Abstract
Aims
The purpose of the current study was to investigate the characteristic expression of circular RNAs (circRNAs) in the peripheral blood of type 2 diabetes mellitus (T2DM) patients and their potential as diagnostic biomarkers for pre-diabetes and T2DM.
Methods
CircRNAs in the peripheral blood from six healthy individuals and six T2DM patients were collected for microarray analysis, and an independent cohort study consisting of 20 normal cases, 20 pre-diabetes patients and 20 T2DM patients was conducted to verify the five chosen circRNAs. We then tested hsa_circ_0054633 in a third cohort (control group, n = 60; pre-diabetes group, n = 63; and T2DM group, n = 64) by quantitative real-time polymerase chain reaction (Q-PCR).
Results
In total, 489 circRNAs were discovered to be differentially expressed between the two groups, and of these, 78 were upregulated and 411 were downregulated in the T2DM group. Five circRNAs were then selected as candidate biomarkers and further verified in a second cohort. Hsa_circ_0054633 was found to have the largest area under the curve (AUC). The diagnostic capacity of hsa_circ_0054633 was tested in a third cohort. After introducing the risk factors of T2DM, the hsa_circ_0054633 AUCs for the diagnosis of pre-diabetes and T2DM slightly increased from 0.751 (95% confidence interval [0.666–0.835], P < 0.001) to 0.841 ([0.773–0.910], P < 0.001) and from 0.793 ([0.716–0.871], P < 0.001) to 0.834 ([0.762–0.905], P < 0.001), respectively.
Conclusions
Hsa_circ_0054633 presented a certain diagnostic capability for pre-diabetes and T2DM.
Similar content being viewed by others
Introduction
According to the International Diabetes Federation (IDF) Diabetes Atlas (Seventh Edition, 2015), nearly 410 million diabetic patients exist worldwide, 46.5% of whom have not been diagnosed. By 2040, the number of patients with diabetes may increase to 642 million [1]. In the advanced stages of T2DM, patients often experience various complications. Therefore, early diagnosis and intervention are urgently needed. However, current diagnostic methods show various insufficiencies for the early diagnosis of T2DM. To improve this situation, researchers have assessed the value of insulin resistance, β-cell function, insulin sensitivity and fasting insulin for the diagnosis of T2DM using the homeostasis model [2]. Some researchers have attempted to identify new indicators of T2DM at early stages, and others have explored the association between genetic variants and early-onset T2DM [3, 4], a new and highly sensitive biomarker that will be of great value.
In recent years, with advancements in genomics, the single-nucleotide polymorphism (SNP) sites of related encoding sequences of some complicated diseases, including T2DM, have been gradually uncovered [5, 6]. Researchers have found that the human genome can be widely transcribed into a large number of non-coding RNAs that are closely linked to the occurrence and development of diseases [7]. CircRNAs are a type of closed circular non-coding RNAs, formed by an exon, an intron, or the reverse splicing of the two [8, 9]. Intracellular circRNAs have higher biological stability than most linear RNAs because of their resistance to RNA exonucleases [10, 11]. CircRNAs have multiple regulatory mechanisms of gene expression [12]: Some circRNAs can be used as microRNA (miRNA) sponges, playing a role in posttranscriptional regulation by engaging in competitive combination with miRNA [13]. CircRNAs can also regulate transcription by interacting with small nuclear RNA (snRNA) or RNA polymerase II in the nucleus [14] and can competitively regulate RNA splicing by binding to transcription factors [15]. Substantial amounts of circRNAs are widely distributed in the cytoplasm and nucleus [16].
CircRNAs play important roles in various diseases, including cancer, atherosclerosis, osteoarthritis, pulmonary fibrosis, myotonic dystrophy and Alzheimer’s disease [17–20]. The high biological stability of circRNAs is a precondition for their usage as biomarkers for various diseases. For example, Li et al. [21] found that hsa_circ_002059 could be used as a new biomarker for gastric cancer, Zhang et al. [22] determined that circ-ITCH could be used for the clinical diagnosis of esophageal cancer, and Qin et al. [23] discovered that hsa_circ_0005075 could be used as a potential biomarker for hepatocellular carcinoma. Regarding diabetes, cerebellar degeneration-related autoantigen 1 (CDR1) is a protein coding gene located in Xq27.1. As the natural antisense transcripts of CDR1, circRNA-CDR1 has been found to affect insulin secretion and β cell renewal [24]. In this study, we compared the expression profile of circRNAs in the peripheral blood of T2DM patients and matched control subjects by microarray analysis and then confirmed our findings in larger independent cohorts. The results demonstrated that hsa_circ_0054633 is a sensitive and specific biomarker for pre-diabetes and T2DM diagnosis.
Materials and methods
Study population
In this study, a total of 259 individuals were classified into three cohorts (their clinical and demographic characteristics are presented in Supplementary Tables 1–3). The participants were enrolled from among the outpatients and inpatients of the cardiology and endocrinology departments of the People’s Hospital of Zhengzhou University from July 2015 to June 2016. Subjects with any of the following characteristics were excluded: (i) malignancies, (ii) liver and kidney dysfunction, (iii) any other clinically systemic acute or chronic inflammatory disease(s), (iv) autoimmune disease, (v) untreated hypertension, and (vi) any endocrine disease other than T2DM.
Study process
The process of this study is shown in Fig. 1 Each study subject was tested using the oral glucose tolerance test (OGTT) to determine whether they were healthy or had pre-diabetes or T2DM. Six control individuals and six T2DM patients were selected to donate venous blood samples, and total RNA was extracted for microarray analysis. The screened circRNAs were then validated in an independent cohort (control group, n = 20; pre-diabetes group, n = 20; and T2DM group, n = 20). After receiver operating characteristic (ROC) curve analysis, the circRNA with the best diagnostic value was selected as the biomarker, and its diagnostic value was validated in another independent cohort (control group, n = 60; pre-diabetes group, n = 63; and T2DM group, n = 64).
The definitions of pre-diabetes and T2DM and collection of whole blood samples
In this study, pre-diabetes and T2DM were diagnosed according to the 1998 Standards of the World Health Organization (WHO) [25]. Thus, patients meeting either of the following criteria could be diagnosed as having T2DM: (i) fasting plasma glucose (FPG) ≥ 125 mg/dL (7.0 mmol/L), where fasting is defined as no caloric intake for at least 8 h or (ii) two-hour post-load plasma glucose ≥200 mg/dL (11.1 mmol/L) during an OGTT.
Additionally, patients meeting either of the following standards were diagnosed as having pre-diabetes: (i) FPG ≥110 mg/dL (6.1 mmol/L) and <125 mg/dL (7.0 mmol/L); or (ii) two-hour post-load plasma glucose ≥140 mg/dL (7.8 mmol/L) and <200 mg/dL (11.1 mmol/L) during an OGTT.
Blood sample collection was performed as follows: After overnight fasting, 2 mL of blood was collected from the median cubital vein of each patient before breakfast and then stored in ethylenediaminetetraacetic acid (EDTA) anticoagulant vacutainers. The total RNA was then extracted as soon as possible.
RNA extraction and Q-PCR
A fast total RNA extraction kit (Biotech, Beijing, China) was used to extract total RNA from 1 mL of whole blood according to the manufacturer’s instructions. RNA was then dissolved in RNase-free water. The yield and purity were measured by a NanoDrop 2000 instrument (Thermo Scientific, Waltham, MA, USA). The integrity of the RNA was determined by 1% formaldehyde denaturing gel electrophoresis. A PrimeScript RT Reagent Kit (Takara Bio, Nojihigashi, Kusatsu, Japan) was used for the production of complementary DNA (cDNA) by reverse transcription, according to the manufacturer’s instructions. Q-PCR was performed using SYBR-Green Premix Ex Taq (Takara Bio, Nojihigashi, Kusatsu, Japan) and monitored by an ABI PRISM 7500 Sequence Detection System (Applied Biosystems, Life Technologies, Waltham, MA, USA). The relative expression levels of circRNAs were determined via Q-PCR. The sequences of the primers used in the Q-PCR assay are shown in Supplementary Table 4.
CircRNA microarray analysis
The RNAs of the peripheral blood of six control subjects and six T2DM patients were extracted for microarray analysis. The purity and concentration of the RNA were determined by a NanoDrop ND-1000 instrument (Thermo Scientific, Waltham, MA, USA). The integrity of the RNA was evaluated using a Bioanalyzer 2100 (Agilent Technologies, Santa Clara, CA, USA). The extracted RNAs were digested, dephosphorylated, denatured, amplified and labeled with Cy3-dCTP according to the manufacturer’s specifications. The purified RNAs were hybridized to a microarray (Agilent human circRNA Array V2.0) containing 170,340 human circRNA probes. The microarray data of the circRNAs were then analyzed using GeneSpring software V13.0 (Agilent Technologies, Santa Clara, CA, USA). The thresholds were as follows: fold change, ≥2 or ≤−2; P < 0.05 according to the t test.
Data analysis
Variables with different distributions were expressed as means ± standard deviations, medians (quartiles) or percentages when they fit. In the scatterplot of circRNA expression, the horizontal lines represent the median values. The Chi-square test was used for categorical variables, whereas the Kolmogorov–Smirnov and Shapiro–Wilk tests were performed to check data normality for continuous variables, followed by the test for homogeneity of variances. The clinical and demographic indicators were checked for significant differences by one-way analysis of variance (ANOVA), if the continuous variables were consistent with the normal distribution and homogeneity of variance; if not, the Kruskal–Wallis H test was used. The clinical diagnostic value of a given circRNA was verified by ROC curve analysis, and when the AUC was equal to 0.5, the circRNA was defined as having no diagnostic value. Furthermore, logistic regression analysis was performed to obtain an odds ratio (OR) when the relative expression of circRNAs was expanded by ten times. P < 0.05 was considered statistically significant. All statistical analyses were conducted using SPSS 22.0 (SPSS Inc., Chicago, IL, USA).
Results
Expression profiles of circRNAs in the peripheral blood of diabetic patients
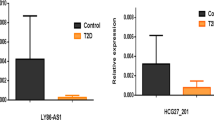
To investigate the expression profiles of circRNAs in healthy individuals and T2DM patients, six healthy subjects and six T2DM patients were selected. Microarray analysis of the expression profiles of circRNAs in peripheral blood was performed using the Agilent human circRNA Array (V2.0). The results showed clear differences in the expression profiles of circRNAs between the two groups (Fig. 2). Differential expression was detected in a total of 489 circRNAs; of these, 78 were upregulated and 411 were downregulated in the T2DM group (Supplementary Table 5). To obtain the biomarkers that would be most applicable in clinical practice, the candidate biomarkers were selected from the 78 upregulated circRNAs utilizing stricter screening criteria: fold change >2.4 and P < 0.01. Five circRNAs met these standards: hsa_circ_0068087, hsa_circ_0054633, hsa_circ_0124636, hsa_circ_0139110 and hsa_circ_0018508 (highlighted in Supplementary Table 5). These circRNAs were used as candidate biomarkers in a subsequent validation utilizing a larger cohort.
Heat map of the circRNA microarray profiles in control individuals and T2DM patients. The expression of circRNAs is hierarchically clustered on the y-axis, and blood samples are hierarchically clustered on the x-axis. The expression levels are presented in red and green, which indicate upregulated and downregulated circRNAs, respectively. Numbers marked with A and B are from control individuals and T2DM patients, respectively
The expression profile of circRNAs verified by Q-PCR
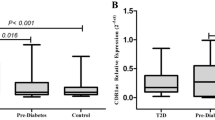
To validate the five selected candidate circRNAs, Q-PCR was conducted in an independent cohort (control group, n = 20; pre-diabetes group, n = 20; and T2DM group, n = 20). The results are shown in Fig. 3. The levels of hsa_circ_0124636 and hsa_circ_0139110 expression among the three groups presented no significant differences. The levels of hsa_circ_0018508 expression in the pre-diabetes and T2DM groups did not differ, but both were higher than that of the control group. The expression levels of hsa_circ_0054633 and hsa_circ_0068087 were significantly different among the three groups and increased gradually from the control group to the pre-diabetes group to the T2DM group.
Expression levels of the selected circRNAs quantified by Q-PCR. a, b and c show the relative expression levels of hsa_circ_0054633, hsa_circ_0068087 and hsa_circ_0018508. *P < 0.05; d, f present the ROC curve analyses of hsa_circ_0054633 and hsa_circ_0068087 for the diagnosis of pre-diabetes; e, g are the ROC curve analyses of hsa_circ_0054633 and hsa_circ_0068087 for the diagnosis of T2DM. The AUC values are given on the graphs
ROC curve analysis of circRNAs with differential expression
To determine the diagnostic values of hsa_circ_0054633 and hsa_circ_0068087 for pre-diabetes and T2DM, ROC curve analysis was performed (Fig. 3). The AUCs of hsa_circ_0054633 for the diagnosis of pre-diabetes and T2DM were 0.747 ([0.589–0.906], P = 0.007) and 0.72 ([0.562–0.878], P = 0.017), respectively. The AUCs of hsa_circ_0068087 for the diagnosis of pre-diabetes and T2DM were 0.692 ([0.529–0.856], P = 0.037) and 0.717 ([0.557–0.878], P = 0.019); the sensitivity and specificity are shown in Table 1. Because hsa_circ_0054633 showed higher AUC and lower P values than hsa_circ_0068087, the former was chosen as the diagnostic biomarker for pre-diabetes and T2DM.
Further clinical validation of the biomarker
To verify its clinical diagnostic capability, hsa_circ_0054633 was tested in another cohort (control group, n = 60; pre-diabetes group, n = 63; and T2DM group, n = 64). The results are shown in Fig. 4. The level of hsa_circ_0054633 expression increased gradually from the control group to the pre-diabetes group to the T2DM group, with a fold change of 1.8 between the first two groups and 1.7 between the latter two groups. Then, the ROC curve analysis was performed. When used as a biomarker for the diagnosis of pre-diabetes and T2DM, the AUC, threshold, sensitivity and specificity of hsa_circ_0054633 were 0.751 ([0.666–0.835], P < 0.001), 0.103, 0.905 and 0.483, respectively, and 0.793 ([0.716–0.871], P < 0.001), 0.270, 0.719 and 0.778, respectively. The crude ORs were 3.05 ([1.803–5.159], P < 0.001) and 2.056 ([1.530–2.762], P < 0.001), respectively. After introducing the risk factors of T2DM (smoking, hypertension, body mass index [BMI], total cholesterol [TC], triglycerides [TG], high-density lipoprotein [HDL] and low-density lipoprotein [LDL]), the AUCs increased to 0.841 ([0.773–0.910], P < 0.001) with a sensitivity of 0.778 and a specificity of 0.783 and 0.834 ([0.762–0.905], P < 0.001) with a sensitivity of 0.766 and a specificity of 0.794, respectively. The adjusted ORs were 6.797 ([3.025–15.273], P < 0.001) and 2.769 ([1.881–4.077], P < 0.001), respectively.
Biomarker expression levels in different gender and age groups
To investigate the levels of hsa_circ_0054633 expression in different gender and age groups, the three groups of the third cohort were divided according to age (cut-off: 50 years old) and gender. The results are shown in Table 2 and indicate that no differential expression of hsa_circ_0054633 was identified between the different gender and age groups.
Discussion
The high morbidity of T2DM and its various complications severely threaten human health. In the advanced stages of T2DM, patients often experience various complications, which severely impact their quality of life. Multiple large-scale investigations have revealed that intensive glucose-lowing therapy in the early phases of T2DM can benefit patients substantially, reducing the incidences of macrovascular and microvascular complications [26, 27]. However, in the early stages of T2DM, most patients are asymptomatic, and they rarely visit hospitals to seek diagnosis and therapy.
Current diagnostic methods have various deficiencies for the early diagnosis of T2DM: OGTT is the gold standard for diagnosing T2DM. However, because this procedure is time-consuming and complicated, it is only considered when there is a strong suspicion that a patient has T2DM. FPG is convenient, but the rate of missed diagnoses is high [28]. Finally, hemoglobin A1c (HbA1c) test has not been standardized in Chinese hospitals. Therefore, a convenient, highly specific and sensitive diagnostic method is urgently needed to facilitate the early diagnosis of T2DM.
Because of their convenience of sampling and low cost, hematological markers play an important role in the diagnosis of many diseases. One of the most important functions of circRNAs is their role as “miRNA sponges,” which competitively bind miRNAs to generate post-transcriptional regulation. Long non-coding RNAs (lncRNAs) can also interact with miRNAs to regulate gene translation, suggesting a potential correlation among the three types of non-coding RNAs [29]. Some lncRNAs and miRNAs have been demonstrated to be involved in the occurrence and development of T2DM; furthermore, they can be used as biomarkers for T2DM diagnosis [30, 31]. CircRNAs are much more stable than linear RNAs in cells, and in some tissues, their expression levels are ten times higher [10]; thus, circRNAs make better biomarkers.
The current study revealed significant differences between the expression levels of circRNAs in the peripheral blood of T2DM patients and that of healthy subjects. To reduce the interference of factors other than T2DM to the minimum, we strictly matched the demographic and clinical characteristics (especially the hematological indicators, such as the blood cell count and percentage of white blood cells) of the first cohort. But individual differences among the patients were huge; the differential expression of circRNA might be attributed to changes in the activation states of different blood cell types. Therefore, we selected the five circRNAs with the most significant differences as candidate biomarkers. Hsa_circ_0054633 showed the highest diagnostic value for pre-diabetes and T2DM among the five candidate biomarkers and was further verified in another cohort. It continued to have reliable diagnostic value, suggesting that hsa_circ_0054633 has the potential to be used as a diagnostic biomarker for pre-diabetes and T2DM in clinical practice.
The field of circRNAs is quite new, and thus, to the best of our knowledge, no definite evidence demonstrating the functions of hsa_circ_0054633 is available. The results of gene ontology (GO) analysis revealed that hsa_circ_0054633 not only participates in biological processes, such as cell cycle and mitotic cell cycle arrest, but is strongly correlated with molecular catabolism. The cell cycle is the basic process of cellular life activities. The proliferation of β cells is regulated by cell cycle progress, and decreased β cell proliferation is the major cause of insufficient insulin secretion [32], which is the basic characteristic of T2DM. Besides, T2DM is a chronic metabolic disease characterized by disordered carbohydrate, lipid and protein metabolism, and we hypothesize that hsa_circ_0054633 may participate in the pathogenesis of T2DM by influencing the cellular metabolism and cell cycle.
To the best of our knowledge, this study is the first to investigate the expression profiles of circRNAs in the peripheral blood of patients with T2DM and to validate the utility of hsa_circ_0054633 as a diagnostic biomarker for pre-diabetes and T2DM. The biomarker identified in this study (hsa_circ_0054633) can be easily tested using peripheral blood. Furthermore, its relatively low cost and high specificity and sensitivity make it a potentially highly useful tool for the diagnosis of T2DM and pre-diabetes.
In the present study, only hsa_circ_0054633 was validated; thus, the expression profiles of other circRNAs in T2DM and pre-diabetes patients remain to be explored. Additionally, this was a single-center study, with a high geographic concentration of subjects. Therefore, whether populations in other regions exhibit similar circRNAs expression profiles is unknown. Thus, the results of our study require further verification in larger and more diverse cohorts.
References
Rahelic D (2016) 7th edition of idf diabetes ATLAS–CALL for immediate action. Lijec Vjesn 138:57–58
Ghasemi A, Tohidi M, Derakhshan A, Hasheminia M, Azizi F et al (2015) Cut-off points of homeostasis model assessment of insulin resistance, beta-cell function, and fasting serum insulin to identify future type 2 diabetes: Tehran Lipid and Glucose Study. Acta Diabetol 52:905–915. doi:10.1007/s00592-015-0730-3
Zaccardi F, Kurl S, Pitocco D, Ronkainen K, Laukkanen JA (2015) Serum fructosamine and risk of type 2 diabetes mellitus among middle-age Finnish men: a 23-year population-based prospective study. Acta Diabetol 52:161–166. doi:10.1007/s00592-014-0625-8
Chidambaram M, Liju S, Saboo B, Sathyavani K, Viswanathan V et al (2016) Replication of genome-wide association signals in Asian Indians with early-onset type 2 diabetes. Acta Diabetol. doi:10.1007/s00592-016-0889-2
Teumer A, Tin A, Sorice R, Gorski M, Yeo NC et al (2016) Genome-wide association studies identify genetic loci associated with albuminuria in diabetes. Diabetes 65:803–817. doi:10.2337/db15-1313
Gaulton KJ, Ferreira T, Lee Y, Raimondo A, Magi R et al (2015) Genetic fine mapping and genomic annotation defines causal mechanisms at type 2 diabetes susceptibility loci. Nat Genet 47:1415–1425. doi:10.1038/ng.3437
Esteller M (2011) Non-coding RNAs in human disease. Nat Rev Genet 12:861–874. doi:10.1038/nrg3074
Barrett SP, Wang PL, Salzman J (2015) Circular RNA biogenesis can proceed through an exon-containing lariat precursor. Elife 4:e07540. doi:10.7554/eLife.07540
Li Z, Huang C, Bao C, Chen L, Lin M et al (2015) Exon-intron circular RNAs regulate transcription in the nucleus. Nat Struct Mol Biol 22:256–264. doi:10.1038/nsmb.2959
Jeck WR, Sorrentino JA, Wang K, Slevin MK, Burd CE et al (2013) Circular RNAs are abundant, conserved, and associated with ALU repeats. RNA 19:141–157. doi:10.1261/rna.035667.112
Schwanhausser B, Busse D, Li N, Dittmar G, Schuchhardt J et al (2011) Global quantification of mammalian gene expression control. Nature 473:337–342. doi:10.1038/nature10098
Memczak S, Jens M, Elefsinioti A, Torti F, Krueger J et al (2013) Circular RNAs are a large class of animal RNAs with regulatory potency. Nature 495:333–338. doi:10.1038/nature11928
Hansen TB, Jensen TI, Clausen BH, Bramsen JB, Finsen B et al (2013) Natural RNA circles function as efficient microRNA sponges. Nature 495:384–388. doi:10.1038/nature11993
Conn SJ, Pillman KA, Toubia J, Conn VM, Salmanidis M et al (2015) The RNA binding protein quaking regulates formation of circRNAs. Cell 160:1125–1134. doi:10.1016/j.cell.2015.02.014
Ashwal-Fluss R, Meyer M, Pamudurti NR, Ivanov A, Bartok O et al (2014) circRNA biogenesis competes with pre-mRNA splicing. Mol Cell 56:55–66. doi:10.1016/j.molcel.2014.08.019
Jeck WR, Sharpless NE (2014) Detecting and characterizing circular RNAs. Nat Biotechnol 32:453–461. doi:10.1038/nbt.2890
Lukiw WJ (2013) Circular RNA (circRNA) in Alzheimer’s disease (AD). Front Genet 4:307. doi:10.3389/fgene.2013.00307
Bachmayr-Heyda A, Reiner AT, Auer K, Sukhbaatar N, Aust S et al (2015) Correlation of circular RNA abundance with proliferation–exemplified with colorectal and ovarian cancer, idiopathic lung fibrosis, and normal human tissues. Sci Rep 5:8057. doi:10.1038/srep08057
Liu Q, Zhang X, Hu X, Dai L, Fu X et al (2016) Circular RNA related to the chondrocyte ECM regulates MMP13 Expression by functioning as a MiR-136 ‘Sponge’ in human cartilage degradation. Sci Rep 6:22572. doi:10.1038/srep22572
Burd CE, Jeck WR, Liu Y, Sanoff HK, Wang Z et al (2010) Expression of linear and novel circular forms of an INK4/ARF-associated non-coding RNA correlates with atherosclerosis risk. PLoS Genet 6:e1001233. doi:10.1371/journal.pgen.1001233
Li P, Chen S, Chen H, Mo X, Li T et al (2015) Using circular RNA as a novel type of biomarker in the screening of gastric cancer. Clin Chim Acta 444:132–136. doi:10.1016/j.cca.2015.02.018
Li F, Zhang L, Li W, Deng J, Zheng J et al (2015) Circular RNA ITCH has inhibitory effect on ESCC by suppressing the Wnt/beta-catenin pathway. Oncotarget 6:6001–6013. doi:10.18632/oncotarget.3469
Qin M, Liu G, Huo X, Tao X, Sun X et al (2016) Hsa_circ_0001649: a circular RNA and potential novel biomarker for hepatocellular carcinoma. Cancer Biomark 16:161–169. doi:10.3233/cbm-150552
Xu H, Guo S, Li W, Yu P (2015) The circular RNA Cdr1as, via miR-7 and its targets, regulates insulin transcription and secretion in islet cells. Sci Rep 5:12453. doi:10.1038/srep12453
Alberti KG, Zimmet PZ (1998) Definition, diagnosis and classification of diabetes mellitus and its complications. Part 1: diagnosis and classification of diabetes mellitus provisional report of a WHO consultation. Diabet Med 15:539–553. doi:10.1002/(sici)1096-9136(199807)15:7<539:aid-dia668>3.0.co;2-s
Holman RR, Paul SK, Bethel MA, Matthews DR, Neil HA (2008) 10-year follow-up of intensive glucose control in type 2 diabetes. N Engl J Med 359:1577–1589. doi:10.1056/NEJMoa0806470
Gaede P, Lund-Andersen H, Parving HH, Pedersen O (2008) Effect of a multifactorial intervention on mortality in type 2 diabetes. N Engl J Med 358:580–591. doi:10.1056/NEJMoa0706245
Gatling W, Begley J (2001) Diagnosing diabetes mellitus in clinical practice: is fasting plasma glucose a good initial test? Pract Diabetes Int 18:89–93
Tay Y, Rinn J, Pandolfi PP (2014) The multilayered complexity of ceRNA crosstalk and competition. Nature 505:344–352. doi:10.1038/nature12986
Moran I, Akerman I, van de Bunt M, Xie R, Benazra M et al (2012) Human beta cell transcriptome analysis uncovers lncRNAs that are tissue-specific, dynamically regulated, and abnormally expressed in type 2 diabetes. Cell Metab 16:435–448. doi:10.1016/j.cmet.2012.08.010
Guay C, Roggli E, Nesca V, Jacovetti C, Regazzi R (2011) Diabetes mellitus, a microRNA-related disease? Transl Res 157:253–264. doi:10.1016/j.trsl.2011.01.009
Rane SG, Reddy EP (2000) Cell cycle control of pancreatic beta cell proliferation. Front Biosci 5:D1–19
Funding
This work is supported by the Department of Science and Technology of Henan province (Grant Number: 122102310620).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Ethical standard
This experiment was approved by the Ethics Committee of the People's Hospital of Zhengzhou University. This study protocol conformed to the ethical guidelines of the 1975 Declaration of Helsinki. All subjects signed informed consent forms.
Human and animal rights
All procedures followed were in accordance with the ethical standards of the responsible committee on human experimentation (institutional and national) and with the Helsinki Declaration of 1975, as revised in 2008.
Informed consent
Informed consent was obtained from all patients for being included in the study.
Additional information
Managed by Antonio Secchi.
Zhenzhou Zhao and Xuejie Li have contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Zhao, Z., Li, X., Jian, D. et al. Hsa_circ_0054633 in peripheral blood can be used as a diagnostic biomarker of pre-diabetes and type 2 diabetes mellitus. Acta Diabetol 54, 237–245 (2017). https://doi.org/10.1007/s00592-016-0943-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00592-016-0943-0