Abstract
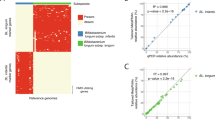
The early bifidobacterial colonization and development of infant gut is considered crucial for the immediate and lifelong health of human host. This study longitudinally analyzed and characterized fecal bifidobacterial profiles in association with feeding regimens observed in six infants during 5 months after birth. The dominant fecal microbiota of bifidobacteria, lactobacilli/enterococci, clostridia, bacteroides and eubacteria were specifically enumerated using fluorescent in situ hybridization (FISH) technique. Breastfeeding exhibited close association with the predomination of bifidobacteria with the highest relative abundance of 32–70% detected in both infants with exclusive breastfeeding. The nested PCR-DGGE technique revealed high diversity existing within a bifidobacterial species with multiple strain variants of B. bifidum, B. longum, B. breve and B. dentium continuously detected in feces of exclusively breast- and combination-fed infants over the period of 5 months. Contrarily, B. breve, B. adolescentis, B. dentium, B. bifidum, B. faecale, B. kashiwanohense and B. lactis detected in all exclusively formula-fed infants seem to be transient species. The persisting strains seem to derive primarily from maternal breastmilk as demonstrated by PCR-DGGE profiles of human milk and feces from three mother-infant pairs. The results suggested the pivotal role of breastfeeding regimen in supporting colonization and succession of bifidobacteria in infant gut.



Similar content being viewed by others
References
Azad MB, Konya T, Persaud RR, Guttman DS, Chari RS, Field CJ, Sears MR, Mandhane PJ, Turvey SE, Subbarao P, Becker AB, Scott JA, Kozyrskyj AL (2016) Impact of maternal intrapartum antibiotics, method of birth and breastfeeding on gut microbiota during the first year of life: a prospective cohort study. BJOG 123:983–993. https://doi.org/10.1111/1471-0528.13601
Ballesté E, Blanch AR (2011) Bifidobacterial diversity and the development of new microbial source tracking indicators. Appl Environ Microbiol 77(10):3518–3525
Baglatzi L, Gavrili S, Stamouli K, Zachaki S, Favre L, Pecquet S, Benyacoub J, Costalos C (2016) Effect of infant formula containing a low dose of the probiotic Bifidobacterium lactis CNCM I-3446 on immune and gut functions in C-Section delivered babies: a pilot study. Clin Med Insights Pediatr 10:11–19. https://doi.org/10.4137/CMPed.S33096
Barrett E, Deshpandey AK, Ryan CA, Dempsey EM, Murphy B, O’Sullivan L, Watkins C, Ross, RP, O’Toole PW, Fitzgerald GF, Stanton C (2015) The neonatal gut harbours distinct bifidobacterial strains. Arch Dis Child Fetal Neonatal Ed 100:405–410. https://doi.org 10.1136/archdischild-2014–306110
Braegger C, Chmielewska A, Decsi T, Kolacek S, Mihatsch W, Moreno L, Piescik M, Puntis J, Shamir R, Szajewska H, Turck D, van Goudoever J (2011) Supplementation of infant formula with probiotics and/or prebiotics: a systematic review and comment by the ESPGHAN committee on nutrition. J Pediatr Gastroenterol Nutr 52:238–250. https://doi.org/10.1097/MPG.0b013e3181fb9e80
Chen J, Cai W, Feng Y (2007) Development of intestinal bifidobacteria and lactobacilli in breastfed neonates. Clin Nutr 26:559–566. https://doi.org/10.1016/j.clnu.2007.03.003
Chichlowski M, De Lartique G, German J, Raybould HE, Mills DA (2012) Bifidobacteria isolated from infants and cultured on human milk oligosaccharides affect intestinal epithelial function. J Pediatr Gastroenterol Nutr 55:321–327. https://doi.org/10.1097/MPG.0b013e31824fb899
Codipilly CN (2014) Role of human milk in the development of gastrointestinal bacterial flora and immunity in preterm infants. MOJ Immunol 1(4)
Euler AR, Mitchell DK, Kline R, Pickering LK (2005) Prebiotic effect of fructo-oligosaccharide infant formula at two concentrations compared with unsupplemented formula and human milk. J Pediatr Gastr Nutr 40:157–164. https://doi.org/10.1097/00005176-200502000-00014
Fanaro S, Boehm G, Garssen J, Knol J, Mosca F, Stahl B, Vigi V (2005) Galacto-oligosaccharides and long-chain fructo-oligosaccharides as prebiotics in infant formulas: a review. Acta Paediatr 94:22–26. https://doi.org/10.1111/j.1651-2227.2005.tb02150.x
Ferretti P, Pasolli E, Tett A, Asnicar F, Gorfer V, Fedi S, Armanini F, Truong DT, Manara S, Zolfo M, Beghini F, Bertorelli R, de Sanctis V, Bertorelli I, Canto R, Clementi R, Cologna M, Crifo T et al (2018) Mother-to-infant microbial transmission from different body sites shapes the developing infant gut microbiome. Cell Host Microobe 24:133–145. https://doi.org/10.1016/j.chom.2018.06.005
Fitzstevens JL, Smith KC, Hagadorn JI, Caimano MJ, Matson AP, Brownell EA (2017) Systematic review of the human milk microiota. Nutr Clin Pract 32:354–364. https://doi.org/10.1177/0884533616670150
Fukuda K, Ogawa M, Taniguchi H, Saito M (2016) Molecular approaches to studying microbial communities: targeting the 16S ribosomal RAN gens. J UOEH 38:223–232. https://doi.org/10.7888/juoeh.38.223
Garrido D, Ruiz-Moyano S, Kirmiz N, Davis JC, Totten SM, Lemay DG, Ugalde JA, German JB, Lebrilla CB, Mills DA (2016) A novel gene cluster allows preferential utilization of fucosylated milk oligosaccharides in Bifidobacterium longum subsp. longum SC596. Sci Rep. https://doi.org/10.1038/srep35045
Gonzalez-Rodriguez I, Ruiz L, Gueimonde M, Margolles A, Sanchez B (2013) Factors involved in the colonization and survival of bifidoacteria in the gastrointestinal tract. FEMS Microbiol Lett 340:1–10. https://doi.org/10.1111/1574-6968.12056
Gronlund MM, Gueimode M, Laitinen K, Kociubinsski G, Gronroos T, Salminen S, Isolauri E (2007) Maternal breast-milk and intestinal bifidobacteria guide the compositional development of the Bifidobacterium microbiota in infants at risk of allergic disease. Clin Exp Allergy 37:1764–1772. https://doi.org/10.1111/j.1365-2222.2007.02849.x
Harmsen HJM, Wildeboer-Veloo AC, Raangs GC, Wagendorp AA, Klijn N, Bindels JG, Welling GW (2000) Analysis of intestinal flora development in breast-fed and formula-fed infants by using molecular identification and detection methods. J Pediatr Gastr Nutr 30:61–67. https://doi.org/10.1097/00005176-200001000-00019
Ishibashi N, Yaeshima T, Hayasawa H (1997) Bifidobacteria: their significance in human intestinal health. Mal J Nutr 3:149–159
Liepke C, Adermann K, Raida M, Magert HJ, Forssman WG, Zucht HD (2002) Human milk provides peptides highly stimulating the growth of bifidobacteria. Eur J Biochem 269:712–718. https://doi.org/10.1046/j.0014-2956.2001.02712.x
Lawson MAE, O’Neill IJ, Kujawska M, Javvadi SG, Wijeyesekere A, Flegg Z, Chalklen L, Hall LJ (2019) Breast milk-derived human milk oligosaccharides promote Bifidobacterium interactions within a single ecosystem. ISME J. https://doi.org/10.1038/s41396-019-0553-2
Makino H, Kushiro A, Ishikawa E, Kubota H, Gawad A, Sakai T, Oishi K, Martin R, Ben-Amor K, Knol J, Tanaka R (2013) Mother-to-infant transmission of intestinal bifidobacterial strains has an impact on the early development of vaginally delivered infant’s microbiota. PLoS ONE 8:e78331. https://doi.org/10.1371/journal.pone.0078331
Mangin I, Suau A, Gotteland M, Brunser O, Pochart P (2010) Amoxicillin treatment modifies the composition of Bifidobacterium species in infant intestinal microbiota. Anaerobe 16:433–438. https://doi.org/10.1016/j.anaerobe.2010.06.005
Marcobal A, Sonnenburg JL (2012) Human milk oligosaccharide consumption by intestinal microbiota. Clin Microbiol Infect 18:12–15. https://doi.org/10.1111/j.1469-0691.2012.03863.x
Marques MT, Wall R, Ross PR, Fitzgerald FG, Ryan AC, Stanton C (2010) Programming infant gut microbiota: influence of dietary and environmental factors. Curr Opin Biotechnol 21:149–156. https://doi.org/10.1016/j.copbio.2010.03.020
Martin CR, Ling PR, Blackburn GL (2016) Review of infant feeding: key features of breast milk and infant formula. Nutrients 8:279. https://doi.org/10.3390/nu8050279
Maynard CL, Elson CO, Hatton RD, Weaver CT (2012) Reciprocal interactions of the intestinal microbiota and immune system. Nature 489:231–241. https://doi.org/10.1038/nature11551
Milani C, Mancabelli L, Lugli GA, Duranti S, Turroni F, Ferrario C, Mangifesta M, Viappiani A, Ferretti P, Gorfer V, Tett A, Segata N, van Sinderen D, Ventura M (2015) Exploring vertical transmission of bifidoacteria from mother the child. Appl Environ Microbiol 81:7078–7087. https://doi.org/10.1128/AEM.02037-15
O’Callaghan A, van Sinderen D (2016) Bifidobacteria and their role as members of the human gut microbiota. Front Microbiol. https://doi.org/10.3389/fmicb.2016.00925
Oliveira DL, Wilbey RA, Grandison AS, Roseiro LB (2015) Milk oligosaccharides: a review. Int J Dairy Technol 68:305–321. https://doi.org/10.1111/1471-0307.12209
Palmer C, Bik EM, DiGiulio DB, Relman DA, Brown PO (2007) Development of the human infant intestinal microbiota. PLoS Biol 5:e177. https://doi.org/10.1371/journal.pbio.0050177
Pannaraj PS, Li F, Cerini C, Bender JM, Yang S, Rollie A, Adisetiyo H, Zabih S, Lincez PJ, Bittinger K, Bailey A, Bushman FD, Sleasman JW, Aldrovandi GM (2017) Association between breast milk bacterial communities and establishment and development of the infant gut microbiome. JAMA Pediatr 171:647–654. https://doi.org/10.1001/jamapediatrics.2017.0378
Penders J, Thijs C, Vink C, Stelma FF, Snijders B, Kummeling I (2006) Factors influencing the composition of the intestinal microbiota in early infancy. Pediatrics 118:511–521. https://doi.org/10.1542/peds.2005-2824
Plaza-Diaz J, Gomez-Llorente C, Fontana L, Gil A (2014) Modulation of immunity and inflammatory gene expression in the gut, in inflammatory diseases of the gut and in the liver by probiotics. World J Gastroenterol 20:15632–15649. https://doi.org/10.3748/wjg.v20.i42.15632
Quigley EMM (2013) Gut bacteria in health and disease. Gastroenterol Hepatol 9:560–569
Rivière A, Selak M, Lantin D, Leroy F, De Vuyst L (2016) Bifidobacteria and butyrate-producing colon bacteria: importance and strategies for their stimulation in the human gut. Front Microbiol. https://doi.org/10.3389/fmicb.2016.00979
Rodriguez JM (2014) The origin of human milk bacteria: is there a bacterial entero-mammary pathway during late pregnancy and lactation? Adv Nutr 14:779–784. https://doi.org/10.3945/an.114.007229
Roger LC, McCartney AL (2010) Longitudinal investigation of the faecal microbiota of healthy full-term infants using fluorescence in situ hybridization and denaturing gradient gel electrophoresis. Microbiology 156:3317–3328. https://doi.org/10.1099/mic.0.041913-0
Ruiz L, Delgado S, Ruas-Madiedo P, Sanchez B, Margolles A (2017) Bifidobacteria and their molecular communication with the immune system. Front Microbiol. https://doi.org/10.3389/fmicb.2017.02345
Ruiz-Moyano S, Totten SM, Garrido D, Smilowitz JT, German JB, Lebrilla CB, Mills DA (2013) Variation in consumption of human milk oligosaccharides by infant-gut associated strains of Bifidobacterium breve. Appl Environ Microbiol 79:6040–6049. https://doi.org/10.1128/AEM.01843-13
Rutayisire E, Huang K, Liu Y, Tao F (2016) The mode of delivery affects the diversity and colonization pattern of the gut microbiota during the first year of infants’ life: a systematic review. BMC Gastroenterol. https://doi.org/10.1186/s1287601604980
Satokari RM, Vaughan EE, Favier FC, Edwards JC, de Vos WM (2002) Diversity of Bifidobacterium and Lactobacillus spp. in breast-fed and formula-fed infants as assessed by 16S rDNA sequence differences. Microb Ecol Health Dis 14:97–105. https://doi.org/10.1080/08910600260081748
Sakata S, Tonooka T, Ishizeki S, Takada M, Sakamoto M, Fukuyama M, Benno Y (2005) Culture-independent analysis of fecal microbiota in infants, with special reference to Bifidobacterium species. FEMS Microbiol Lett 243:417–423. https://doi.org/10.1016/j.femsle.2005.01.002
Servin AL (2004) Antagonistic activities of lactobacilli and bifidobacteria against microbial pathogens. FEMS Microbiol Rev 28:405–440. https://doi.org/10.1016/j.femsre.2004.01.003
Solis G, delosReyes-Gavilan CG, Fernadez N, Margolles A, Gueimonde M (2010) Establishment and development of lactic acid bacteria and bifidobacteria microbiota in breast-milk and infant gut. Anaerobe 16:307–310. https://doi.org/10.1016/j.anaerobe.2010.02.004
Stewart CJ, Ajami NJ, O’Brien JL, Hutchinson DS, Smith DP, Wong MC, Ross MC, Lloyd RE, Doddapaneni HV, Metcalf GA, Muzny D, Gibbs RA, Vatanen T, Huttenhower C, Xavier RJ, Rewers M, Hagopian W, Toppari J, Ziegler AG, She JX, Akolkar B, Lernmark A, Hyoty H, Vehik K (2018) Temporal development of the gut microiome in early childhood from the TEDDY study. Nature 7728:583–588. https://doi.org/10.1038/s41586-018-0617-x
Sun J, Chang EB (2014) Exploring gut microbes in human health and disease: pushing the envelope. Genes Dis 1:132–139. https://doi.org/10.1016/j.gendis.2014.08.001
Tannock GW, Lawley B, Munro K, Gowri-Pathmanathan S, Zhou SJ, Makrides M, Gison RA, Sullivan T, Prosser CG, Lowry D, Hodgkinson AJ (2013) Comparison of the compositions of the stool microbiotas of infants fed goat milk formula, cow milk-based formula, or breast milk. Appl Environ Microbiol 79:3040–3048. https://doi.org/10.1128/AEM.03910-12
Tannock GW, Lee PS, Wong KH, Lawley B (2016) Why don't all infants have bifidobacteria in their stool? Front Microbiol. https://doi.org/10.3389/fmicb.2016.00834
Thompson-Chagoyan CO, Maldonado J, Gil A (2007) Colonization and impact of disease and other factors on intestinal microbiota. Dig Dis Sci 52:2069–2077. https://doi.org/10.1007/s10620-006-9285-z
Thomson P, Medina DA, Garrido D (2018) Human milk oligosaccharides and infant gut bifidobacteria: molecular strategies for their utilization. Food Microiol 75:37–46. https://doi.org/10.1016/j.fm.2017.09.001
Timmerman HM, Rutten NBMM, Boekhorst J, Saulnier DM, Kortman GAM, Contractor N, Kullen M, Floris E, Harmsen HJM, Vlieger AM, Kleerebezem M, Rijkers GT (2017) Intestinal colonization patterns in breast-fed and formula-fed infants during the first 12 weeks of life reveal sequential microbiota signatures. Sci Rep. https://doi.org/10.1038/s41598-017-08268-4
Turroni F, Peano C, Pass DA, Foroni E, Severgnini M, Claesson MJ, Kerr C, Hourihane J, Murray D, Fuligni F, Gueimonde M, Margolles A, De Bellis G, O'Toole PW, van Sinderen D, Marchesi JR, Ventura M (2012) Diversity of bifidobacteria within the Infant Gut Microbiota. PLoS ONE 7:e36957. https://doi.org/10.1371/journal.pone.0036957
Turroni F, Milani C, Duranti S, Mahony J, van Sinderen D, Ventura M (2018) Glycan utilization and cross-feeding activities by bifidobacteria. Trends Microbiol 26:339–350. https://doi.org/10.1016/j.tim.2017.10.001
Underwood MA, Kalanetra KM, Bokulich NA, Lewis ZT, Mirmiran M, Tancredi DJ, Mills DA (2013) A comparison of two proiotic strains of bifidobacteria in premature infants. J Pediatr. https://doi.org/10.1016/j.jpeds.2013.07.017
Uraipan S, Brigidi P, Hongpattarakere T (2014) Antagonistic mechanism of synbiosis between Lactobacillus plantarum CIF17AN2 and green banana starch in the proximal colon model challenged with Salmonella Typhimurium. Anaerobe 28:44–53. https://doi.org/10.1016/j.anaerobe.2014.05.002
Uraipan S, Hongpattarakere T (2015) Antagonistic characteristics against food-borne pathogenic bacteria of lactic acid bacteria and bifidobacteria isolated from feces of healthy Thai infants. Jundishapur J Microbiol 8:e18264. https://doi.org/10.5812/jjm.8(6)2015.18264
Walter J, Hertel C, Tannock GW, Lis CM, Munro K, Hammes WP (2001) Detection of Lactobacillus, Pediococcus, Leuconostoc, and Weissella species in human feces by using group-specific PCR primers and denaturing gradient gel electrophoresis. Appl Environ Microbiol 6:2578–2585. https://doi.org/10.1128/AEM.67.6.2578-2585.2001
Zwielehner J, Lassl C, Hippe B, Pointner A, Switzeny OJ, Remely M, Kitzeeger E, Ruckser R, Haslberger AG (2011) Changes in human fecal microbiota due to chemotherapy analyzed by TaqMan-PCR, 454 sequencing and PCR-DGGE fingerprinting. PLoS ONE 6:e28654. https://doi.org/10.1371/journal.pone.0028654
Zúñiga M, Monedero V, Yebra MJ (2018) Utilization of host-derived glycans by intestinal Lactobacillus and Bifidobacterium species. Front Microbiol 9:1917
Acknowledgements
This work was financially supported by Graduate School, Prince of Songkla University and the Office of the Higher Education Commission under the Strategic Scholarships for Frontier Research Networks (Specific for Southern region) for the Joint Ph.D. Program Thai Doctoral Degree Scholarship (CHE-SSR-Ph.D SW # 03–2553 awarded to Khanitta Kongnum under supervision of Associate Prof. Tipparat Hongapttarakere, Ph.D.).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
The protocol of this study was reviewed and approved by the Ethics Committee of Faculty of Medicine, Prince of Songkla University (EC Numbers: 55-244-19-2-3 and 55-243-19-2-3). All infant’s parents provided verbal informed consent on behalf of the participating babies. The verbal consent was granted by the Ethics Committee because this research presented minimum risks to the participants. In addition, they were not identifiable from the data collected. Infants’ parents directly rung and handed the researcher to pick up the samples right after defecation or before breastfeeding (< 1 h) to prevent the samples from oxygen exposure for too long period.
Additional information
Communicated by Erko Stackebrandt.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Kongnum, K., Taweerodjanakarn, S. & Hongpattarakere, T. Longitudinal characterization of bifidobacterial abundance and diversity profile developed in Thai healthy infants. Arch Microbiol 202, 1425–1438 (2020). https://doi.org/10.1007/s00203-020-01856-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00203-020-01856-5