Abstract
The centromere is essential for the segregation of chromosomes, as it serves as attachment site for microtubules to mediate chromosome segregation during mitosis and meiosis. In most organisms, the centromere is restricted to one chromosomal region that appears as primary constriction on the condensed chromosome and is partitioned into two chromatin domains: The centromere core is characterized by the centromere-specific histone H3 variant CENP-A (also called cenH3) and is required for specifying the centromere and for building the kinetochore complex during mitosis. This core region is generally flanked by pericentric heterochromatin, characterized by nucleosomes containing H3 methylated on lysine 9 (H3K9me) that are bound by heterochromatin proteins. During mitosis, these two domains together form a three-dimensional structure that exposes CENP-A-containing chromatin to the surface for interaction with the kinetochore and microtubules. At the same time, this structure supports the tension generated during the segregation of sister chromatids to opposite poles. In this review, we discuss recent insight into the characteristics of the centromere, from the specialized chromatin structures at the centromere core and the pericentromere to the three-dimensional organization of these regions that make up the functional centromere.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The centromere is a conserved and essential feature of eukaryotic chromosomes that enables the equal segregation of genetic material into daughter cells during cell division. The centromere generally appears as primary constriction of mitotic chromosomes, first described by Walther Flemming in 1882 (Flemming 1882). It was apparent early on that chromosomes are segregated by attachment of microtubules to the primary constriction. The kinetochore—the proteinaceous structure that links the chromosomes to the microtubules for the segregation of chromosomes—was described in the 1960s (Luykx 1965; Brinkley and Stubblefield 1966; Jokelainen 1967) and has since been studied in great detail. The structure and organization of the chromatin forming the centromere, however, have remained more obscure. One key feature of centromeric chromatin is the incorporation of the histone H3 variant centromere protein A (CENP-A), also called cenH3 (Earnshaw and Rothfield 1985; Palmer et al. 1987; Talbert et al. 2012; Earnshaw et al. 2013; Talbert and Henikoff 2013). CENP-A is an almost universally conserved centromeric protein that plays a key role in specifying the centromere, where it replaces H3 in centromeric nucleosomes and is necessary for centromere function (McKinley and Cheeseman 2016). As is the case for CENP-A, many protein components of centromeric chromatin are conserved, but the organization of CENP-A nucleosomes and the DNA sequences on which the centromere is established vary widely between taxa (Steiner and Henikoff 2015). DNA at the centromere is often characterized by tandemly repeated sequence. Tandem repeat monomer lengths vary between species but in many cases roughly correspond to nucleosomal units (Melters et al. 2013). These repeats, however, are not required for centromere function, as neocentromeres can form in virtually any sequence context (Scott and Sullivan 2014). Despite the divergence in DNA sequence and CENP-A organization, centromeric chromatin can in most organisms be separated into two regions: the centromere core, marked by CENP-A nucleosomes, and a pericentromeric region, usually associated with heterochromatin. Functional studies have shown that disruption of the specialized chromatin in both regions leads to defects in chromosome segregation, but the mechanistic details of the interplay between the two regions are not well understood. Here, we summarize recent progress in understanding how the centromere core and the pericentric region work together in establishing a three-dimensional structure that enables attachment to microtubules via kinetochore complex and that supports the tension generated during the segregation of chromosomes.
Characterization of the centromere core
Centromere cores are defined by the presence of CENP-A. The number of CENP-A nucleosomes varies between species from a single one that occupies ∼120 bp sequence in Saccharomyces cerevisiae to several hundred that are embedded in megabases of satellite DNA in humans (Pluta et al. 1995; Furuyama and Biggins 2007; Aldrup-Macdonald and Sullivan 2014; Bodor et al. 2014). Together with additional centromeric chromatin components, CENP-A nucleosomes create a permissive environment for centromere establishment and maintenance and are involved in the formation of three-dimensional structures that link to kinetochores and serve as capture devices for the mitotic microtubules. Recent research has advanced our understanding of how CENP-A nucleosomes are able to define the centromere core and revealed new features of the centromere core chromatin that are essential for centromere function.
CENP-A residues that specify the centromere
CENP-A is conserved among virtually all eukaryotic lineages and specifically marks the centromere. However, in contrast to other histone variants, there is little or no amino acid sequence conservation outside the C-terminal histone fold domain, and the N-terminal tails vary widely both in size and sequence (Malik and Henikoff 2003). Despite this divergence, S. cerevisiae CENP-A can complement human CENP-A, and distant plant CENP-As can complement Arabidopsis thaliana CENP-A, indicating that the functional features remain conserved (Wieland et al. 2004; Ravi et al. 2010; Maheshwari et al. 2015).
The in vivo structure and composition of CENP-A nucleosomes remain a controversial area of research that has been extensively covered in other reviews, e.g., Quénet and Dalal (2012), Henikoff and Furuyama (2012), Kurumizaka et al. (2013), Padeganeh et al. (2013), Fukagawa and Earnshaw (2014), and Steiner and Henikoff (2015). In vitro, human CENP-A is incorporated into an octameric nucleosome that is largely superimposable with the H3 nucleosome (Tachiwana et al. 2011). Comparison of the crystal structure of a human CENP-A-containing octamer with that of the H3-containing octamer revealed two major differences in the histone fold domain: CENP-A loop 1 protrudes from the CENP-A nucleosome and exposes two extra amino acid residues (Arg 80 and Gly 81), and the αN helix of CENP-A is about one turn shorter than in H3, which affects how DNA is wrapped around the CENP-A-containing nucleosome (Tachiwana et al. 2011; Roulland et al. 2016). These findings are in line with experiments using chimeras of H3 and CENP-A that identified a region covering loop 1 and α2 helix, termed CENP-A targeting domain (CATD), as the main CENP-A domain to drive centromere localization and function (Black et al. 2004; Black et al. 2007). A series of recent papers have refined our understanding of how the CATD and additional residues of CENP-A interact with other centromeric proteins to mediate centromere establishment and function (Fig. 1a).
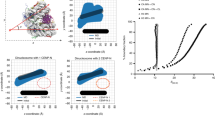
CENP-A residues that specify the centromere. a Primary sequence and schematic representation of the secondary structure of human CENP-A (top). The CATD is highlighted in purple. Regions of the CENP-A protein that interact with HJURP and different CCAN components (bottom). b Structural model of the CENP-A/H4 heterodimer in complex with HJURP (PDB ID 3R45) (Hu et al. 2011). c Structural model of the CENP-A nucleosome (PDB ID 3AN2) (Tachiwana et al. 2011). The CENP-A/H4 heterodimer on the left is shown in the same orientation as in b (framed by red dotted line), illustrating how HJURP binding prevents formation of a (CENP-A/H4)2 tetramer and association with DNA. d Structural model of CENP-C bound to the CENP-A nucleosome (CENP-C from PDB ID 4X23 modeled on CENP-A nucleosome from PDB ID 3AN2) (Tachiwana et al. 2011; Kato et al. 2013)
Human CENP-A interacts with its histone chaperone HJURP via CATD, and this interaction is essential for targeting CENP-A to the centromere (Foltz et al. 2009; Fachinetti et al. 2015). Crystal structures of human and S. cerevisiae HJURP/CENP-A/H4 heterotrimers revealed that the CATD interacts with the N-terminal part of HJURP, while the C-terminal domain of HJURP caps the DNA-binding region of the CENP-A/H4 heterodimer, preventing formation of a (CENP-A/H4)2 tetramer and premature association with DNA (Zhou et al. 2011; Hu et al. 2011; Cho and Harrison 2011) (Fig. 1b). Six surface-exposed human CENP-A CATD residues (four in S. cerevisiae CENP-A) are the main drivers of interaction, while binding of HJURP at the α1 helix restricts the conformational flexibility of the CENP-A/H4 dimer (Zhou et al. 2011; Bassett et al. 2012; Zhao et al. 2016). In addition to the CATD, interaction with HJURP also involves serine 68 in the α1 helix (Hu et al. 2011; Logsdon et al. 2015), which needs to be dephosphorylated for stable interaction (Yu et al. 2015). These findings explain how CENP-A is discriminated from H3 for targeted incorporation at the centromere. Since interaction of HJURP with CENP-A is incompatible with DNA binding, HJURP has to dissociate from the CENP-A/H4 dimer for incorporation into the nucleosome, thereby making the CATD available for interaction with other factors (Fig. 1c). In most organisms, CENP-A serves as an epigenetic mark for the inheritance of centromeres during mitosis and meiosis, and new CENP-A nucleosomes are recruited to sites of pre-existing CENP-A. For recent reviews on the regulation of CENP-A loading, refer to Nechemia-Arbely et al. (2012), Stellfox et al. (2013), Müller and Almouzni (2014), McKinley and Cheeseman (2016), Chen and Mellone (2016), and Nagpal and Fukagawa (2016).
CENP-A specifies chromatin at the centromere core, but centromere function also requires a set of proteins that form the constitutive centromere-associated network (CCAN). Once deposited, CENP-A interacts with CENP-B and a number of CCAN components, including CENP-C, CENP-N, and CENP-T, and these, in turn, interact with other CCAN components, forming a network of inner kinetochore proteins (Figs. 1 and 2a).
Three-dimensional arrangement of the centromere during mitosis. a Cartoon of kinetochore complex linking CENP-A chromatin to the microtubules. CENP-A nucleosomes coordinate the protein network of the CCAN, which recruits the outer kinetochore (KMN network) that attaches to the microtubules. b The pericentromere provides cohesion between sister chromatids and acts as a foundation for the centromere core, which assembles the kinetochore complexes for the attachment of microtubules. c Chromatin of the centromere core is folded to expose the CENP-A nucleosomes to the surface of the primary constriction. Several models for this assembly have been proposed: solenoid model (top) (Blower et al. 2002), layered boustrophedon (middle) (Ribeiro et al. 2010), and looping model (bottom) (Blower et al. 2002)
Of the CCAN components, CENP-C is most closely associated with the CENP-A nucleosome. Human CENP-C also possesses a DNA-binding motif (Sugimoto et al. 1994), but it is unclear if it binds DNA independently of the CENP-A nucleosomes. Using H3/CENP-A chimeras, it was shown that CENP-C recognizes the C-terminal part of CENP-A in humans and that just the C-terminal six amino acids of CENP-A are sufficient for CENP-C recruitment and kinetochore assembly in Xenopus laevis egg extracts (Carroll et al. 2010; Guse et al. 2011). This suggested that the CATD is mainly required for CENP-A targeting, but once it is deposited, the kinetochore is recruited via C-terminal domain and CENP-C. However, follow-up Xenopus studies showed that the CATD also has a role in CENP-C recruitment, especially at lower densities of CENP-A nucleosomes (Westhorpe et al. 2015). Experiments targeting human CENP-A/H3 chimeras to ectopic loci confirmed the need for both the C-terminus and the CATD for CENP-C recruitment and uncovered that residues within the N-terminus further enhance this interaction (Logsdon et al. 2015). Although CENP-C homologs have been identified in all model organisms, not all parts of the protein are fully conserved, and the way it binds to the CENP-A nucleosome differs between species. The central region of CENP-C is critical for this interaction in mammals, while in chicken cells, only the C-terminal part of CENP-C is required (Carroll et al. 2010; Kato et al. 2013; Nagpal et al. 2015). By NMR and crystal structure, the CENP-C central domain was shown to bind to the nucleosome surface by docking to the acidic patch (Kato et al. 2013) (Fig. 1d). These results explain the low conservation but universally hydrophobic nature of CENP-A C-termini (in contrast to H3 C-termini) across species. Binding of human CENP-C flattens and rigidifies CENP-A nucleosomes and limits the rate at which they turn over at centromeres (Falk et al. 2015; Falk et al. 2016). CENP-C interacts with CENP-H-K-I-M and CENP-L-N complexes via its central domain and is important for bridging the centromeric chromatin to the outer kinetochore, as shown in human and chicken cells (Klare et al. 2015; Nagpal et al. 2015; McKinley et al. 2015) (Fig. 2a).
The CATD of human CENP-A is also required for interaction with CENP-N, which is independent of CENP-C recruitment (Carroll et al. 2009; Logsdon et al. 2015; Fang et al. 2015). This suggests that there may be competition between CENP-C and CENP-N for binding at the CENP-A CATD, but the significance of this competition is unclear. The exposed RG loop of CENP-A plays a key role in CENP-N recruitment (Fang et al. 2015). The RG loop is concealed by centromeric chromatin compaction, and access to the RG loop by CENP-N requires decompaction of CENP-A-containing chromatin fiber, which occurs mainly at S phase.
The interaction of human CENP-A with CENP-T is mediated by a combination of binding to the N-terminal and C-terminal regions and the CATD but requires the presence of CENP-C and CENP-N and may therefore be indirect (Logsdon et al. 2015; Tachiwana et al. 2015). In Schizosaccharomyces pombe, the CENP-A N-terminus seems to be the major driver for the interaction with CENP-T (Folco et al. 2015). CENP-T particles have been mapped to the linker regions between CENP-A nucleosomes at both S. pombe and human centromeres (Thakur et al. 2015; Thakur et al. 2016).
Removal of ectopic CENP-A deposited outside of the centromere is best studied in S. cerevisiae. The E3 ubiquitin ligase Psh1 mediates degradation of ectopic CENP-A that otherwise accumulates at euchromatic loci such as promoters and can lead to segregation defects (Hewawasam et al. 2010; Ranjitkar et al. 2010; Hildebrand and Biggins 2016). Interestingly, targeting of CENP-A by Psh1 depends on the CATD, adding another function to this region of the protein. As a consequence, nucleosomal CENP-A is a poor substrate for Psh1 and requires the facilitates chromatin transcription (FACT) complex for efficient targeting (Choi et al. 2012; Deyter and Biggins 2014). FACT thus can play an important role in chromosome segregation both by ensuring the degradation of ectopically incorporated CENP-A and by facilitating the deposition of CENP-A at the centromere core (discussed below) (Chen et al. 2015; Okada et al. 2009). CENP-A ubiquitination is antagonized by the SAGA complex, and the right balance between the two activities is important, as the mitotic instabilities observed in SAGA mutants can be rescued by Psh1 deletion (Canzonetta et al. 2015).
CENP-T, CENP-W, CENP-S, CENP-X, and CENP-B contribute to centromeric chromatin
Centromeric chromatin is generally defined by the presence of CENP-A nucleosomes. However, other centromere-specific proteins may help to shape the chromatin at the centromere by binding centromeric DNA independently of CENP-A. The CCAN components CENP-T, CENP-W, CENP-S, and CENP-X have been identified independently, CENP-T and CENP-S as components of CENP-A chromatin (Foltz et al. 2006; Izuta et al. 2006), CENP-X as a subcomplex component with CENP-S (Foltz et al. 2006; Amano et al. 2009), and CENP-W as an interactor with CENP-T (Hori et al. 2008). Crystal structures revealed that CENP-T, CENP-W, CENP-S, and CENP-X are histone fold proteins that heterotetramerize, bind to, and supercoil DNA (Nishino et al. 2012). The resulting particle wraps about 100 bp of DNA, presumably in a single wrap. Both heterotetramerization and DNA binding are required for the assembly of a functional kinetochore. Interestingly, the CENP-T-W-S-X complex induces positive supercoils (Takeuchi et al. 2014), as has been shown for S. cerevisiae point centromeres (Furuyama and Henikoff 2009; Díaz-Ingelmo et al. 2015). CENP-T, CENP-W, CENP-S, and CENP-X are essential for centromere function, and CENP-T can recruit a functional centromere when tethered to DNA (Gascoigne et al. 2011; Nishino et al. 2012; Hori et al. 2013). They may thus provide an alternative mechanism to link the kinetochore to chromatin that partially bypasses CENP-A nucleosomes. How this works on a molecular level remains to be addressed, especially given that some lineages lack some or all of these proteins. While tethering of CENP-T recruits only outer kinetochore components, tethering of CENP-C, CENP-I, or HJURP recruits CENP-A and establishes a fully functional CCAN in chicken cells (Hori et al. 2013) and highlights the interdependence of CENP-A deposition and CCAN formation.
As mentioned above, DNA sequences that underlie the centromeres vary greatly between organisms, while the centromere-specific proteins are relatively well conserved. As a consequence, most centromeric proteins do not show affinity for a specific sequence. The best-known exception to this rule is CENP-B, which specifically binds to a conserved DNA sequence called CENP-B box within the satellite repeats at centromeres (Masumoto et al. 1989; Muro et al. 1992). CENP-B has been independently domesticated from pogo-like transposase in several lineages including mammals, S. pombe, and insects (Mateo and González 2014). CENP-B interacts with the N-terminal tail of CENP-A (Fachinetti et al. 2013), and its binding to CENP-B boxes in the vicinity of CENP-A stabilizes the CENP-A nucleosome on alphoid DNA (Fujita et al. 2015). Based on MNase digestion patterns, CENP-B separates two tandem CENP-A nucleosome on human alpha-satellite arrays in vivo (Henikoff et al. 2015). The functional importance of the CENP-B box and CENP-B itself is not entirely understood. There are examples of centromeres without CENP-B boxes, e.g., on the human Y-chromosome, at neocentromeres, or at all centromeres in some primates (Masumoto et al. 1989; Voullaire et al. 1993; Haaf et al. 1995; Goldberg et al. 1996), and the CENP-B protein is not essential in mice (Kapoor et al. 1998; Hudson et al. 1998; Perez-Castro et al. 1998). Moreover, S. pombe CENP-B homologs play a role in host genome surveillance for retrotransposons and replication fork progression through transposons but have no significant role in chromosome segregation (Cam et al. 2008; Zaratiegui et al. 2011). However, a recent study has shown that human CENP-B binds to and stabilizes CENP-A and CENP-C and enhances the fidelity of centromere function (Fachinetti et al. 2015). Moreover, CENP-B boxes are required for the formation of functional human artificial chromosomes (Ohzeki et al. 2002). Therefore, CENP-B may not be strictly required for centromere function but may contribute to the stability and maintenance of centromeres.
“Active” chromatin and transcription at centromere cores
Post-translational histone modifications correlate with domains of chromatin organization and can influence how chromatin is packaged. Histone marks that correlate with transcriptional activity tend to be associated with less tightly packaged chromatin than histone marks that correlate with transcriptional inactivity. Chromatin at the centromere core is characterized by H3K4me2, generally associated with active chromatin, that is interspersed with CENP-A. This contrasts with the predominant H3K9me3 found at pericentric chromatin (Sullivan and Karpen 2004; Lam et al. 2006). Centromeric H3K4me2 is functionally important, as it is required for HJURP targeting and CENP-A assembly on a synthetic human kinetochore (Bergmann et al. 2012). H2B monoubiquitination, another modification associated with actively transcribed chromatin, is needed for centromere integrity, as H2Bub depletion leads to heterochromatization of centromere cores and deficient chromosome segregation (Sadeghi et al. 2014). However, the centromere core is not homogeneously covered, and H3-rich domains associated with high H3K9me3 and low H3K4me2 levels are also observed at the centromere core (Ribeiro et al. 2010). The balance between modifications associated with active or silent chromatin seems to be important: At human artificial chromosomes, altering the chromatin state to more transcriptionally active or inactive configurations by introducing transcriptional activators or silencers results in an imbalance of H3K4me2 and H3K9me3 and leads to loss of the centromere (Nakano et al. 2008). However, there seems to be some plasticity, as inducible establishment of H3K9me3 or H3K27me marks at human artificial chromosomes causes a reduction of H3K4me2, but centromere function is maintained (Martins et al. 2016). This (temporary) tolerance for repressive marks may be important to prevent inactivation of centromeres by spreading of silent chromatin states from the pericentromere to the centromere core. One of the main contributors identified so far to antagonize spreading of heterochromatin to the centromere core is the histone acetyltransferase KAT7/HBO1/MYST2 (Ohzeki et al. 2016). It is associated with Mis18—part of the CENP-A loading machinery—at the centromere core in G1 phase and promotes nucleosome turnover, thus preventing accumulation of H3K9me3 and centromere inactivation. A finely tuned chromatin landscape is presumably required to provide the required stability and three-dimensional arrangement of centromeric chromatin during mitosis.
The importance of histone modifications associated with transcriptionally active chromatin indicated that at least some regions of the centromere are actively transcribed. Indeed, centromeric RNA polymerase II (RNA Pol II) transcripts have been reported in a variety of organisms including humans, rice, S. pombe, maize, and Drosophila. In many cases, inhibition of transcription leads to loss of centromere function (Chan et al. 2012a,b; Choi et al. 2011; Lam et al. 2006; Quénet and Dalal 2012; Rošić et al. 2014; Sadeghi et al. 2014; Saffery et al. 2003; Topp et al. 2004; Yan et al. 2006; Quénet and Dalal 2014). These studies thus uncovered an essential function of transcription for the integrity of centromeres. Centromere function is relatively tolerant to changes in levels of transcription. Chromatin with high levels of histone H3 acetylated on lysine 9 (H3K9ac) and 10-fold elevation in transcript levels had no substantial effect on kinetochore assembly or function. However, there seems to be an upper limit to the levels that are tolerated, as an increase by ∼150-fold rapidly inactivated centromere function and caused loss of CENP-A (Bergmann et al. 2012). In the tammar wallaby, hypermorphic expression of centromeric small RNAs results in disruption of CENP-A localization (Carone et al. 2013). Whether the centromere defects are a direct consequence of increased transcriptional activity or are caused by the over-abundance of centromeric transcripts remains an open question.
The fact that the DNA sequence underlying centromeres is neither necessary nor sufficient for centromere function makes it unlikely that transcripts fulfill specific sequence requirements. Indeed, transcription is also important for the function of human neocentromeres (Chueh et al. 2009). The contributions of the centromeric RNA transcripts and the process of transcription itself to the formation and the maintenance of centromeres have been difficult to clearly separate. Centromeric transcripts have been shown to be required for CENP-A loading in humans, as depletion of these transcripts leads to mitotic defects (Quénet and Dalal 2014). RNA Pol II is enriched on central domain DNA in S. pombe, but only relatively low levels of transcripts are detected, consistent with RNA Pol II stalling during transcription of centromeric DNA (Catania et al. 2015). This stalling may enable chromatin remodeling and the deposition of CENP-A nucleosomes. The chromatin remodeler FACT plays an important role in the incorporation of CENP-A (Okada et al. 2009). Mutation of FACT impairs the maintenance of H3 chromatin on transcribed regions and promotes widespread CENP-A incorporation at non-centromeric sites in S. pombe (Choi et al. 2012). FACT interacts with the CENP-A histone chaperone CAL-1 in Drosophila and is important for transcription and CENP-A incorporation at the centromere (Chen et al. 2015). Moreover, FACT interacts directly with the CCAN components CENP-T-W and is required for their incorporation at centromeres (Prendergast et al. 2016). In addition to the importance of centromere transcription and centromeric transcripts, processing of centromeric RNA, but not translation, has also been shown to play a role in centromere function (Grenfell et al. 2016).
The presence of chromatin marks associated with open chromatin and transcription likely counteracts the more compacted chromatin found at the pericentric regions and creates a permissive environment for CENP-A incorporation and kinetochore assembly. The precise role of centromeric transcripts in these protein complexes is currently unknown. While it is possible that they simply serve a structural role, it is also possible that they function in the epigenetic maintenance of centromeres via sequence complementarity to centromeric DNA.
Structure of the centromere core
The centromere in animal cells displays a layered architecture with a central core of pericentric heterochromatin and the two CENP-A-containing chromatin domains of sister chromatids peripherally attached, onto which the kinetochores build to engage with spindle microtubules (Vagnarelli and Earnshaw 2001; McIntosh et al. 2002; Guenatri et al. 2004; Vagnarelli et al. 2008) (Fig. 2b). While the CENP-A domains provide the substrates for attachment of kinetochores, the pericentromeric domain provides elasticity and resistance to tension mediated by cohesin (Gerlich et al. 2006; Ribeiro et al. 2009). The pericentromeric domain is further critical for tension sensing and signaling to the mitotic checkpoint as the chromosome passenger complex including aurora B localizes to the pericentromeric domain. For a detailed review on this topic, see van der Horst and Lens (2014). A well-studied example for this layered architecture is found in mouse, where mouse major satellite repeats are packaged into heterochromatin and form the inner centromere that provides cohesion of sister chromatids, while the minor satellite repeats assemble the CENP-A-containing core centromere and link to the kinetochore (Guenatri et al. 2004).
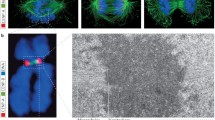
Quantitative approaches have shown that CENP-A nucleosomes make up only a fraction of core centromeres, arguing that these nucleosomes are interspersed with H3-containing nucleosomes (Bodor et al. 2014). This is also evident from work using three-dimensional deconvolution and super-resolution microscopy of stretched chromatin fibers that revealed alternating blocks of CENP-A-containing and H3-containing nucleosomes in Drosophila, chicken, and humans (Blower et al. 2002; Ribeiro et al. 2010). S. pombe regional centromeres deviate from this pattern, as their core regions are devoid of H3 nucleosomes (Thakur et al. 2015).
To identify the three-dimensional arrangement of the CENP-A-containing and H3-containing chromatin, a number of groups have analyzed mitotic chromosomes by electron microscopy. CENP-A domains were found to be localized at the surface of the primary constriction and occupy roughly two thirds of the constriction length, one third of the constriction height, and one third of the constriction width, both at human alphoid and neocentromeres (Marshall et al. 2008). A similar pattern is found at barley, wheat, and spelt chromosomes, where CENP-A is restricted to the surface of the primary constriction (Wanner et al. 2015). Interestingly, Wanner et al. also reported that microtubules do attach not only to the primary constriction but also to the chromosomal areas flanking it. In the holocentric plant Luzula elegans, metaphase chromosomes form a groove of chromatin to which CENP-A almost exclusively localizes (Heckmann et al. 2011; Wanner et al. 2015; Schubert et al. 2016). In pea chromosomes, the primary constrictions are unusually elongated, exhibit three to five explicit CENP-A-containing regions, and can thus be seen as an intergrade between regional and holocentromeres. Correlative fluorescence and scanning electron microscopy showed that the CENP-A within these domains is also oriented towards the surface, as is seen in organisms with smaller primary constrictions, but interrupted by areas without visible CENP-A (Neumann et al. 2012).
In summary, these studies confirm that CENP-A-containing chromatin occupies only a limited fraction of the centromeric chromatin and reveal that the CENP-A-containing chromatin is mainly exposed towards the surface of the chromosome. How chromatin is folded so that chromosomes form a constriction and that CENP-A nucleosomes are exposed on the surface in a back-to-back orientation during mitosis remains subject to research. Several models have been proposed (Fig. 2c): The chromatin could be coiled into cylindrical structures (solenoid model) or organized in parallel loops (looping model), exposing CENP-A to the outside (Blower et al. 2002). Alternatively, the chromatin could be folded as a layered boustrophedon, with planar sinusoids containing interspersed CENP-A-rich and H3-rich subdomains oriented towards the outer kinetochore (Ribeiro et al. 2010). The coiled centromeric chromatin may be folded or looped back upon itself multiple times in the length dimension to form a multilayered structure (Marshall et al. 2008). Regardless of which model turns out to be closest to reality, the exposed CENP-A nucleosomes coordinate the CCAN protein network described above, which together with the KNL-1/Mis12 complex/Ndc80 complex (KMN) network of the outer kinetochore forms the platform to which microtubules attach (Fig. 2a). Details on the kinetochore structures are described in recent reviews (Pesenti et al. 2016; Nagpal and Fukagawa 2016).
Imaging higher order chromatin to derive folding patterns seems currently beyond the limit of microscopy. Chromosome capture methods could alternatively be used to probe centromeric chromatin of mitotic chromosomes (Naumova et al. 2013). However, this approach will only be feasible once the sequences at satellite centromeres are better assembled, so that interactions can be mapped to the assembly.
Characterization of pericentromeric chromatin
Centromeres can be divided into centromere core and pericentromeric regions, where the core regions provide the attachment site for the kinetochore and the pericentromere is responsible for sister chromatid cohesion. The major task of centromeres is the capture of microtubules from both spindle poles and to establish bi-orientation of chromatids during mitosis or of bivalents during meiosis. Once attached, centromeres provide the load-bearing point for the spindle forces that act on chromosomes. The mechanical resistance is provided by pericentric chromatin and the ring-like molecules cohesin and condensin that act as tethers for chromatin loops. The tension generated by proper bi-orientation serves as a signal for the spindle assembly checkpoint for control of the cell cycle. Once mitosis progresses to anaphase, the mechanical resistance is rapidly dissipated by proteolytic cleavage of cohesin through the protease separase, thus releasing chromatids to migrate to separate poles. Here, we will review the current knowledge of the chromatin structure at the pericentromere and its relationship to cohesin and condensin.
Pericentromeric chromatin stabilizes cohesin
Cohesion of sister chromatids is mediated by the stabilization of the cohesin complex on pericentromeric chromatin (Nasmyth and Haering 2009). Cohesin consists of the two structural maintenance of chromatin (SMC) subunits Smc1 and Smc3, both of them folded back onto themselves to form a long coiled coil with N-terminal and C-terminal ABC-type ATPase domains at one end and the central hinge domain at the other (Fig. 3a). Smc1 and Smc3 heterodimerize through the hinge domains, thereby forming long, V-shaped heterodimers. The open ends of the V are bridged by the kleisin subunit Scc1 and the Scc3 subunit to form a ring that can trap two strands of chromatin (Gruber et al. 2003; Haering et al. 2008). During G1 and S phases, cohesin is mobile and is dynamically loaded by the cohesin loader Scc2/Scc4 and released from chromosomes by Wapl/Rad61 (Chan et al. 2012b; Lopez-Serra et al. 2013). During DNA replication, cohesin becomes stabilized through acetylation of the Smc3 subunit by the Eco1 acetyltransferase, which blocks Wapl from triggering the opening of the cohesin ring (Marston 2014).
Cohesin shapes the centromere. a Model of the S. cerevisiae cohesin molecule. b Schematic representation of two S. cerevisiae sister chromosomes attached to the mitotic spindle showing how cohesin shapes the centromeres in yeast mitosis. Cohesin connects sister chromatids at chromosome arms but also links together centromeric chromatin, both by intrachromosome (blue) and interchromosome (purple) strand trapping, thereby forming a barrel-shaped structure underneath the kinetochore
While stabilization of cohesin during replication links together the entire arms of the sister chromatids, pericentromeres have additional mechanisms to preferentially enrich cohesin. In S. cerevisiae, a 25–50 kb region around the kinetochore attachment point is highly enriched in cohesin (Blat and Kleckner 1999; Megee et al. 1999; Tanaka et al. 1999) and corresponds to the pericentromere that, in contrast to regional centromeres, lacks typical heterochromatin. Scc2/Scc4 interacts with the kinetochore (Fernius and Marston 2009; Ng et al. 2009) and loads cohesin in the core region from where it distributes into the surrounding pericentromere (Hu et al. 2015).
In contrast to S. cerevisiae, S. pombe has extensive pericentric heterochromatin similar to animal and plant centromeres. The S. pombe heterochromatin system is extremely well characterized and relies on silencing by both the chromatin and RNA interference (RNAi) machinery to control transcription, recombination, and structural aspects of chromosomes at centromeres, telomeres, and the silent mating-type loci (Grewal and Jia 2007). The RNAi pathway degrades transcripts in cis, while histone deacetylation and histone H3K9 methylation established by chromatin-modifying machinery result in transcriptional gene silencing (Alper et al. 2012; Holoch and Moazed 2015). H3K9 is methylated by a single histone methyltransferase, Clr4, which provides a binding site for the heterochromatin protein 1 (HP1) homologs Swi6 and Chp2. Heterochromatin formation is required for cohesin enrichment at S. pombe centromeres, as Swi6 provides the platform for cohesin binding (Bernard et al. 2001; Nonaka et al. 2002), and phosphorylation of Swi6 regulates its effect on sister chromatid cohesion (Bailis et al. 2003). The cohesin subunit Psc3 (Scc3 in S. cerevisiae and STAG1/2/3 in humans) interacts with Swi6, and localization of Psc3 through fusion of two chromodomains restores cohesion in the absence of Swi6 (Yamagishi et al. 2008).
Cohesin on pericentromeres is protected from turnover by the recruitment of shugoshin proteins, thus ensuring sister chromatid cohesion (Kitajima et al. 2004; Katis et al. 2004; Kiburz et al. 2005). Shugoshin’s protective function is mediated by the recruitment of the PP2A phosphatase that prevents phosphorylation of the Scc1 subunit of cohesin and thereby keeps separase cleavage at a low level until anaphase (Salic et al. 2004; Kitajima et al. 2006; Riedel et al. 2006). While S. pombe shugoshin is required exclusively for cohesion of homologous chromosomes in meiosis I, in S. cerevisiae and animals, shugoshin plays a critical role in protecting cohesion during mitosis.
Suv39h and HP1 influence the cohesion of the pericentromere, as loss of H3K9me and HP1 binding lead to increased separation of major satellites during metaphase. However, some groups have reported that the HP1-Suv39 axis does not affect the localization of cohesin to the pericentromere (Koch et al. 2008; Serrano et al. 2009), while more recent reports document a significant contribution of the Suv39-HP1 system to cohesin recruitment, mediated by the H4K20 methyltransferase Suv4-20h2 (Shimura et al. 2011; Hahn et al. 2013). In mammalian cells, cohesin recruitment and stabilization at centromeres during mitosis are more complex, likely because shugoshin-based and HP1-based pathways intermingle, and seem to depend on tissue type and developmental stages.
The structure and function of pericentromeric chromatin in yeasts
The centromere in S. cerevisiae has been studied extensively in order to understand the properties of chromatin at the centromere. The 16 chromosomes have well-defined centromeres that are accessible to sequencing methods since they are non-repetitive. Upon bi-orientation during metaphase, the 32 centromeres of the replicated chromosomes cluster into two foci (Goshima and Yanagida 2000) and connect to 16 microtubules from each side (Winey et al. 1995). While the chromosome arms remain together, chromatids need to dissociate in a ∼10 kb region around the centromere core resulting in the separation of the kinetochores by ∼1 μm (Goshima and Yanagida 2000; He et al. 2000; Tanaka et al. 2000). In order to reconcile the separation of sister kinetochores with cohesin enrichment, Bloom and colleagues proposed that in contrast to the intermolecular tethering that holds together chromatids in the chromosome arms, cohesin at the centromere is engaged in intramolecular trapping of pericentromeric chromatin to produce a loop with the kinetochore at its tip (Verdaasdonk and Bloom 2011; Stephens et al. 2011; Stephens et al. 2013), thereby giving the mitotic chromosome a cruciform structure (Fig. 3b).
The kinetochores and the intervening chromatin structure act against the spindle forces in an elastic manner. One component contributing elasticity is chromatin itself (Bouck and Bloom 2007), which cooperates with condensin and cohesin to establish the spring-like bi-oriented kinetochore structure (Stephens et al. 2011). GFP-tagged cohesin observed in live cells during mitosis was found to form a barrel that extends between the kinetochores (Yeh et al. 2008). This cohesin barrel overlaps with pericentromeric chromatin and encircles the spindle microtubules. In contrast, condensin is found running down the center of the barrel in close proximity to the spindle microtubules (Stephens et al. 2011).
In S. pombe, a picture is emerging where pericentromeric chromatin interacts extensively among arms of the same chromosome and among arms of different chromosomes. Recent data from chromosome conformation capture (Duan et al. 2010; Burton et al. 2014; Mizuguchi et al. 2014) show typical “X” patterns off the diagonal that mark centromere-centromere interactions. This prototypical pattern can be used to infer the centromere locations in yeast species for which centromeres have not been mapped previously (Varoquaux et al. 2015). In S. pombe, the proximity of centromeres is only mildly reduced in cohesin mutants but almost completely lost in mutants deficient in pericentric heterochromatin formation (Mizuguchi et al. 2014). This indicates that heterochromatin plays a dominant role in promoting the interaction between centromeres in this system.
Taken together, these data suggest that yeast pericentromeres form an intrachromosomal and interchromosomal meshwork at centromeres that is held together by cohesin and heterochromatin proteins (Fig. 3b). This meshwork is important for the sister chromatid cohesion and the establishment of a tension-resisting elastic structure that is required for proper chromosome segregation.
Conclusions
The three-dimensional organization of the centromeric chromatin is a critical factor for a functional chromosome segregation machinery. It is based on the specialized structural features of the CENP-A nucleosomes that allow assembly of the kinetochore and is evident in the arrangement of the core centromere and pericentromere domains in the context of segregating chromosomes that present the spindle attachment points and monitor their correct engagement and timely resolution of cohesion. Recent progress has started to elucidate many of the details of how the kinetochore interacts with the CENP-A nucleosome. Chromosome conformation capture and microscopy are revealing the higher order structures that centromeres adopt and show that in yeast, centromeres cluster together in interphase and also during mitosis. How the centromere chromatin is folded in detail to establish a tension-resistant structure and how the spatial arrangement into inner and outer centromeric chromatin is established remain open questions. In this review, we have synthesized the wealth of new insight that has been obtained in recent years, revealing an intricately balanced system of transcription, chromatin variants, and modifications that tightly cooperates with the structural and regulatory machinery of the kinetochore in order to guarantee faithful and robust segregation of the genomic material.
References
Aldrup-Macdonald ME, Sullivan BA (2014) The past, present, and future of human centromere genomics. Genes 5:33–50
Alper BJ, Lowe BR, Partridge JF (2012) Centromeric heterochromatin assembly in fission yeast—balancing transcription, RNA interference and chromatin modification. Chromosom Res 20:521–534
Amano M, Suzuki A, Hori T et al (2009) The CENP-S complex is essential for the stable assembly of outer kinetochore structure. J Cell Biol 186:173–182
Bailis JM, Bernard P, Antonelli R et al (2003) Hsk1-Dfp1 is required for heterochromatin-mediated cohesion at centromeres. Nat Cell Biol 5:1111–1116
Bassett EA, DeNizio J, Barnhart-Dailey MC et al (2012) HJURP uses distinct CENP-A surfaces to recognize and to stabilize CENP-A/histone H4 for centromere assembly. Dev Cell 22:749–762
Bergmann JH, Jakubsche JN, Martins NM et al (2012) Epigenetic engineering: histone H3K9 acetylation is compatible with kinetochore structure and function. J Cell Sci 125:411–421
Bernard P, Maure JF, Partridge JF et al (2001) Requirement of heterochromatin for cohesion at centromeres. Science 294:2539–2542
Black BE, Foltz DR, Chakravarthy S et al (2004) Structural determinants for generating centromeric chromatin. Nature 430:578–582
Black BE, Jansen LET, Maddox PS et al (2007) Centromere identity maintained by nucleosomes assembled with histone H3 containing the CENP-A targeting domain. Mol Cell 25:309–322
Blat Y, Kleckner N (1999) Cohesins bind to preferential sites along yeast chromosome III, with differential regulation along arms versus the centric region. Cell 98:249–259
Blower MD, Sullivan BA, Karpen GH (2002) Conserved organization of centromeric chromatin in flies and humans. Dev Cell 2:319–330
Bodor DL, Mata JF, Sergeev M et al (2014) The quantitative architecture of centromeric chromatin. Elife 3:e02137
Bouck DC, Bloom K (2007) Pericentric chromatin is an elastic component of the mitotic spindle. Curr Biol 17:741–748
Brinkley BR, Stubblefield E (1966) The fine structure of the kinetochore of a mammalian cell in vitro. Chromosoma 19:28–43
Burton JN, Liachko I, Dunham MJ, Shendure J (2014) Species-level deconvolution of metagenome assemblies with Hi-C-based contact probability maps. G3 4:1339–1346
Cam HP, Noma K-I, Ebina H et al (2008) Host genome surveillance for retrotransposons by transposon-derived proteins. Nature 451:431–436
Canzonetta C, Vernarecci S, Iuliani M, et al. (2015) SAGA DUB-Ubp8 deubiquitylates centromeric histone variant Cse4. G3 6:287–298
Carone DM, Zhang C, Hall LE et al (2013) Hypermorphic expression of centromeric retroelement-encoded small RNAs impairs CENP-A loading. Chromosom Res 21:49–62
Carroll CW, Milks KJ, Straight AF (2010) Dual recognition of CENP-A nucleosomes is required for centromere assembly. J Cell Biol 189:1143–1155
Carroll CW, Silva MCC, Godek KM et al (2009) Centromere assembly requires the direct recognition of CENP-A nucleosomes by CENP-N. Nat Cell Biol 11:896–902
Catania S, Pidoux AL, Allshire RC (2015) Sequence features and transcriptional stalling within centromere DNA promote establishment of CENP-A chromatin. PLoS Genet 11:e1004986
Chan FL, Marshall OJ, Saffery R et al (2012a) Active transcription and essential role of RNA polymerase II at the centromere during mitosis. Proc Natl Acad Sci U S A 109:1979–1984
Chan K-L, Roig MB, Hu B et al (2012b) Cohesin’s DNA exit gate is distinct from its entrance gate and is regulated by acetylation. Cell 150:961–974
Chen C-C, Bowers S, Lipinszki Z et al (2015) Establishment of centromeric chromatin by the CENP-A assembly factor CAL1 requires FACT-mediated transcription. Dev Cell 34:73–84
Chen C-C, Mellone BG (2016) Chromatin assembly: journey to the CENter of the chromosome. J Cell Biol 214:13–24
Choi ES, Strålfors A, Castillo AG et al (2011) Identification of noncoding transcripts from within CENP-A chromatin at fission yeast centromeres. J Biol Chem 286:23600–23607
Choi ES, Strålfors A, Catania S et al (2012) Factors that promote H3 chromatin integrity during transcription prevent promiscuous deposition of CENP-A(Cnp1) in fission yeast. PLoS Genet 8:e1002985
Cho U-S, Harrison SC (2011) Recognition of the centromere-specific histone Cse4 by the chaperone Scm3. Proc Natl Acad Sci U S A 108:9367–9371
Chueh AC, Northrop EL, Brettingham-Moore KH et al (2009) LINE retrotransposon RNA is an essential structural and functional epigenetic component of a core neocentromeric chromatin. PLoS Genet 5:e1000354
Deyter GMR, Biggins S (2014) The FACT complex interacts with the E3 ubiquitin ligase Psh1 to prevent ectopic localization of CENP-A. Genes Dev 28:1815–1826
Díaz-Ingelmo O, Martínez-García B, Segura J et al (2015) DNA topology and global architecture of point centromeres. Cell Rep 13:667–677
Duan Z, Andronescu M, Schutz K et al (2010) A three-dimensional model of the yeast genome. Nature 465:363–367
Earnshaw WC, Allshire RC, Black BE et al (2013) Esperanto for histones: CENP-A, not CenH3, is the centromeric histone H3 variant. Chromosom Res 21:101–106
Earnshaw WC, Rothfield N (1985) Identification of a family of human centromere proteins using autoimmune sera from patients with scleroderma. Chromosoma 91:313–321
Fachinetti D, Folco HD, Nechemia-Arbely Y et al (2013) A two-step mechanism for epigenetic specification of centromere identity and function. Nat Cell Biol 15:1056–1066
Fachinetti D, Han JS, McMahon MA et al (2015) DNA sequence-specific binding of CENP-B enhances the Fidelity of human centromere function. Dev Cell 33:314–327
Falk SJ, Guo LY, Sekulic N et al (2015) Chromosomes. CENP-C reshapes and stabilizes CENP-A nucleosomes at the centromere Science 348:699–703
Falk SJ, Lee J, Sekulic N et al (2016) CENP-C directs a structural transition of CENP-A nucleosomes mainly through sliding of DNA gyres. Nat Struct Mol Biol 23:204–208
Fang J, Liu Y, Wei Y et al (2015) Structural transitions of centromeric chromatin regulate the cell cycle-dependent recruitment of CENP-N. Genes Dev 29:1058–1073
Fernius J, Marston AL (2009) Establishment of cohesion at the pericentromere by the Ctf19 kinetochore subcomplex and the replication fork-associated factor, Csm3. PLoS Genet 5:e1000629
Flemming W (1882) Zellsubstanz, Kern und Zelltheilung. F. C. W, Vogel
Folco HD, Campbell CS, May KM et al (2015) The CENP-A N-tail confers epigenetic stability to centromeres via the CENP-T branch of the CCAN in fission yeast. Curr Biol 25:348–356
Foltz DR, Jansen LET, Bailey AO et al (2009) Centromere-specific assembly of CENP-a nucleosomes is mediated by HJURP. Cell 137:472–484
Foltz DR, Jansen LET, Black BE et al (2006) The human CENP-A centromeric nucleosome-associated complex. Nat Cell Biol 8:458–469
Fujita R, Otake K, Arimura Y et al (2015) Stable complex formation of CENP-B with the CENP-A nucleosome. Nucleic Acids Res 43:4909–4922
Fukagawa T, Earnshaw WC (2014) The centromere: chromatin foundation for the kinetochore machinery. Dev Cell 30:496–508
Furuyama S, Biggins S (2007) Centromere identity is specified by a single centromeric nucleosome in budding yeast. Proc Natl Acad Sci U S A 104:14706–14711
Furuyama T, Henikoff S (2009) Centromeric nucleosomes induce positive DNA supercoils. Cell 138:104–113
Gascoigne KE, Takeuchi K, Suzuki A et al (2011) Induced ectopic kinetochore assembly bypasses the requirement for CENP-A nucleosomes. Cell 145:410–422
Gerlich D, Hirota T, Koch B et al (2006) Condensin I stabilizes chromosomes mechanically through a dynamic interaction in live cells. Curr Biol 16:333–344
Goldberg IG, Sawhney H, Pluta AF et al (1996) Surprising deficiency of CENP-B binding sites in African green monkey alpha-satellite DNA: implications for CENP-B function at centromeres. Mol Cell Biol 16:5156–5168
Goshima G, Yanagida M (2000) Establishing biorientation occurs with precocious separation of the sister kinetochores, but not the arms, in the early spindle of budding yeast. Cell 100:619–633
Grenfell AW, Heald R, Strzelecka M (2016) Mitotic noncoding RNA processing promotes kinetochore and spindle assembly in Xenopus. J Cell Biol. doi:10.1083/jcb.201604029
Grewal SIS, Jia S (2007) Heterochromatin revisited. Nat Rev Genet 8:35–46
Gruber S, Haering CH, Nasmyth K (2003) Chromosomal cohesin forms a ring. Cell 112:765–777
Guenatri M, Bailly D, Maison C, Almouzni G (2004) Mouse centric and pericentric satellite repeats form distinct functional heterochromatin. J Cell Biol 166:493–505
Guse A, Carroll CW, Moree B et al (2011) In vitro centromere and kinetochore assembly on defined chromatin templates. Nature 477:354–358
Haaf T, Thomas H, Gregory Mater A et al (1995) Presence and abundance of CENP-B box sequences in great ape subsets of primate-specific alpha satellite DNA. J Mol Evol 41:487–491
Haering CH, Farcas A-M, Arumugam P et al (2008) The cohesin ring concatenates sister DNA molecules. Nature 454:297–301
Hahn M, Dambacher S, Dulev S et al (2013) Suv4-20h2 mediates chromatin compaction and is important for cohesin recruitment to heterochromatin. Genes Dev 27:859–872
Heckmann S, Schroeder-Reiter E, Kumke K et al (2011) Holocentric chromosomes of Luzula elegans are characterized by a longitudinal centromere groove, chromosome bending, and a terminal nucleolus organizer region. Cytogenet Genome Res 134:220–228
Henikoff JG, Thakur J, Kasinathan S, Henikoff S (2015) A unique chromatin complex occupies young α-satellitearrays of human centromeres. Sci Adv. doi:10.1126/sciadv.1400234
Henikoff S, Furuyama T (2012) The unconventional structure of centromeric nucleosomes. Chromosoma 121:341–352
Hewawasam G, Shivaraju M, Mattingly M et al (2010) Psh1 is an E3 ubiquitin ligase that targets the centromeric histone variant Cse4. Mol Cell 40:444–454
He X, Asthana S, Sorger PK (2000) Transient sister chromatid separation and elastic deformation of chromosomes during mitosis in budding yeast. Cell 101:763–775
Hildebrand EM, Biggins S (2016) Regulation of budding yeast CENP-A levels prevents misincorporation at promoter nucleosomes and transcriptional defects. PLoS Genet 12:e1005930
Holoch D, Moazed D (2015) RNA-mediated epigenetic regulation of gene expression. Nat Rev Genet 16:71–84
Hori T, Amano M, Suzuki A et al (2008) CCAN makes multiple contacts with centromeric DNA to provide distinct pathways to the outer kinetochore. Cell 135:1039–1052
Hori T, Shang W-H, Takeuchi K, Fukagawa T (2013) The CCAN recruits CENP-A to the centromere and forms the structural core for kinetochore assembly. J Cell Biol 200:45–60
Hu B, Petela N, Kurze A et al (2015) Biological chromodynamics: a general method for measuring protein occupancy across the genome by calibrating ChIP-seq. Nucleic Acids Res 43:e132
Hudson DF, Fowler KJ, Earle E et al (1998) Centromere protein B null mice are mitotically and meiotically normal but have lower body and testis weights. J Cell Biol 141:309–319
Hu H, Liu Y, Wang M et al (2011) Structure of a CENP-A-histone H4 heterodimer in complex with chaperone HJURP. Genes Dev 25:901–906
Izuta H, Ikeno M, Suzuki N et al (2006) Comprehensive analysis of the ICEN (interphase centromere complex) components enriched in the CENP-A chromatin of human cells. Genes Cells 11:673–684
Jokelainen PT (1967) The ultrastructure and spatial organization of the metaphase kinetochore in mitotic rat cells. J Ultrastruct Res 19:19–44
Kapoor M, Mini K, de Oca Luna RM et al (1998) The cenpB gene is not essential in mice. Chromosoma 107:570–576
Katis VL, Galova M, Rabitsch KP et al (2004) Maintenance of cohesin at centromeres after meiosis I in budding yeast requires a kinetochore-associated protein related to MEI-S332. Curr Biol 14:560–572
Kato H, Jiang J, Zhou B-R et al (2013) A conserved mechanism for centromeric nucleosome recognition by centromere protein CENP-C. Science 340:1110–1113
Kiburz BM, Reynolds DB, Megee PC et al (2005) The core centromere and Sgo1 establish a 50-kb cohesin-protected domain around centromeres during meiosis I. Genes Dev 19:3017–3030
Kitajima TS, Kawashima SA, Watanabe Y (2004) The conserved kinetochore protein shugoshin protects centromeric cohesion during meiosis. Nature 427:510–517
Kitajima TS, Sakuno T, Ishiguro K-I et al (2006) Shugoshin collaborates with protein phosphatase 2A to protect cohesin. Nature 441:46–52
Klare K, Weir JR, Basilico F et al (2015) CENP-C is a blueprint for constitutive centromere-associated network assembly within human kinetochores. J Cell Biol 210:11–22
Koch B, Kueng S, Ruckenbauer C et al (2008) The Suv39h-HP1 histone methylation pathway is dispensable for enrichment and protection of cohesin at centromeres in mammalian cells. Chromosoma 117:199–210
Kurumizaka H, Horikoshi N, Tachiwana H, Kagawa W (2013) Current progress on structural studies of nucleosomes containing histone H3 variants. Curr Opin Struct Biol 23:109–115
Lam AL, Boivin CD, Bonney CF et al (2006) Human centromeric chromatin is a dynamic chromosomal domain that can spread over noncentromeric DNA. Proc Natl Acad Sci U S A 103:4186–4191
Logsdon GA, Barrey EJ, Bassett EA et al (2015) Both tails and the centromere targeting domain of CENP-A are required for centromere establishment. J Cell Biol 208:521–531
Lopez-Serra L, Lengronne A, Borges V et al (2013) Budding yeast Wapl controls sister chromatid cohesion maintenance and chromosome condensation. Curr Biol 23:64–69
Luykx P (1965) The structure of the kinetochore in meiosis and mitosis in Urechis eggs. Exp Cell Res 39:643–657
Maheshwari S, Tan EH, West A et al (2015) Naturally occurring differences in CENH3 affect chromosome segregation in zygotic mitosis of hybrids. PLoS Genet 11:e1004970
Malik HS, Henikoff S (2003) Phylogenomics of the nucleosome. Nat Struct Biol 10:882–891
Marshall OJ, Marshall AT, Choo KHA (2008) Three-dimensional localization of CENP-A suggests a complex higher order structure of centromeric chromatin. J Cell Biol 183:1193–1202
Marston AL (2014) Chromosome segregation in budding yeast: sister chromatid cohesion and related mechanisms. Genetics 196:31–63
Martins NMC, Bergmann JH, Shono N et al (2016) Epigenetic engineering shows that a human centromere resists silencing mediated by H3K27me3/K9me3. Mol Biol Cell 27:177–196
Masumoto H, Masukata H, Muro Y et al (1989) A human centromere antigen (CENP-B) interacts with a short specific sequence in alphoid DNA, a human centromeric satellite. J Cell Biol 109:1963–1973
Mateo L, González J (2014) Pogo-like transposases have been repeatedly domesticated into CENP-B-related proteins. Genome Biol Evol 6:2008–2016
McIntosh JR, Grishchuk EL, West RR (2002) Chromosome-microtubule interactions during mitosis. Annu Rev Cell Dev Biol 18:193–219
McKinley KL, Cheeseman IM (2016) The molecular basis for centromere identity and function. Nat Rev Mol Cell Biol 17:16–29
McKinley KL, Sekulic N, Guo LY et al (2015) The CENP-L-N complex forms a critical node in an integrated meshwork of interactions at the centromere-kinetochore interface. Mol Cell 60:886–898
Megee PC, Mistrot C, Guacci V, Koshland D (1999) The centromeric sister chromatid cohesion site directs Mcd1p binding to adjacent sequences. Mol Cell 4:445–450
Melters DP, Bradnam KR, Young HA et al (2013) Comparative analysis of tandem repeats from hundreds of species reveals unique insights into centromere evolution. Genome Biol 14:R10
Mizuguchi T, Fudenberg G, Mehta S et al (2014) Cohesin-dependent globules and heterochromatin shape 3D genome architecture in S. pombe. Nature 516:432–435
Müller S, Almouzni G (2014) A network of players in H3 histone variant deposition and maintenance at centromeres. Biochim Biophys Acta 1839:241–250
Muro Y, Masumoto H, Yoda K et al (1992) Centromere protein B assembles human centromeric alpha-satellite DNA at the 17-bp sequence, CENP-B box. J Cell Biol 116:585–596
Nagpal H, Fukagawa T (2016) Kinetochore assembly and function through the cell cycle. Chromosoma. doi:10.1007/s00412-016-0608-3
Nagpal H, Hori T, Furukawa A et al (2015) Dynamic changes in CCAN organization through CENP-C during cell-cycle progression. Mol Biol Cell 26:3768–3776
Nakano M, Cardinale S, Noskov VN et al (2008) Inactivation of a human kinetochore by specific targeting of chromatin modifiers. Dev Cell 14:507–522
Nasmyth K, Haering CH (2009) Cohesin: its roles and mechanisms. Annu Rev Genet 43:525–558
Naumova N, Imakaev M, Fudenberg G et al (2013) Organization of the mitotic chromosome. Science 342:948–953
Nechemia-Arbely Y, Fachinetti D, Cleveland DW (2012) Replicating centromeric chromatin: spatial and temporal control of CENP-A assembly. Exp Cell Res 318:1353–1360
Neumann P, Navrátilová A, Schroeder-Reiter E et al (2012) Stretching the rules: monocentric chromosomes with multiple centromere domains. PLoS Genet 8:e1002777
Ng TM, Waples WG, Lavoie BD, Biggins S (2009) Pericentromeric sister chromatid cohesion promotes kinetochore biorientation. Mol Biol Cell 20:3818–3827
Nishino T, Takeuchi K, Gascoigne KE et al (2012) CENP-T-W-S-X forms a unique centromeric chromatin structure with a histone-like fold. Cell 148:487–501
Nonaka N, Kitajima T, Yokobayashi S et al (2002) Recruitment of cohesin to heterochromatic regions by Swi6/HP1 in fission yeast. Nat Cell Biol 4:89–93
Ohzeki J-I, Nakano M, Okada T, Masumoto H (2002) CENP-B box is required for de novo centromere chromatin assembly on human alphoid DNA. J Cell Biol 159:765–775
Ohzeki J-I, Shono N, Otake K et al (2016) KAT7/HBO1/MYST2 regulates CENP-A chromatin assembly by antagonizing Suv39h1-mediated centromere inactivation. Dev Cell 37:413–427
Okada M, Okawa K, Isobe T, Fukagawa T (2009) CENP-H-containing complex facilitates centromere deposition of CENP-A in cooperation with FACT and CHD1. Mol Biol Cell 20:3986–3995
Padeganeh A, De Rop V, Maddox PS (2013) Nucleosomal composition at the centromere: a numbers game. Chromosom Res 21:27–36
Palmer DK, O’Day K, Wener MH et al (1987) A 17-kD centromere protein (CENP-A) copurifies with nucleosome core particles and with histones. J Cell Biol 104:805–815
Perez-Castro AV, Shamanski FL, Meneses JJ et al (1998) Centromeric protein B null mice are viable with no apparent abnormalities. Dev Biol 201:135–143
Pesenti ME, Weir JR, Musacchio A (2016) Progress in the structural and functional characterization of kinetochores. Curr Opin Struct Biol 37:152–163
Pluta AF, Mackay AM, Ainsztein AM et al (1995) The centromere: hub of chromosomal activities. Science 270:1591–1594
Prendergast L, Lisa P, Sebastian M et al (2016) The CENP-T/-W complex is a binding partner of the histone chaperone FACT. Genes Dev 30:1313–1326
Quénet D, Dalal Y (2012) The CENP-A nucleosome: a dynamic structure and role at the centromere. Chromosom Res 20:465–479
Quénet D, Dalal Y (2014) A long non-coding RNA is required for targeting centromeric protein A to the human centromere. Elife 3:e03254
Ranjitkar P, Press MO, Yi X et al (2010) An E3 ubiquitin ligase prevents ectopic localization of the centromeric histone H3 variant via the centromere targeting domain. Mol Cell 40:455–464
Ravi M, Kwong PN, Menorca RMG et al (2010) The rapidly evolving centromere-specific histone has stringent functional requirements in Arabidopsis thaliana. Genetics 186:461–471
Ribeiro SA, Gatlin JC, Dong Y et al (2009) Condensin regulates the stiffness of vertebrate centromeres. Mol Biol Cell 20:2371–2380
Ribeiro SA, Vagnarelli P, Dong Y et al (2010) A super-resolution map of the vertebrate kinetochore. Proc Natl Acad Sci U S A 107:10484–10489
Riedel CG, Katis VL, Katou Y et al (2006) Protein phosphatase 2A protects centromeric sister chromatid cohesion during meiosis I. Nature 441:53–61
Rošić S, Köhler F, Erhardt S (2014) Repetitive centromeric satellite RNA is essential for kinetochore formation and cell division. J Cell Biol 207:335–349
Roulland Y, Ouararhni K, Naidenov M et al (2016) The flexible ends of CENP-A nucleosome are required for mitotic fidelity. Mol Cell 63(4):674–685
Sadeghi L, Siggens L, Svensson JP, Ekwall K (2014) Centromeric histone H2B monoubiquitination promotes noncoding transcription and chromatin integrity. Nat Struct Mol Biol 21:236–243
Saffery R, Sumer H, Hassan S et al (2003) Transcription within a functional human centromere. Mol Cell 12:509–516
Salic A, Waters JC, Mitchison TJ (2004) Vertebrate shugoshin links sister centromere cohesion and kinetochore microtubule stability in mitosis. Cell 118:567–578
Schubert V, Zelkowski M, Klemme S, Houben A (2016) Similar sister chromatid arrangement in mono- and holocentric plant chromosomes. Cytogenet Genome Res. doi:10.1159/000447681
Scott KC, Sullivan BA (2014) Neocentromeres: a place for everything and everything in its place. Trends Genet 30:66–74
Serrano A, Rodríguez-Corsino M, Losada A (2009) Heterochromatin protein 1 (HP1) proteins do not drive pericentromeric cohesin enrichment in human cells. PLoS One 4:e5118
Shimura M, Toyoda Y, Iijima K et al (2011) Epigenetic displacement of HP1 from heterochromatin by HIV-1 Vpr causes premature sister chromatid separation. J Cell Biol 194:721–735
Steiner FA, Henikoff S (2015) Diversity in the organization of centromeric chromatin. Curr Opin Genet Dev 31:28–35
Stellfox ME, Bailey AO, Foltz DR (2013) Putting CENP-A in its place. Cell Mol Life Sci 70:387–406
Stephens AD, Haase J, Vicci L et al (2011) Cohesin, condensin, and the intramolecular centromere loop together generate the mitotic chromatin spring. J Cell Biol 193:1167–1180
Stephens AD, Snider CE, Haase J et al (2013) Individual pericentromeres display coordinated motion and stretching in the yeast spindle. J Cell Biol 203:407–416
Sugimoto K, Yata H, Muro Y, Himeno M (1994) Human centromere protein C (CENP-C) is a DNA-binding protein which possesses a novel DNA-binding motif. J Biochem 116:877–881
Sullivan BA, Karpen GH (2004) Centromeric chromatin exhibits a histone modification pattern that is distinct from both euchromatin and heterochromatin. Nat Struct Mol Biol 11:1076–1083
Tachiwana H, Kagawa W, Shiga T et al (2011) Crystal structure of the human centromeric nucleosome containing CENP-A. Nature 476:232–235
Tachiwana H, Müller S, Blümer J et al (2015) HJURP involvement in de novo CenH3(CENP-A) and CENP-C recruitment. Cell Rep 11:22–32
Takeuchi K, Nishino T, Mayanagi K et al (2014) The centromeric nucleosome-like CENP-T-W-S-X complex induces positive supercoils into DNA. Nucleic Acids Res 42:1644–1655
Talbert PB, Ahmad K, Almouzni G et al (2012) A unified phylogeny-based nomenclature for histone variants. Epigenetics Chromatin 5:7
Talbert PB, Henikoff S (2013) Phylogeny as the basis for naming histones. Trends Genet 29:499–500
Tanaka T, Cosma MP, Wirth K, Nasmyth K (1999) Identification of cohesin association sites at centromeres and along chromosome arms. Cell 98:847–858
Tanaka T, Fuchs J, Loidl J, Nasmyth K (2000) Cohesin ensures bipolar attachment of microtubules to sister centromeres and resists their precocious separation. Nat Cell Biol 2:492–499
Thakur J, Jitendra T, Steven H (2016) CENPT bridges adjacent CENPA nucleosomes on young human α-satellite dimers. Genome Res gr.204784.116
Thakur J, Talbert PB, Henikoff S (2015) Inner kinetochore protein interactions with regional centromeres of fission yeast. Genetics 201:543–561
Topp CN, Zhong CX, Dawe RK (2004) Centromere-encoded RNAs are integral components of the maize kinetochore. Proc Natl Acad Sci U S A 101:15986–15991
van der Horst A, Lens SMA (2014) Cell division: control of the chromosomal passenger complex in time and space. Chromosoma 123:25–42
Vagnarelli PB, Earnshaw WC (2001) INCENP loss from an inactive centromere correlates with the loss of sister chromatid cohesion. Chromosoma 110:393–401
Vagnarelli P, Ribeiro SA, Earnshaw WC (2008) Centromeres: old tales and new tools. FEBS Lett 582:1950–1959
Varoquaux N, Liachko I, Ay F et al (2015) Accurate identification of centromere locations in yeast genomes using Hi-C. Nucleic Acids Res 43:5331–5339
Verdaasdonk JS, Bloom K (2011) Centromeres: unique chromatin structures that drive chromosome segregation. Nat Rev Mol Cell Biol 12:320–332
Voullaire LE, Slater HR, Petrovic V, Choo KH (1993) A functional marker centromere with no detectable alpha-satellite, satellite III, or CENP-B protein: activation of a latent centromere? Am J Hum Genet 52:1153–1163
Wanner G, Schroeder-Reiter E, Ma W et al (2015) The ultrastructure of mono- and holocentric plant centromeres: an immunological investigation by structured illumination microscopy and scanning electron microscopy. Chromosoma 124:503–517
Westhorpe FG, Fuller CJ, Straight AF (2015) A cell-free CENP-A assembly system defines the chromatin requirements for centromere maintenance. J Cell Biol 209:789–801
Wieland G, Orthaus S, Ohndorf S et al (2004) Functional complementation of human centromere protein a (CENP-A) by Cse4p from Saccharomyces cerevisiae. Mol Cell Biol 24:6620–6630
Winey M, Mamay CL, O’Toole ET et al (1995) Three-dimensional ultrastructural analysis of the Saccharomyces cerevisiae mitotic spindle. J Cell Biol 129:1601–1615
Yamagishi Y, Sakuno T, Shimura M, Watanabe Y (2008) Heterochromatin links to centromeric protection by recruiting shugoshin. Nature 455:251–255
Yan H, Ito H, Nobuta K et al (2006) Genomic and genetic characterization of rice Cen3 reveals extensive transcription and evolutionary implications of a complex centromere. Plant Cell 18:2123–2133
Yeh E, Haase J, Paliulis LV et al (2008) Pericentric chromatin is organized into an intramolecular loop in mitosis. Curr Biol 18:81–90
Yu Z, Zhou X, Wang W et al (2015) Dynamic phosphorylation of CENP-A at Ser68 orchestrates its cell-cycle-dependent deposition at centromeres. Dev Cell 32:68–81
Zaratiegui M, Vaughn MW, Irvine DV et al (2011) CENP-B preserves genome integrity at replication forks paused by retrotransposon LTR. Nature 469:112–115
Zhao H, Winogradoff D, Bui M et al (2016) Promiscuous histone mis-assembly is actively prevented by chaperones. J Am Chem Soc 138:13207–13218
Zhou Z, Feng H, Zhou B-R et al (2011) Structural basis for recognition of centromere histone variant CenH3 by the chaperone Scm3. Nature 472:234–237
Acknowledgements
The Schalch and Steiner labs are supported by the Swiss National Science Foundation (Grant Nos. 31003A_156774 to F.S. and PP00P3_139137 and PP00P3_163760 to T.S.) and the Republic and Canton of Geneva.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
This article does not contain any studies with human participants or animals performed by any of the authors.
Conflict of interest
The authors declare that they have no conflict of interest.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Schalch, T., Steiner, F.A. Structure of centromere chromatin: from nucleosome to chromosomal architecture. Chromosoma 126, 443–455 (2017). https://doi.org/10.1007/s00412-016-0620-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00412-016-0620-7