Abstract
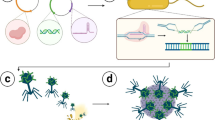
We developed a genetic approach to efficiently add an affinity tag to every copy of protein IX (pIX) of M13 filamentous bacteriophage in a population. Affinity-tagged phages can be immobilized on a surface in a uniform monolayer in order to position the pIII–displayed peptides or proteins for optimal interaction with ligands. The tagging consists of two major steps. First, gene IX (gIX) of M13 phage is mutated in Escherichia coli via genetic recombineering with the gIX::aacCI insertion allele. Second, a plasmid that co-produces the affinity-tagged pIX and native pVIII is transformed into the strain carrying the defective M13 gIX. This genetic complementation allows the formation of infective phage particles that carry a full complement (five copies per virion) of the affinity-tagged pIX. To demonstrate the efficacy of our method, we tagged a M13 derivative phage, M13KE, with Strep-tag II. In order to tag pIX with Strep-tag II, the phage genes for pIX and pVIII were cloned and expressed from pASG-IBA4 which contains the E. coli OmpA signal sequence and Strep-Tag II under control of the tetracycline promoter/operator system. We achieved the maximum phage production of 3 × 1011 pfu/ml when Strep-Tag II-pIX-pVIII fusion was induced with 10 ng/ml of anhydrotetracycline. The complete process of affinity tagging a phage probe takes less than 5 days and can be utilized to tag any M13 or fd pIII-displayed oligopeptide probes to improve their performance.




Similar content being viewed by others
References
Agrawal S, Kulabhusan PK, Joshi M, Bodas D, Paknikar KM (2016) A high affinity phage-displayed peptide as a recognition probe for the detection of Salmonella Typhimurium. J Biotechnol 231:40–45. https://doi.org/10.1016/j.jbiotec.2016.05.027
Ausubel FM, Brent R, Kingston RE, Moore DD, Seidman JG, Smith JA, Struhl K (eds)(1993) Curr Prot Mol Biol. Greene Publishing Associates, Inc. and John Wiley & Sons, Inc.
Bertani G (1951) Studies on lysogenesis. I The mode of phage liberation by lysogenic Escherichia coli. J Bacteriol 62(3):293–300
Bishop-Hurley SL, Rea PJ, McSweeney CS (2010) Phage-displayed peptides selected for binding to Campylobacter jejuni are antimicrobial. Protein Eng Des Sel 23(10):751–757. https://doi.org/10.1093/protein/gzq050
Brigati J, Williams DD, Sorokulova IB, Nanduri V, Chen IH, Turnbough CL Jr, Petrenko VA (2004) Diagnostic probes for Bacillus anthracis spores selected from a landscape phage library. Clin Chem 50(10):1899–1906
Chai Y, Li S, Horikawa S, Park MK, Vodyanoy V, Chin BA (2012) Rapid and sensitive detection of Salmonella Typhimurium on eggshells by using wireless biosensors. J Food Prot 75(4):631–636. https://doi.org/10.4315/0362-028x.jfp-11-339
Chan S-W, Bye JM, Jackson P, Allain J-P (1996) Human recombinant antibodies specific for hepatitis C virus core and envelope E2 peptides from an immune phage display library. J Gen Virol 77(10):2531–2539. https://doi.org/10.1099/0022-1317-77-10-2531
Chung WJ, Lee DY, Yoo SY (2014) Chemical modulation of M13 bacteriophage and its functional opportunities for nanomedicine. Int J Nanomedicine 9:5825–5836. https://doi.org/10.2147/IJN.S73883
Datsenko KA, Wanner BL (2000) One-step inactivation of chromosomal genes in Escherichia coli K-12 using PCR products. Proc Natl Acad Sci U S A 97(12):6640–6645
Deng X, Wang L, You X, Dai P, Zeng Y (2018) Advances in the T7 phage display system (review). Mol Med Rep 17(1):714–720. https://doi.org/10.3892/mmr.2017.7994
Hanahan D (1983) Studies on transformation of Escherichia coli with plasmids. J Mol Biol 166(4):557–580
Houbiers MC, Spruijt RB, Demel RA, Hemminga MA, Wolfs CJ (2001) Spontaneous insertion of gene 9 minor coat protein of bacteriophage M13 in model membranes. Biochim Biophys Acta 1511(2):309–316
Huang S, Li SQ, Yang H, Johnson ML, Wan J, Chen I, Petrenko VA, Barbaree JM, Chin BA (2008) Optimization of phage-based magnetoelastic biosensor performance. Sensor Trans J on Microsyst: Technol Appl 3:87–96
Huang S, Yang H, Lakshmanan RS, Johnson ML, Wan J, Chen IH, Wikle HC 3rd, Petrenko VA, Barbaree JM, Chin BA (2009) Sequential detection of Salmonella typhimurium and Bacillus anthracis spores using magnetoelastic biosensors. Biosens Bioelectron 24(6):1730–1736. https://doi.org/10.1016/j.bios.2008.09.006
Lakshmanan RS, Guntupalli R, Hu J, Kim DJ, Petrenko VA, Barbaree JM, Chin BA (2007) Phage immobilized magnetoelastic sensor for the detection of Salmonella typhimurium. J Microbiol Methods 71(1):55–60. https://doi.org/10.1016/j.mimet.2007.07.012
Li SQ, Johnson ML, Wan J, Petrenko VA, Chin BA (2008) Microfabricated magnetoelastic biosensors for the detection of Bacillus anthracis spores. ECS Trans 16:177–185
Loset GA, Roos N, Bogen B, Sandlie I (2011) Expanding the versatility of phage display II: improved affinity selection of folded domains on protein VII and IX of the filamentous phage. PLoS One 6(2):e17433. https://doi.org/10.1371/journal.pone.0017433
Lu X, Weiss P, Block T (2004) A phage with high affinity for hepatitis B surface antigen for the detection of HBsAg. J Virol Methods 119(1):51–54
Martinez E, Bartolome B, de la Cruz F (1988) pACYC184-derived cloning vectors containing the multiple cloning site and lacZ alpha reporter gene of pUC8/9 and pUC18/19 plasmids. Gene 68(1):159–162
Mechaly A, Zahavy E, Fisher M (2008) Development and implementation of a single-chain Fv antibody for specific detection of Bacillus anthracis spores. Appl Environ Microbiol 74(3):818–822. https://doi.org/10.1128/aem.01244-07
Mimmi S, Maisano D, Quinto I, Iaccino E (2019) Phage display: an overview in context to drug discovery. Trends Pharmacol Sci 40(2):87–91. https://doi.org/10.1016/j.tips.2018.12.005
Moon JS, Choi EJ, Jeong NN, Sohn JR, Han DW, Oh JW (2019) Research progress of M13 bacteriophage-based biosensors. Nanomaterials (Basel) 9(10). https://doi.org/10.3390/nano9101448
Muzard J, Platt M, Lee GU (2012) M13 bacteriophage-activated superparamagnetic beads for affinity separation. Small 8(15):2403–2411. https://doi.org/10.1002/smll.201200099
Nam KT, Peelle BR, Lee SW, Belcher AM (2004) Genetically driven assembly of nanorings based on the M13 virus. Nano Lett 4(1):23–27. https://doi.org/10.1021/nl0347536
Nanduri V, Bhunia AK, Tu S-I, Paoli GC, Brewster JD (2007) SPR biosensor for the detection of L. monocytogenes using phage-displayed antibody. Biosens Bioelectron 23(2):248–252
Niyomdecha S, Limbut W, Numnuam A, Kanatharana P, Charlermroj R, Karoonuthaisiri N, Thavarungkul P (2018) Phage-based capacitive biosensor for Salmonella detection. Talanta 188:658–664. https://doi.org/10.1016/j.talanta.2018.06.033
Olsen EV, Sorokulova IB, Petrenko VA, Chen IH, Barbaree JM, Vodyanoy VJ (2006) Affinity-selected filamentous bacteriophage as a probe for acoustic wave biodetectors of Salmonella typhimurium. Biosens Bioelectron 21(8):1434–1442
Park M-K, Wikle HC, Chai Y, Horikawa S, Shen W, Chin BA (2012) The effect of incubation time for Salmonella Typhimurium binding to phage-based magnetoelastic biosensors. Food Control 26:539–545
Park M-K, Li S, Chin BA (2013a) Detection of Salmonella typhimurium grown directly on tomato surface using phage-based magnetoelastic biosensors. Food Bioprocess Technol 6:682–689
Park M-K, Park JH, Wikle HC 3rd, Chin BA (2013b) Evaluation of phage-based magnetoelastic biosensors for direct detection of Salmonella Typhimurium on spinach leaves. Sensors Actuators B Chem 176:1134–1140
Petrenko VA (2008) Landscape phage as a molecular recognition interface for detection devices. Microelectron J 39(2):202–207. https://doi.org/10.1016/j.mejo.2006.11.007
Petrenko VA, Smith GP, Gong X, Quinn T (1996) A library of organic landscapes on filamentous phage. Protein Eng 9:797–801
Rahbarnia L, Farajnia S, Babaei H, Majidi J, Veisi K, Ahmadzadeh V, Akbari B (2017) Evolution of phage display technology: from discovery to application. J Drug Target 25(3):216–224. https://doi.org/10.1080/1061186X.2016.1258570
Ramji S, Lakshmanan RS, Guntupalli R, Petrenko VA, Chin BA (2007a) Phage based magnetoelastic sensor for the detection of Salmonella typhimurium. J Microbiol Methods 71(1):55–60
Ramji S, Lakshmanan RS, Hu J, Petrenko VA, Barbaree JM, Chin BA (2007b) Detection of Salmonella typhimurium in fat free milk using a phage immobilized magnetoelastic sensor. Sensors Actuators B 126(2):544–550
Sawada T, Serizawa T (2018) Filamentous viruses as building blocks for hierarchical self-assembly toward functional soft materials. Bull Chem Soc Jpn 91(3):455–466. https://doi.org/10.1246/bcsj.20170428
Schmidt TG, Koepke J, Frank R, Skerra A (1996) Molecular interaction between the strep-tag affinity peptide and its cognate target, streptavidin. J Mol Biol 255(5):753–766. https://doi.org/10.1006/jmbi.1996.0061
Schweizer HP (1993) Small broad-host-range gentamycin resistance gene cassettes for site-specific insertion and deletion mutagenesis. Biotechniques 15(5):831–834
Shen W, Lakshmanan RS, Mathison LC, Petrenko VA, Chin BA (2009) Phage coated magnetoelastic micro-biosensors for real-time detection of Bacillus anthracis spores. Sensors Actuators B Chem 137:501–506
Smith GP (1985) Filamentous phage fusion: novel expression vectors that display cloned antigens on the surface of the viron. Science 228:1315–1317
Sorokulova IB, Olsen EV, Chen IH, Fiebor B, Barbaree JM, Vodyanoy VJ, Chin BA, Petrenko VA (2005) Landscape phage probes for Salmonella typhimurium. J Microbiol Methods 63(1):55–72
Tan WS, Tan GH, Yusoff K, Seow HF (2005) A phage-displayed cyclic peptide that interacts tightly with the immunodominant region of hepatitis B surface antigen. J Clin Virol 34(1):35–41
Tan Y, Tian T, Liu W, Zhu Z, C JY (2016) Advance in phage display technology for bioanalysis. Biotechnol J 11(6):732–745. https://doi.org/10.1002/biot.201500458
Turnbough CL Jr (2003) Discovery of phage display peptide ligands for species-specific detection of Bacillus spores. J Microbiol Methods 53(2):263–271
Wan J, Shu H, Huang S, Fiebor B, Chen IH, Petrenko VA, Chin BA (2007a) Phage-based magnetoelastic wireless biosensors for detecting Bacillus anthracis spores. IEEE Sensors J 7(3):470–477
Wan JH, Johnson ML, Guntupalli VA, Petrenko VA, Chin BA (2007b) Detection of Bacillus anthracis spores in liquid using phage-based magnetoelastic micro-resonators. Sensors Actuators B Chem 127(2):559–566
Wu CH, Liu IJ, Lu RM, Wu HC (2016) Advancement and applications of peptide phage display technology in biomedical science. J Biomed Sci 23:8. https://doi.org/10.1186/s12929-016-0223-x
Funding
This research was partially supported by the United States Department of Agriculture National Institute of Food and Agriculture grant USDA-2016-67,021-25,005 and Alabama Agricultural Experimental Station Hatch grant ALA0SUH to Suh and Chin. Zhou Tong was supported by the Chinese Scholarship Council fellowship. Anwar Kalalah was supported by a scholarship from the Kingdom of Saudi Arabia.
Author information
Authors and Affiliations
Contributions
Zhou Tong, Laura Silo-Suh, Anwar Kalalah, and Paul Dawson performed the experiments. Bryan Chin and Sang-Jin Suh conceived and designed the study. Laura Silo-Suh and Sang-Jin Suh analyzed the data and supervised the project. Zhou Tong wrote the initial draft of the paper, and Laura Silo-Suh and Sang-Jin Suh revised the manuscript. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Tong, Z., Silo-Suh, L.A., Kalalah, A. et al. Efficient affinity-tagging of M13 phage capsid protein IX for immobilization of protein III-displayed oligopeptide probes on abiotic platforms. Appl Microbiol Biotechnol 104, 1201–1209 (2020). https://doi.org/10.1007/s00253-019-10338-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-019-10338-8