Abstract
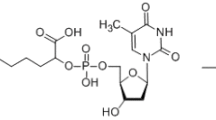
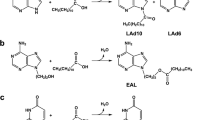
The hammerhead ribozyme (HHR) is one of smallest catalytic RNAs, composed of a catalytic core and three stems; it undergoes self-cleavage in the presence of divalent magnesium ions (Mg2+) or other cations. It is hypothesized that the function and metabolism of RNAs might be regulated via interaction with lipid membranes in the prebiotic world. Using synthetic RNAs that model the core fragment of hammerhead ribozyme-like assembly (HHR-a), we investigated the enhancement of the self-cleavage reaction of HHR-a induced by the liposomes, both in the absence and presence of Mg2+. The HHR-a activity was enhanced by 1,2-dioleoyl-sn-glycero-3-phosphoethanolamine (DOPE)/1,2-dioleoyl-sn-glycero-3-phosphocholine (DPPC) = 8/2 liposome with Mg2+, while other liposomes did not so significant. In the presence of DOPE/DPPC = 8/2 liposome, the HHR-a activity was observed without Mg2+, revealed by the conformational change of the HHR inhibitor complex induced by the interaction with the liposome. The UV resonance Raman spectroscopy analysis investigated the interaction between lipid molecules and nucleobases, suggesting that the ethanolamine group of DOPE molecules are assumed to act as monovalent cations alternative to Mg2+, depending on the liposome membrane characteristics.
Graphical abstract





Similar content being viewed by others
Abbreviations
- HHR:
-
Hammerhead ribozyme
- HHR-a:
-
Hammerhead ribozyme-like RNA assembly
- HHR–IC:
-
HHR–inhibitor complex
- MALDI TOF/MS:
-
Matrix-assisted laser desorption/ionization time-of-flight/mass spectrometry
- CD:
-
Circular dichroism
- UVRR:
-
UV resonance Raman
- DOPE:
-
1,2-Dioleoyl-sn-glycero-3-phosphoethanolamine
- DPPE:
-
1,2-Dipalmitoyl-sn-glycero-3-phosphoethanolamine
- DOPC:
-
1,2-Dioleoyl-sn-glycero-3-phosphocholine
- DPPC:
-
1,2-Dipalmitoyl-sn-glycero-3-phosphocholine
- DPH:
-
1,6-Diphenyl-1,3,5-hexatriene
- Laurdan:
-
6-Lauroyl-2-dimethylaminonaphthalene
- TNS:
-
2-(p-Toluidino)naphthalene-6-sulfonate
- SGI:
-
SYBR Green I
References
Ausländer S, Ketzer P, Hartig JS (2010) A ligand-dependent hammerhead ribozyme switch for controlling mammalian gene expression. Mol BioSyst 6:807–814
Bassi GS, Murchie AIH, Lilley DMJ (1996) The ion-induced folding of the hammerhead ribozyme: core sequence changes that perturb folding into the active conformation. RNA 2:756–768
Billinghurst BE, Oladepo SA, Loppnow GR (2009) pH-Dependent UV resonance Raman spectra of cytosine and uracil. J Phys Chem B 113:7392–7397
Boots JL, Canny MD, Azimi E, Pardi A (2008) Metal ion specificities for folding and cleavage activity in the Schistosoma hammerhead ribozyme. RNA 14:2212–2222
Carmona P, Rodriguez-Casado A, Molina M (1999) Conformational structure and bindingmode of glyceraldehydes-3-phosphate dehydrogenase to tRNA studied by Raman and CD spectroscopy. Biochim Biophys Acta 1432:222–233
Chen IA (2015) Replicating towards complexity. Nat Chem 7:191–192
Chen IA, Salehi-Ashtiani K, Szostak JW (2005) RNA catalysis in model protocell vesicles. J Am Chem Soc 127:13213–13219
Curtis EA, Bartel DP (2001) The hammerhead cleavage reaction in monovalent cations. RNA 7:546–552
De Almeida RFM, Loura LMS, Fedorov A, Prieto M (2005) Lipid rafts have different sizes depending on membrane composition: a time-resolved fluorescence resonance energy transfer study. J Mol Biol 346:1109–1120
De la Peña M, Gago S, Flores R (2003) Peripheral regions of natural hammerhead ribozymes greatly increase their self-cleavage activity. EMBO J 22:5561–5570
Ferrari D, Peracchi A (2002) A continuous kinetic assay for RNA-cleaving deoxyribozymes, exploiting ethidium bromide as an extrinsic fluorescent probe. Nucleic Acid Res 30:e112
Florián J, Baumruk V, Leszczyński J (1996) IR and Raman spectra, tautomeric stabilities, and scaled quantum mechanical force fields of protonated cytosine. J Phys Chem 100:5578–5589
Hammann C, Lilley DMJ (2002) Folding and activity of the hammerhead ribozyme. ChemBioChem 3:690–700
Janas T, Janas T, Yarus M (2006) Specific RNA binding to ordered phospholipid bilayers. Nucleic Acid Res 34:2128–2136
Khvorova A, Lescoute A, Westhof E, Jayasena SD (2003) Sequence elements outside the hammerhead ribozyme catalytic core enable intracellular activity. Nat Struct Biol 10:708–712
Lanir A, Yu NT (1979) A Raman spectroscopic study of the interaction of divalent metal ions with adenine moiety of adenosine 5′-triphosphate. J Biol Chem 254:5882–5887
Lilley DMJ (2005) Structure, folding and mechanisms of ribozymes. Curr Opin Struct Biol 15:313–323
Liu Y, Nagle JF (2004) Diffuse scattering provides material parameters and electron density profiles of biomembranes. Phys Rev E 69:1–4. article id:040901
MacDonald RC, MacDonald RI, Menco BP, Takeshita K, Subbarao NK, Hu LR (1991) Small-volume extrusion apparatus for preparation of large, unilamellar vesicles. Biochim Biophys Acta 1061:297–303
Marrink SJ, De Vries AH, Mark AE (2004) Coarse grained model for semiquantitative lipid simulations. J Phys Chem B 108:750–760
Mathlouthi M, Seuvre AM, Koenig JL (1983) FT-IR and laser-Raman spectra of d-ribose and 2-deoxy-d-erythro-pentose (“2-deoxy-d-ribose”). Carbohyd Res 122:31–47
Müller UF, Bartel DP (2008) Improved polymerase ribozyme efficiency on hydrophobic assemblies. RNA 14:552–562
Nagatomo S, Nagai M, Kitagawa T (2011) A new way to understand quaternary structure changes of hemoglobin upon ligand binding on the basis of UV-resonance Raman evaluation of intersubunit interactions. J Am Chem Soc 133:10101–10110
Pley HW, Flaherty KM, McKay DB (1994) Three-dimensional structure of a hammerhead ribozyme. Nature 372:68–74
Ricardo A, Szostak JW (2009) Origin of life on earth. Sci Am 301:54–61
Riepe A, Beier H, Gross HJ (1999) Enhancement of RNA self-cleavage by micellar catalysis. FEBS Lett 457:193–199
Schneider CA, Rasband WS, Eliceiri KW (2012) NIH Image to ImageJ: 25 years of image analysis. Nat Method 9:671–675
Singh JS (2012) FTIR and Raman spectra and fundamental frequencies of 5-halosubstituted uracils: 5-X-uracil (X = F, Cl, Br and I). Spectrochim Acta A 87:106–111
Strobel SA, Cochrane JC (2007) RNA catalysis: ribozymes, ribosomes, and riboswitches. Curr Opin Chem Biol 11:636–643
Strulson CA, Molden RC, Keating CD, Bevilacqua PC (2012) RNA catalysis through compartmentalization. Nat Chem 4:941–946
Suga K, Umakoshi H (2013) Detection of nanosized ordered domains in DOPC/DPPC and DOPC/Ch binary lipid mixture systems of large unilamellar vesicles using a TEMPO quenching method. Langmuir 29:4830–4837
Suga K, Umakoshi H, Tomita H et al (2010) Liposomes destabilize tRNA during heat stress. Biotechnol J 5:526–529
Suga K, Tanabe T, Tomita H et al (2011) Conformational change of single-stranded RNAs induced by liposome binding. Nucl Acid Res 39:8891–8900
Suga K, Tanabe T, Umakoshi H (2013) Heterogeneous cationic liposomes modified with 3β-{N-[(N′, N′-dimethylamino) ethyl] carbamoyl} cholesterol can induce partial conformational changes in messenger RNA and regulate translation in an Escherichia coli cell-free translation system. Langmuir 29:1899–1907
Tanaka Y, Kasai Y, Mochizuki S et al (2004) Nature of the chemical bond formed with the structural metal ion at the A9/G10.1 motif derived from hammerhead ribozymes. J Am Chem Soc 126:744–752
Tessera M (2009) Life began when evolution began: a lipidic vesicle-based scenario. Orig Life Evol Biosph 39:559–564
Tsuji A, Yoshikawa K (2010) Real-time monitoring of RNA synthesis in a phospholipid-coated microdroplet as a live-cell model. ChemBioChem 11:351–357
Veatch SL, Keller SL (2005) Miscibility phase diagrams of giant vesicles containing sphingomyelin. Phys Rev Lett 94:148101
Walde P, Umakoshi H, Stano P, Mavellid F (2014) Emergent proper-ties arising from the assembly of amphiphiles. Artificial vesicle membranes as reaction promoters and regulators. Chem Commun 50:10177–10197
Wang S, Karbstein K, Paracchi A et al (1999) Identification of the hammerhead ribozyme metal ion binding site responsible for rescue of the deleterious effect of a cleavage site phosphorothioate. Biochem 38:14363–14378
Wieland M, Ausländer D, Fussenegger M (2012) Engineering of ribozyme-based riboswitches for mammalian cells. Methods 56:351–357
Zhelyaskov V, Yue KT (1992) A Raman study of the binding of Fe(III) to ATP and AMP. Biochem J 287:561–566
Zipper H, Brunner H, Bernhagen J, Vitzthum F (2004) Investigations on DNA intercalation and surface binding by SYBR Green I, its structure determination and methodological implications. Nucl Acid Res 32:e103
Acknowledgments
We thank Dr. Toshinori Shimanouchi (Graduate School of Environmental Science, Okayama University) for his constructive comments. This work was supported by the Funding Program for Next Generation World-Leading Researchers of the Council for Science and Technology Policy (CSTP) (GR066), JSPS Grant-in-Aid for Scientific Research A (26249116), and JSPS Grant-in-Aid for Research Activity Start-up (25889039). One of the authors (K.S.) also expresses his gratitude for the Japan Society for the Promotion of Science (JSPS) and GCOE scholarships.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing financial interest.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Suga, K., Tanaka, S. & Umakoshi, H. Liposome membrane can induce self-cleavage of RNA that models the core fragments of hammerhead ribozyme. Eur Biophys J 45, 55–62 (2016). https://doi.org/10.1007/s00249-015-1076-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00249-015-1076-z