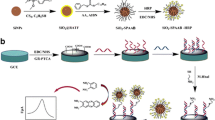
A nanocomposite was prepared from β-cyclodextrin (β-CD) functionalized graphene oxide and magnetic nanoparticles (GO/Fe3O4/β-CD). In parallel, a polyamidoamine dendrimer conjugated to avidinylated alkaline phosphatase (PAMAM-avidin-ALP) was prepared and exploited as a signal amplification unit in a voltammetric immunoassay for 5-methylcytosine (5mC) in genomic DNA. The GO/Fe3O4/β-CD as a substrate material exhibited good solubility, electrical conductivity and large surface. This is beneficial for the further modification of antibodies (Ab) by host-guest interaction and amide bonds. By taking advantage of three-dimensional structure to capture avidin-ALP by amide linkages, PAMAM was used as a catalytic signal amplification element in this assay. Under the optimized condition and at a typical working potential of 0.94 V, the response to 5mC is linear in the 0.01–50 nM concentration range with a detection limit of 3.2 pM (at S/N = 3). The method is stable, selective and reproducible. It was applied to the determination of 5mC in genomic DNA of human tissue.

An electrochemical immunoassay was constructed for 5-methylcytosine detection based on nanocomposite of graphene oxide, magnetite nanoparticles and β-cyclodextrin, and enzymatic signal amplification.





Similar content being viewed by others
References
Bird A (2002) DNA methylation patterns and epigenetic memory. Genes Dev 16:6–21. https://doi.org/10.1101/gad.947102
Frigola J, Song J, Stirzaker C, Hinshelwood RA, Peinado MA, Clark SJ (2006) Epigenetic remodeling in colorectal cancer results in coordinate gene suppression across an entire chromosome band. Nat Genet 38:540–549. https://doi.org/10.1038/ng1781
Robertson KD (2005) DNA methylation and human disease. Nat Rev Genet 6:597–610. https://doi.org/10.1038/nrg1655
Yan P, Hao Y, Shu Z, Gu C, Zhou X, Liu X, Xiang H (2018) Double signal enhancement strategy based on rolling circle amplification and photoinduced electron transfer for ultrasensitive fluorometric detection of methylated DNA. Microchim Acta 185:299. https://doi.org/10.1007/s00604-018-2839-x
Su F, Wang L, Sun Y, Liu C, Duan X, Li Z (2015) Highly sensitive detection of CpG methylation in genomic DNA by AuNP-based colorimetric assay with ligase chain reaction. Chem Commun 51:3371–3374. https://doi.org/10.1039/c4cc07688e
Liutkevičiūtė Z, Lukinavičius G, Masevičius V, Daujotytė D, Klimaauskas S (2009) Cytosine-5-methyltransferases add aldehydes to DNA. Nat Chem Biol 5:400–402. https://doi.org/10.1038/nchembio.172
Bi S, Zhao T, Luo B, Zhu JJ (2013) Hybridization chain reaction-based branched rolling circle amplification for chemiluminescence detection of DNA methylation. Chem Commun 49:6906–6908. https://doi.org/10.1039/c3cc43353f
Liu X, Wei C, Luo J, Wu Y, Guo X, Ying Y, Wen Y, Yang H (2018) Photoelectrochemical determination of the activity of M.SssI methyltransferase, and a method for inhibitor screening. Microchim Acta 185:498. https://doi.org/10.1007/s00604-018-3033-x
Wang Y, He X, Wang K, Su J, Chen Z, Yan G, Du Y (2013) A label-free electrochemical assay for methyltransferase activity detection based on the controllable assembly of single wall carbon nanotubes. Biosens Bioelectron 41:238–243. https://doi.org/10.1016/j.bios.2012.08.034
Bhattacharjee R, Moriam S, Nguyen N-T, Shiddiky MJA (2019) A bisulfite treatment and PCR-free global DNA methylation detection method using electrochemical enzymatic signal engagement. Biosens Bioelectron 126:102–107. https://doi.org/10.1016/j.bios.2018.10.020
Ren H, Cheyne CG, Fleming AM, Burrows CJ, White HS (2018) Single-molecule titration in a protein nanoreactor reveals the protonation/deprotonation mechanism of a C:C mismatch in DNA. J Am Chem Soc 140:5153–5160. https://doi.org/10.1021/jacs.8b00593
Liu H, Weng L, Yang C (2017) A review on nanomaterial-based electrochemical sensors for H2O2, H2S and NO inside cells or released by cells. Microchim Acta 184:1267–1283. https://doi.org/10.1007/s00604-017-2179-2
Rasheed PA, Sandhyarani N (2017) Electrochemical DNA sensors based on the use of gold nanoparticles: a review on recent developments. Microchim Acta 184:981–1000. https://doi.org/10.1007/s00604-017-2143-1
Zhang Z, Sheng S, Cao X, Li Y, Yao J, Wang T, Xie G (2015) Proximity-based electrochemical biosensor for highly sensitive determination of methyltransferase activity using gold nanoparticle-based cooperative signal amplification. Microchim Acta 182:2329–2336. https://doi.org/10.1007/s00604-015-1564-y
Gao F, Fan T, Ou S, Wu J, Zhang X, Luo J, Li N, Yao Y, Mou Y, Liao X, Geng D (2018) Highly efficient electrochemical sensing platform for sensitive detection DNA methylation, and methyltransferase activity based on Ag NPs decorated carbon nanocubes. Biosens Bioelectron 99:201–208. https://doi.org/10.1016/j.bios.2017.07.063
Liu P, Wang D, Zhou Y, Wang H, Yin H, Ai S (2016) DNA methyltransferase detection based on digestion triggering the combination of poly adenine DNA with gold nanoparticles. Biosens Bioelectron 80:74–78. https://doi.org/10.1016/j.bios.2015.12.100
Yang Z, Wang F, Wang M, Yin H, Ai S (2015) A novel signal-on strategy for M.SssI methyltransfease activity analysis and inhibitor screening based on photoelectrochemical immunosensor. Biosens Bioelectron 66:109–114. https://doi.org/10.1016/j.bios.2014.11.015
Zhou Y, Xu Z, Wang M, Sun B, Yin H, Ai S (2014) DNA methyltransferase activity assay based on visible light-activated photoelectrochemical biosensor. Biosens Bioelectron 53:263–267. https://doi.org/10.1016/j.bios.2013.09.065
Wang M, Xu Z, Chen L, Yin H, Ai S (2012) An electrochemical immunosensing platform for DNA methyltransferase activity analysis and inhibitor screening. Anal Chem 84:9072–9078. https://doi.org/10.1021/ac301620m
Roy E, Patra S, Kumar D, Madhuri R, Sharma PK (2015) Multifunctional magnetic reduced graphene oxide dendrites: synthesis, characterization and their applications. Biosens Bioelectron 68:726–735. https://doi.org/10.1016/j.bios.2015.01.072
Liang RP, Liu CM, Meng XY, Wang JW, Qiu JD (2012) A novel open-tubular capillary electrochromatography using β-cyclodextrin functionalized graphene oxide-magnetic nanocomposites as tunable stationary phase. J Chromatogr A 1266:95–102. https://doi.org/10.1016/j.chroma.2012.09.101
Song J, Yang C, Ma J, Han Q, Ran P, Fu Y (2018) Voltammetric chiral discrimination of tryptophan using a multilayer nanocomposite with implemented amino-modified β-cyclodextrin as recognition element. Microchim Acta 185:230. https://doi.org/10.1007/s00604-018-2765-y
Niu X, Mo Z, Yang X, Sun M, Zhao P, Li Z, Ouyang M, Liu Z, Gao H, Guo R, Liu N (2018) Advances in the use of functional composites of β-cyclodextrin in electrochemical sensors. Microchim Acta 185:328. https://doi.org/10.1007/s00604-018-2859-6
Peng Y, Shen H, Tang S, Huang Z, Hao Y, Luo Z, Zhou F, Wang T, Feng W (2018) Colorimetric determination of BCR/ABL fusion genes using a nanocomposite consisting of Au@Pt nanoparticles covered with a PAMAM dendrimer and acting as a peroxidase mimic. Microchim Acta 185:401. https://doi.org/10.1007/s00604-018-2940-1
Hu L, Dong T, Zhao K, Deng A, Li J (2018) Ultrasensitive electrochemiluminescent brombuterol immunoassay by applying a multiple signal amplification strategy based on a PAMAM-gold nanoparticle conjugate as the bioprobe and Ag@Au core shell nanoparticles as a substrate. Microchim Acta 184:3415–3423. https://doi.org/10.1007/s00604-017-2359-0
Kavosi B, Salimi A, Hallaj R, Amani K (2014) A highly sensitive prostate-specific antigen immunosensor based on gold nanoparticles/PAMAM dendrimer loaded on MWCNTS/chitosan/ionic liquid nanocomposite. Biosens Bioelectron 52:20–28. https://doi.org/10.1016/j.bios.2013.08.012
Wang Y-Z, Hao N, Feng Q-M, Shi H-W, Xu J-J, Chen H-Y (2016) A ratiometric electrochemiluminescence detection for cancer cells using g-C3N4 nanosheets and Ag–PAMAM–luminol nanocomposites. Biosens Bioelectron 77:76–82. https://doi.org/10.1016/j.bios.2015.08.057
Zhang X, Shen J, Ma H, Jiang Y, Huang C, Han E, Yao B, He Y (2016) Optimized dendrimer-encapsulated gold nanoparticles and enhanced carbon nanotube nanoprobes for amplified electrochemical immunoassay of E. coli in dairy product based on enzymatically induced deposition of polyaniline. Biosens Bioelectron 80:666–673. https://doi.org/10.1016/j.bios.2016.02.043
Gao J, Guo Z, Su F, Liang G, Pang X, Cao W, Du B, Wei Q (2015) Ultrasensitive electrochemical immunoassay for CEA through host–guest interaction of β-cyclodextrin functionalized graphene and Cu@Ag core–shell nanoparticles with adamantine-modified antibody. Biosens Bioelectron 63:465–471. https://doi.org/10.1016/j.bios.2014.07.081
Yin H, Sun B, Zhou Y, Wang M, Xu Z, Fu Z, Ai S (2014) A new strategy for methylated DNA detection based on photoelectrochemical immunosensor using Bi2S3 nanorods, methyl bonding domain protein and anti-his tag antibody. Biosens Bioelectron 51:103–108. https://doi.org/10.1016/j.bios.2013.07.040
Cao A, C-y Z (2012) Sensitive and label-free DNA methylation detection by ligation-mediated hyperbranched rolling circle amplification. Anal Chem 84:6199–6205. https://doi.org/10.1021/ac301186j
Ellison EM, Abner EL, Lovell MA (2017) Multiregional analysis of global 5-methylcytosine and 5-hydroxymethylcytosine throughout the progression of Alzheimer's disease. J Neurochem 140:383–394. https://doi.org/10.1111/jnc.13912
Zeng T, Liu L, Li T, Li Y, Gao J, Zhao Y, Wu H-C (2015) Detection of 5-methylcytosine and 5-hydroxymethylcytosine in DNA via host–guest interactions inside α-hemolysin nanopores. Chem Sci 6:5628–5634. https://doi.org/10.1039/c5sc01436k
Acknowledgements
This work was supported by the National Natural Science Foundation of China (No. 21775090).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
The author(s) declare that they have no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
ESM 1
(DOC 144 kb)
Rights and permissions
About this article
Cite this article
Zhou, Y., Jiang, W., Wu, H. et al. Amplified electrochemical immunoassay for 5-methylcytosine using a nanocomposite prepared from graphene oxide, magnetite nanoparticles and β-cyclodextrin. Microchim Acta 186, 488 (2019). https://doi.org/10.1007/s00604-019-3575-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00604-019-3575-6