Abstract
Purpose
K-ras mutations predict resistance against epidermal growth factor receptor (EGFR)-directed therapy of metastatic colorectal cancer (CRC). The purpose of this study was to analyze the distribution of K-ras mutations in primary tumors and corresponding sentinel lymph nodes (SLNs) from colon cancer patients.
Methods
Tumor biopsies and SLNs from 158 patients with non-metastatic colon cancer were analyzed for K-ras mutations in codons 12 and 13 by a sensitive and quantitative peptide nucleic acid clamp PCR assay.
Results
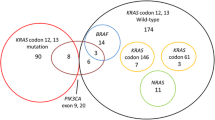
Analyses of single fresh-frozen tumor biopsies revealed K-ras mutations in 67 (42%) of the patients. Apparently low levels of K-ras mutations in 13 of the mutated primary tumors and the presence of K-ras mutations in SLNs from seven patients with a wild-type primary tumor biopsy suggested possible intratumoral heterogeneity for 20 of the patients. To confirm this hypothesis, we analyzed tissue sections from all available formalin-fixed, paraffin-embedded (FFPE) tumor blocks from these 20 patients. Ten of the patients had a mixture of tissue sections positive and tissue sections negative for K-ras mutations, two patients had K-ras mutations in all sections, and eight patients had no detectable K-ras mutations in tumor FFPE tissue blocks. Among these eight patients, five had K-ras mutations detected in SLNs. Thus, evidence supporting a heterogeneous distribution of K-ras mutations was obtained for 15 patients.
Conclusions
Heterogeneous distribution of K-ras codon 12 and 13 mutations within primary tumor, or between primary tumor and lymph node metastases, was demonstrated for 15 (20%) of 74 colon cancer patients having K-ras mutations. This may have implications for tissue sampling routines with regard to EGFR-directed therapy of CRC, both in adjuvant and metastatic settings.




Similar content being viewed by others
References
Jemal A, Siegel R, Ward E, Hao Y, Xu J, Thun MJ (2009) Cancer statistics, 2009. CA Cancer J Clin 59:225–249
Jiang Y, Kimchi ET, Staveley-O’Carroll KF, Cheng H, Ajani JA (2009) Assessment of K-ras mutation: a step toward personalized medicine for patients with colorectal cancer. Cancer 115:3609–3617
Allegra CJ, Jessup JM, Somerfield MR, Hamilton SR, Hammond EH, Hayes DF, McAllister PK, Morton RF, Schilsky RL (2009) American Society of Clinical Oncology provisional clinical opinion: testing for KRAS gene mutations in patients with metastatic colorectal carcinoma to predict response to anti-epidermal growth factor receptor monoclonal antibody therapy. J Clin Oncol 27:2091–2096
Amado RG, Wolf M, Peeters M, Van Cutsem E, Siena S, Freeman DJ, Juan T, Sikorski R, Suggs S, Radinsky R, Patterson SD, Chang DD (2008) Wild-type KRAS is required for panitumumab efficacy in patients with metastatic colorectal cancer. J Clin Oncol 26:1626–1634
Karapetis CS, Khambata-Ford S, Jonker DJ, O’Callaghan CJ, Tu D, Tebbutt NC, Simes RJ, Chalchal H, Shapiro JD, Robitaille S, Price TJ, Shepherd L, Au HJ, Langer C, Moore MJ, Zalcberg JR (2008) K-ras mutations and benefit from cetuximab in advanced colorectal cancer. N Engl J Med 359:1757–1765
Siena S, Sartore-Bianchi A, Di Nicolantonio F, Balfour J, Bardelli A (2009) Biomarkers predicting clinical outcome of epidermal growth factor receptor-targeted therapy in metastatic colorectal cancer. J Natl Cancer Inst 101:1308–1324
Andreyev HJ, Norman AR, Cunningham D, Oates JR, Clarke PA (1998) Kirsten ras mutations in patients with colorectal cancer: the multicenter “RASCAL” study. J Natl Cancer Inst 90:675–684
Vogelstein B, Fearon ER, Hamilton SR, Kern SE, Preisinger AC, Leppert M, Nakamura Y, White R, Smits AM, Bos JL (1988) Genetic alterations during colorectal-tumor development. N Engl J Med 319:525–532
Fearon ER, Vogelstein B (1990) A genetic model for colorectal tumorigenesis. Cell 61:759–767
Losi L, Baisse B, Bouzourene H, Benhattar J (2005) Evolution of intratumoral genetic heterogeneity during colorectal cancer progression. Carcinogenesis 26:916–922
Baisse B, Bouzourene H, Saraga EP, Bosman FT, Benhattar J (2001) Intratumor genetic heterogeneity in advanced human colorectal adenocarcinoma. Int J Cancer 93:346–352
Giaretti W, Monaco R, Pujic N, Rapallo A, Nigro S, Geido E (1996) Intratumor heterogeneity of K-ras2 mutations in colorectal adenocarcinomas: association with degree of DNA aneuploidy. Am J Pathol 149:237–245
Ishii M, Sugai T, Habano W, Nakamura S (2004) Analysis of Ki-ras gene mutations within the same tumor using a single tumor crypt in colorectal carcinomas. J Gastroenterol 39:544–549
Fukunari H, Iwama T, Sugihara K, Miyaki M (2003) Intratumoral heterogeneity of genetic changes in primary colorectal carcinomas with metastasis. Surg Today 33:408–413
Al-Mulla F, Going JJ, Sowden ET, Winter A, Pickford IR, Birnie GD (1998) Heterogeneity of mutant versus wild-type Ki-ras in primary and metastatic colorectal carcinomas, and association of codon-12 valine with early mortality. J Pathol 185:130–138
Baldus SE, Schaefer KL, Engers R, Hartleb D, Stoecklein NH, Gabbert HE (2010) Prevalence and heterogeneity of KRAS, BRAF, and PIK3CA mutations in primary colorectal adenocarcinomas and their corresponding metastases. Clin Cancer Res 16:790–799
Shibata D, Schaeffer J, Li ZH, Capella G, Perucho M (1993) Genetic heterogeneity of the c-K-ras locus in colorectal adenomas but not in adenocarcinomas. J Natl Cancer Inst 85:1058–1063
Dix BR, Robbins PD, Spagnolo DV, Padovan GL, House AK, Iacopetta BJ (1995) Clonal analysis of colorectal tumors using K-ras and p53 gene mutations as markers. Diagn Mol Pathol 4:261–265
Thiede C, Bayerdorffer E, Blasczyk R, Wittig B, Neubauer A (1996) Simple and sensitive detection of mutations in the ras proto-oncogenes using PNA-mediated PCR clamping. Nucleic Acids Res 24:983–984
Taback B, Bilchik AJ, Saha S, Nakayama T, Wiese DA, Turner RR, Kuo CT, Hoon DS (2004) Peptide nucleic acid clamp PCR: a novel K-ras mutation detection assay for colorectal cancer micrometastases in lymph nodes. Int J Cancer 111:409–414
Gilje B, Heikkilä R, Oltedal S, Tjensvoll K, Nordgård O (2008) High-fidelity DNA polymerase enhances the sensitivity of a peptide nucleic acid clamp PCR assay for K-ras mutations. J Mol Diagn 10:325–331
Oltedal S, Gilje B, Kørner H, Aasprong OG, Tjensvoll K, Heikkila R, Smaaland R, Nordgard O (2010) Detection of occult metastases in sentinel lymph nodes from colon cancer patients by K-ras mutation peptide nucleic acid clamp PCR. Ann Surg 251:1087–1091
Nordgård O, Oltedal S, Kørner H, Aasprong OG, Tjensvoll K, Gilje B, Heikkilä R (2009) Quantitative RT-PCR detection of tumor cells in sentinel lymph nodes isolated from colon cancer patients with an ex vivo approach. Ann Surg 249:602–607
Tortola S, Steinert R, Hantschick M, Peinado MA, Gastinger I, Stosiek P, Lippert H, Schlegel W, Reymond MA (2001) Discordance between K-ras mutations in bone marrow micrometastases and the primary tumor in colorectal cancer. J Clin Oncol 19:2837–2843
Zauber P, Sabbath-Solitare M, Marotta SP, Bishop DT (2003) Molecular changes in the Ki-ras and APC genes in primary colorectal carcinoma and synchronous metastases compared with the findings in accompanying adenomas. Mol Pathol 56:137–140
Santini D, Loupakis F, Vincenzi B, Floriani I, Stasi I, Canestrari E, Rulli E, Maltese PE, Andreoni F, Masi G, Graziano F, Baldi GG, Salvatore L, Russo A, Perrone G, Tommasino MR, Magnani M, Falcone A, Tonini G, Ruzzo A (2008) High concordance of KRAS status between primary colorectal tumors and related metastatic sites: implications for clinical practice. Oncologist 13:1270–1275
Italiano A, Hostein I, Soubeyran I, Fabas T, Benchimol D, Evrard S, Gugenheim J, Becouarn Y, Brunet R, Fonck M, Francois E, Saint-Paul MC, Pedeutour F (2010) KRAS and BRAF mutational status in primary colorectal tumors and related metastatic sites: biological and clinical implications. Ann Surg Oncol 17:1429–1434
Al-Mulla F, Milner-White EJ, Going JJ, Birnie GD (1999) Structural differences between valine-12 and aspartate-12 Ras proteins may modify carcinoma aggression. J Pathol 187:433–438
Seeburg PH, Colby WW, Capon DJ, Goeddel DV, Levinson AD (1984) Biological properties of human c-Ha-ras1 genes mutated at codon 12. Nature 312:71–75
Visvader JE, Lindeman GJ (2008) Cancer stem cells in solid tumours: accumulating evidence and unresolved questions. Nat Rev Cancer 8:755–768
Weinberg RA (2006) The biology of cancer. Garland science. Taylor and Francis, New York
Artale S, Sartore-Bianchi A, Veronese SM, Gambi V, Sarnataro CS, Gambacorta M, Lauricella C, Siena S (2008) Mutations of KRAS and BRAF in primary and matched metastatic sites of colorectal cancer. J Clin Oncol 26:4217–4219
Mancuso A, Sollami R, Recine F, Cerbone L, Macciomei MC, Leone A (2010) Patient with colorectal cancer with heterogeneous KRAS molecular status responding to cetuximab-based chemotherapy. J Clin Oncol 28:e756–e758
Bouchahda M, Karaboue A, Saffroy R, Innominato P, Gorden L, Guettier C, Adam R, Levi F (2010) Acquired KRAS mutations during progression of colorectal cancer metastases: possible implications for therapy and prognosis. Cancer Chemother Pharmacol 66:605–609
Maheswaran S, Sequist LV, Nagrath S, Ulkus L, Brannigan B, Collura CV, Inserra E, Diederichs S, Iafrate AJ, Bell DW, Digumarthy S, Muzikansky A, Irimia D, Settleman J, Tompkins RG, Lynch TJ, Toner M, Haber DA (2008) Detection of mutations in EGFR in circulating lung-cancer cells. N Engl J Med 359:366–377
Acknowledgments
We thank Anne Elin Varhaugvik for technical assistance. The study was supported by the Western Norway Regional Health Authorities, the Norwegian Cancer Society, the Folke Hermansen Fund, and Stavanger University Hospital.
Conflict of interest
The authors have no conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Oltedal, S., Aasprong, O.G., Møller, J.H. et al. Heterogeneous distribution of K-ras mutations in primary colon carcinomas: implications for EGFR-directed therapy. Int J Colorectal Dis 26, 1271–1277 (2011). https://doi.org/10.1007/s00384-011-1233-5
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00384-011-1233-5