Abstract
Retinal degeneration diseases, such as age-related macular degeneration and retinitis pigmentosa, affect millions of people worldwide and are major causes of irreversible blindness. Effective treatments for retinal degeneration, including drug therapy, gene augmentation or transplantation approaches, have been widely investigated. Nevertheless, more research should be dedicated to therapeutic methods to improve future clinical treatments. Recently, with the rapid development of genome-editing technology, gene therapy has become a potentially effective treatment for retinal degeneration diseases. A clustered regularly interspaced short palindromic repeats (CRISPR)/CRISPR-associated protein 9 (Cas9) system has been developed as a powerful genome-editing tool in ophthalmic studies. The CRISPR/Cas9 system has been widely applied in basic research to develop animal models and gene therapies in vivo. With the ability to self-renew and the potential to differentiate into different types of cells, induced pluripotent stem cells (iPSCs) have already been used as a promising tool for understanding disease pathophysiology and evaluating the effect of drug and gene therapeutics. iPSCs are also a cell source for autologous transplantation. In this review, we compared genome-editing strategies and highlighted the advantages and concerns of the CRISPR/Cas9 system. Moreover, the latest progress and applications of the CRISPR/Cas9 system and its combination with iPSCs for the treatment of retinal degenerative diseases are summarized.



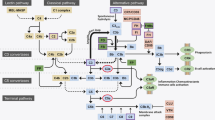
(reprinted from Liao et al. (2017), Copyright (2017), with permission from Elsevier)
Similar content being viewed by others
References
Acland GM, Aguirre GD, Ray J, Zhang Q, Aleman TS, Cideciyan AV, Pearcekelling SE, Anand V, Zeng Y, Maguire AM (2001) Gene therapy restores vision in a canine model of childhood blindness. Nat Genet 28:92–95
Acland GM, Aguirre GD, Bennett J, Aleman TS, Cideciyan AV, Bennicelli J, Dejneka NS, Pearcekelling SE, Maguire AM, Palczewski K (2005) Long-term restoration of rod and cone vision by single dose rAAV-mediated gene transfer to the retina in a canine model of childhood blindness. Mol Ther J Am Soc Gene Ther 12:1072
Arno G, Agrawal SA, Eblimit A, Bellingham J, Xu M, Wang F, Chakarova C, Parfitt DA, Lane A, Burgoyne T (2016) Mutations in REEP6 cause autosomal-recessive retinitis pigmentosa. Am J Hum Genet 99:1305
Auer TO, Duroure K, De CA, Concordet JP, Del BF (2014) Highly efficient CRISPR/Cas9-mediated knock-in in zebrafish by homology-independent DNA repair. Genome Res 24:142
Bassuk AG, Zheng A, Li Y, Tsang SH, Mahajan VB (2016) Precision medicine: genetic repair of retinitis pigmentosa in patient-derived stem cells. Sci Rep 6:19969
Beltran WA, Cideciyan AV, Boye SE, Ye GJ, Iwabe S, Dufour VL, Marinho LF, Swider M, Kosyk MS, Sha J (2017) Optimization of retinal gene therapy for X-linked retinitis pigmentosa due to RPGR mutations. Mol Ther 25(8):1866–1880. https://doi.org/10.1016/j.ymthe.2017.05.004
Benjamin B, Lv W, Lu B, Jones MK, Yuchun T, Kim KJ, Rachelle L, Abbasi AA, Breunig JJ, Svendsen CN (2016) In vivocrispr/Cas9 gene Editing corrects retinal dystrophy in the S334ter-3 rat model of autosomal dominant retinitis pigmentosa. Mol Ther J Am Soc Gene Ther 24:556
Bennicelli J, Wright JF, Komaromy A, Jacobs JB, Hauck B, Zelenaia O, Mingozzi F, Hui D, Chung D, Rex TS (2008) Reversal of blindness in animal models of leber congenital amaurosis using optimized AAV2-mediated gene transfer. Mol Ther J Am Soc Gene Ther 16:458
Burnight ER, Wiley LA, Drack AV, Braun TA, Anfinson KR, Kaalberg EE, Halder JA, Affatigato LM, Mullins RF, Stone EM (2014) CEP290 gene transfer rescues Leber congenital amaurosis cellular phenotype. Gene Ther 21:662
Camara MFDL, Nanda A, Salvetti AP, Fischer MD, Maclaren RE (2018) Gene therapy for the treatment of X-linked retinitis pigmentosa. Expert Opin Orphan Drugs 6(3):167–177. https://doi.org/10.1080/21678707.2018.1444476
Cereso N, Pequignot MO, Robert L, Becker F, Luca VD, Nabholz N, Rigau V, Vos JD, Hamel CP, Kalatzis V (2014) Proof of concept for AAV2/5-mediated gene therapy in iPSC-derived retinal pigment epithelium of a choroideremia patient. Mol Ther Methods Clin Dev 1:14011
Chang N, Sun C, Gao L, Zhu D, Xu X, Zhu X, Xiong JW, Xi JJ (2013) Genome editing with RNA-guided Cas9 nuclease in Zebrafish embryos. Cell Res 23:465
Chavala SH, Sari A, Lewis H, Pauer GJ, Simpson E, Hagstrom SA, Traboulsi EI (2005) An Arg311Gln NR2E3 mutation in a family with classic Goldmann–Favre syndrome. Br J Ophthalmol 89:1065–1066
Chrenek MA, Nickerson JM, Boatright JH (2016) CRISPR challenges in treating retinal disease. Asia Pac J Ophthalmol (Philadelphia Pa) 5:304
Cox DBT, Platt RJ, Zhang F (2015) Therapeutic genome editing: prospects and challenges. Nat Med 21:121
Da CR, Röger C, Segelken J, Barben M, Grimm C, Neidhardt J (2016) A novel method combining vitreous aspiration and intravitreal AAV2/8 injection results in retina-wide transduction in adult mice. Invest Ophthalmol Vis Sci 57:5326–5334
Day TP, Byrne LC, Schaffer DV, Flannery JG (2014) Advances in AAV vector development for gene therapy in the retina. Adv Exp Med Biol 801:687
Fischer MD, Mcclements ME, de la Camara CMF, Bellingrath JS, Dauletbekov D, Ramsden SC, Hickey DG, Barnard AR, Maclaren RE (2017) Codon-optimized RPGR improves stability and efficacy of AAV8 gene therapy in two mouse models of X-linked retinitis pigmentosa. Mol Ther 25(8):1854–1865. https://doi.org/10.1016/j.ymthe.2017.05.005
Fu Y, Foden JA, Khayter C, Maeder ML, Reyon D, Joung JK, Sander JD (2013) High frequency off-target mutagenesis induced by CRISPR-Cas nucleases in human cells. Nat Biotechnol 31:822
He Y, Zhang Y, Su G (2015) Recent advances in treatment of retinitis pigmentosa. Curr Stem Cell Res Ther 10:258–265
Homma K, Okamoto S, Mandai M, Gotoh N, Rajasimha HK, Chang YS, Chen S, Li W, Cogliati T, Swaroop A (2014) Developing rods transplanted into the degenerating retina of Crx-knockout mice exhibit neural activity similar to native photoreceptors. Stem Cells 31:1149–1159
Huang Y, Enzmann V, Ildstad ST (2011) Stem cell-based therapeutic applications in retinal degenerative diseases. Stem Cell Rev Rep 7:434–445
Jacobson SG, Marmor MF, Kemp CM, Knighton RW (1990) SWS (blue) cone hypersensitivity in a newly identified retinal degeneration. Invest Ophthalmol Vis Sci 31:827
Jager RD, Mieler WF, Miller JW (2008) Age-related macular degeneration. N Engl J Med 358:2606
Jasin M, Haber JE (2016) The Democratization of gene editing: insights from site-specific cleavage and double-strand break repair. DNA Repair 44:6
Jinek M, East A, Cheng A, Lin S, Ma E, Doudna J (2013) RNA-programmed genome editing in human cells. Elife 2:e00471
Kim K, Park SW, Kim JH, Lee SH, Kim D, Koo T, Kim KE, Kim JH, Kim JS (2017) Genome surgery using Cas9 ribonucleoproteins for the treatment of age-related macular degeneration. Genome Res 27:419
Kleinstiver BP, Pattanayak V, Prew MS, Tsai SQ, Nguyen NT, Zheng Z, Joung JK (2016) High-fidelity CRISPR–Cas9 nucleases with no detectable genome-wide off-target effects. Nature 529:490
Latella MC, Salvo MTD, Cocchiarella F, Benati D, Grisendi G, Comitato A, Marigo V, Recchia A (2016) In vivo editing of the human mutant rhodopsin gene by electroporation of plasmid-based CRISPR/Cas9 in the mouse retina. Mol Ther Nucleic Acids 5:e389
Liao HK, Hatanaka F, Araoka T, Reddy P, Wu MZ, Sui Y, Yamauchi T, Sakurai M, O’Keefe DD, Núñez-Delicado E (2017) In vivo target gene activation via CRISPR/Cas9-mediated trans-epigenetic modulation. Cell 171:1495–1507
Lim Y, Bak SY, Sung K, Jeong E, Lee SH, Kim JS, Bae S, Kim SK (2016) Structural roles of guide RNAs in the nuclease activity of Cas9 endonuclease. Nat Commun 7:13350
Ma Y, Zhang X, Shen B, Lu Y, Chen W, Ma J, Bai L, Huang X, Zhang L (2014) Generating rats with conditional alleles using CRISPR/Cas9. Cell Res 24:122
Maeder ML, Linder SJ, Cascio VM, Fu Y, Ho QH, Joung JK (2013) Crispr rna-guided activation of endogenous human genes. Nat Methods 10:977
Mali P, Yang L, Esvelt KM, Aach J, Guell M, Dicarlo JE, Norville JE, Church GM (2013) RNA-guided human genome engineering via Cas9. Science 339:823
Marmor MF, Jacobson SG, Foerster MH, Kellner U, Weleber RG (1990) Diagnostic clinical findings of a new syndrome with night blindness, maculopathy, and enhanced S cone sensitivity. Am J Ophthalmol 110(2):124–134
Nelson CE, Gersbach CA (2016) Engineering delivery vehicles for genome editing. Ann Rev Chem Biomol Eng 7:637
Pawlyk BS, Adamian M, Sun X, Bulgakov OV, Shu X, Smith AJ, Berson EL, Ali RR, Khani S, Wright AF (2016) photoreceptor rescue by an abbreviated human RPGR gene in a murine model of X-linked retinitis pigmentosa. Gene Ther 23:196–204
Ran FA, Hsu PD, Lin CY, Gootenberg JS, Konermann S, Trevino AE, Scott DA, Inoue A, Matoba S, Zhang Y (2013) Double nicking by RNA-guided CRISPR Cas9 for enhanced genome editing specificity. Cell 154:1380–1389
Schaefer KA, Wu WH, Colgan DF, Tsang SH, Bassuk AG, Mahajan VB (2017) Unexpected mutations after CRISPR-Cas9 editing in vivo. Nat Methods 14:547
Schaefer KA, Darbro BW, Colgan DF, Tsang SH, Bassuk AG, Mahajan VB (2018) Corrigendum and follow-up: whole genome sequencing of multiple CRISPR-edited mouse lines suggests no excess mutations. https://doi.org/10.1101/154450
Schwank G, Koo BK, Sasselli V, Dekkers JF, Heo I, Demircan T, Sasaki N, Boymans S, Cuppen E, Ck VDE (2013) Functional repair of CFTR by CRISPR/Cas9 in intestinal stem cell organoids of cystic fibrosis patients. Cell Stem Cell 13:653
Senís E, Fatouros C, Große S, Wiedtke E, Niopek D, Mueller AK, Börner K, Grimm D (2015) CRISPR/Cas9-mediated genome engineering: an adeno-associated viral (AAV) vector toolbox. Biotechnol J 9:1402–1412
Shin J, Jiang F, Liu JJ, Bray NL, Rauch BJ, Baik SH, Nogales E, Bondydenomy J, Corn JE, Doudna JA (2017) Disabling Cas9 by an anti-CRISPR DNA mimic. Sci Adv 3:e1701620
Stemmer M, Thumberger T, Del SKM, Wittbrodt J, Mateo JL (2015) CCTop: an intuitive, flexible and reliable CRISPR/Cas9 target prediction tool. PLoS One 10:e0124633
Suzuki K, Tsunekawa Y, Hernandezbenitez R, Wu J, Zhu J, Kim EJ, Hatanaka F, Yamamoto M, Araoka T, Li Z (2016) In vivo genome editing via CRISPR/Cas9 mediated homology-independent targeted integration. Nature 540:144
Thomas N, Robin C, Tom VN, Dionysia D, Jannick L, Jurgen H, Annekatrien B, Wouter S, Trees L, Dieter D (2016) CRISPR/Cas9 mediated knockout ofrb1andrbl1leads to rapid and penetrant retinoblastoma development inXenopus tropicalis. Sci Rep 6:35264
Tucker BA, Park IH, Qi SD, Klassen HJ, Jiang C, Yao J, Redenti S, Daley GQ, Young MJ (2011) Transplantation of adult mouse iPS cell-derived photoreceptor precursors restores retinal structure and function in degenerative mice. PLoS One 6:e18992
Tucker BA, Mullins RF, Stone EM (2014) Stem cells for investigation and treatment of inherited retinal disease. Hum Mol Genet 23:R9
Wiley LA, Burnight ER, Songstad AE, Drack AV, Mullins RF, Stone EM, Tucker BA (2015) Patient-specific induced pluripotent stem cells (iPSCs) for the study and treatment of retinal degenerative diseases. Prog Retin Eye Res 44:15–35
Wu WH, Tsai YT, Justus S, Cho GY, Sengillo JD, Xu Y, Cabral T, Lin CS, Bassuk AG, Mahajan VB (2016) CRISPR repair reveals causative mutation in a preclinical model of retinitis pigmentosa: a brief methodology. Mol Ther J Am Soc Gene Ther 24:1388–1394
Yanik M, Müller B, Song F, Gall J, Wagner F, Wende W, Lorenz B, Stieger K (2016) In vivo genome editing as a potential treatment strategy for inherited retinal dystrophies. Prog Retin Eye Res 56:1
Yanik M, Müller B, Fei S, Gall J, Wagner F, Wende W, Lorenz B, Stieger K (2017) In vivo genome editing as a potential treatment strategy for inherited retinal dystrophies. Prog Retin Eye Res 56:1
Zetsche B, Heidenreich M, Mohanraju P, Fedorova I, Kneppers J, Degennaro EM, Winblad N, Choudhury SR, Abudayyeh OO, Gootenberg JS (2017) Multiplex gene editing by CRISPR-Cpf1 using a single crRNA array. Nat Biotechnol 35:31
Zheng A, Li Y, Tsang SH (2015) Personalized therapeutic strategies for patients with retinitis pigmentosa. Expert Opin Biol Ther 15:391–402
Zhong H, Chen Y, Li Y, Chen R, Mardon G (2015) CRISPR-engineered mosaicism rapidly reveals that loss of Kcnj13 function in mice mimics human disease phenotypes. Sci Rep 5:8366
Zhou J, Shen B, Zhang W, Wang J, Yang J, Chen L, Zhang N, Zhu K, Xu J, Hu B (2014) One-step generation of different immunodeficient mice with multiple gene modifications by CRISPR/Cas9 mediated genome engineering. Int J Biochem Cell Biol 46:49
Zhu H, Misel L, Graham M, Robinson ML, Liang C (2016) CT-finder: a web service for CRISPR optimal target prediction and visualization. Sci Rep 6:25516
Acknowledgements
We thank Jin Yang, Xiaomin Zhang and Xiaorong Li for comments and suggestions, Jin Yang and Zhiqing Li for editing the manuscript, Bincui Cai for writing the manuscript, and Shuo Sun for assistance in generating the table. This work was supported by the National Natural Science Foundation (81670875; 81500745), China; the Natural Science Foundation of Tianjin City (18JCQNJC10700); the Natural Science Foundation of Tianjin City (17JCYBJC27200); and a grant from the Dr. Henry Norman Bethune: LangMu Young Scientist Scholarship (BJ-LM2015008L).
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Cai, B., Sun, S., Li, Z. et al. Application of CRISPR/Cas9 technologies combined with iPSCs in the study and treatment of retinal degenerative diseases. Hum Genet 137, 679–688 (2018). https://doi.org/10.1007/s00439-018-1933-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00439-018-1933-9