Abstract
Aims/hypothesis
Circulating metabolites have been shown to reflect metabolic changes during the development of type 2 diabetes. In this study we examined the association of metabolite levels and pairwise metabolite ratios with insulin responses after glucose, glucagon-like peptide-1 (GLP-1) and arginine stimulation. We then investigated if the identified metabolite ratios were associated with measures of OGTT-derived beta cell function and with prevalent and incident type 2 diabetes.
Methods
We measured the levels of 188 metabolites in plasma samples from 130 healthy members of twin families (from the Netherlands Twin Register) at five time points during a modified 3 h hyperglycaemic clamp with glucose, GLP-1 and arginine stimulation. We validated our results in cohorts with OGTT data (n = 340) and epidemiological case–control studies of prevalent (n = 4925) and incident (n = 4277) diabetes. The data were analysed using regression models with adjustment for potential confounders.
Results
There were dynamic changes in metabolite levels in response to the different secretagogues. Furthermore, several fasting pairwise metabolite ratios were associated with one or multiple clamp-derived measures of insulin secretion (all p < 9.2 × 10−7). These associations were significantly stronger compared with the individual metabolite components. One of the ratios, valine to phosphatidylcholine acyl-alkyl C32:2 (PC ae C32:2), in addition showed a directionally consistent positive association with OGTT-derived measures of insulin secretion and resistance (p ≤ 5.4 × 10−3) and prevalent type 2 diabetes (ORVal_PC ae C32:2 2.64 [β 0.97 ± 0.09], p = 1.0 × 10−27). Furthermore, Val_PC ae C32:2 predicted incident diabetes independent of established risk factors in two epidemiological cohort studies (HRVal_PC ae C32:2 1.57 [β 0.45 ± 0.06]; p = 1.3 × 10−15), leading to modest improvements in the receiver operating characteristics when added to a model containing a set of established risk factors in both cohorts (increases from 0.780 to 0.801 and from 0.862 to 0.865 respectively, when added to the model containing traditional risk factors + glucose).
Conclusions/interpretation
In this study we have shown that the Val_PC ae C32:2 metabolite ratio is associated with an increased risk of type 2 diabetes and measures of insulin secretion and resistance. The observed effects were stronger than that of the individual metabolites and independent of known risk factors.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Recent technological advances allow simultaneous detection of a wide range of metabolites in blood samples from healthy and diabetic individuals [1]. Studies on type 2 diabetes have provided strong evidence for the association of several blood metabolites with both prevalent and incident type 2 diabetes. In particular, the branched-chain amino acids (BCAAs; valine, leucine and isoleucine) and several phospholipids have consistently been shown to associate with disease progression [1,2,3,4]. Furthermore, there is evidence from OGTTs that these metabolites also associate with insulin secretion and/or insulin sensitivity [5,6,7]. However, OGTT-derived measures do not allow detailed analysis of insulin secretion, for example the response to various non-glucose insulin secretagogues such as glucagon-like peptide-1 (GLP-1) and arginine. GLP-1 is a gut hormone that stimulates insulin secretion from the pancreas, and arginine can be used as a measure of (near maximal) functional beta cell mass [8]. Alterations in the ratios between two single metabolites may point at perturbations in pathways relevant for a certain disease or phenotype and metabolite ratios are indeed known to associate with specific phenotypes [9,10,11,12]. The analysis of metabolite profiles and ratios in response to different insulin secretagogues are thus relevant for further elucidating the underlying biology of the development of type 2 diabetes. Furthermore, they may be useful for early identification of individuals with an increased risk of type 2 diabetes beyond what can be achieved with currently known risk factors.
To the best of our knowledge, this is the first study to analyse metabolite ratios in relation to insulin secretion phenotypes and type 2 diabetes risk.
Methods
Study design
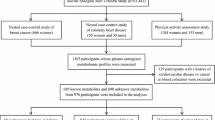
A schematic outline of the study and the rationale for selecting the cohorts is provided in Fig. 1 and in the electronic supplementary material (ESM) Methods. All studies were approved by the appropriate local institutional review boards and participants provided written informed consent before participating in the study.
Schematic overview of the design used in the discovery (blue) and validation (green) phases of the study. MZ, monozygotic; DZ, dizygotic; sibs, siblings. Further details on the study samples can be found in ESM Methods. aMost replication cohorts had only ten of the 18 ratios available
Discovery hyperglycaemic clamp study sample
Metabolite profiles and their responses to glucose, GLP-1 and arginine stimulation were studied using a modified 3 h hyperglycaemic clamp in 130 participants of the Netherlands Twin Register (NTR) [13]. Of the 130 participants, 100 were twins and 30 were non-twin siblings from 54 families. Six of the participants had impaired glucose tolerance, while the remaining individuals had normal glucose tolerance as determined by OGTT. The clinical characteristics of the study group and details of the procedure are described in ESM Methods, ESM Table 1 and schematically presented in Figs 1, 2.
(a) Insulin responses. First- and second-phase GSIS (red and green, respectively), GLP-1-SIS (orange) and arginine-SIS (blue). Blood samples for metabolomics measurements were drawn at t = 0, 30, 120, 180 and 190 min as indicated by the black arrows. (b) Glucose levels. Hyperglycaemia was established and maintained at 10 mmol/l glucose via variable infusion of glucose. After 2 h, insulin secretion was further stimulated using i.v. GLP-1 infusion (1.5 pmol/kg bolus for 1 min at t = 120 followed by a continuous infusion of 0.5 pmol kg−1 min−1 for 1 h). The near maximal insulin response was assessed by injecting a bolus of 5 g arginine hydrochloride at t = 180 min
Validation OGTT study samples
Next we validated our results in two independent cohorts with OGTT data: the Leiden Longevity Study (LLS) [14] and the POGO (Postpartum Outcomes in mothers with Gestational diabetes and their Offspring) study [15] (see ESM Methods for further details). Clinical characteristics of the study participants can be found in ESM Tables 2 and 3. From these studies we included a total of 340 non-diabetic participants who all underwent a standardised OGTT. We calculated six surrogate measures of insulin secretion and insulin resistance (ESM Table 4).
Validation type 2 diabetes study sample
The metabolites that demonstrated significant associations in the clamp phase of the study were further investigated in four independent epidemiological studies where we studied associations with prevalent (LLS [14, 16], NTR [17, 18]; the cooperative health research in the region of Augsburg, Germany [KORA F4] study [19, 20]) or incident (KORA S4_to_F4 prospective follow-up [19, 20] and the European Prospective Investigation into Cancer and Nutrition-Potsdam [EPIC-Potsdam] study [21]) type 2 diabetes. Both the KORA S4_to_F4 and the EPIC-Potsdam studies have an average of 7 years follow-up. Further details of the studies, sampling methods and data collection can be found in references [17,18,19,20,21] and ESM Methods, ESM Tables 2, 5–8 and ESM Figs 1, 2. In the analysis for prevalent diabetes we included a total of 306 individuals with prevalent type 2 diabetes and 4619 non-diabetic volunteers. For the analysis of incident diabetes, we included 910 participants who were free of diabetes at baseline when blood was drawn but who developed type 2 diabetes during follow-up, and 3367 non-diabetic volunteers.
Metabolomic measurements
Plasma concentrations of metabolites in the hyperglycaemic clamp cohort were determined with a commercial assay (AbsoluteIDQ p180 Kit; Biocrates Life Sciences, Innsbruck, Austria). The assay allows the quantification of 188 metabolites. The metabolite abbreviations are provided in ESM Table 9, metabolite naming was as described in Römisch-Margl et al [22]. Fasting and samples at four subsequent time points during the clamp (Fig. 2) were analysed according to the manufacturer’s protocol. A detailed description of the method can be found in the ESM Methods [23]. After quality control, 143 metabolites (135 metabolites and eight calculated compositions) remained for analysis. In the LLS, NTR, KORA F4 and EPIC-Potsdam cohorts, the AbsoluteIDQ p150 Kit was used, according to the methods and quality control procedures as described previously [17, 22]. ESM Table 9 describes all metabolites measured with either the p180 or p150 kits including metabolites that failed quality control in the discovery sample.
Statistics
Discovery phase
In order to account for the family relationships in the hyperglycaemic clamp study we fitted generalised estimating equations (GEEs) using the R package GEEpack, v1.2-0.1 [24] (https://cran.r-project.org/web/packages/geepack/index.html). To analyse dynamic changes in metabolite levels between the different time points the linear regression models were adjusted for age, sex and BMI. In order to reduce the chance of false positives we applied stringent Bonferroni correction to correct for multiple testing (p ≤ 3.5 × 10−4; using α = 0.05 and 143 metabolites/tests). All six clamp-derived phenotypes were quantile normalised before analysis. To study the associations of fasting metabolites or their ratios we applied linear regression models (GEE) unadjusted, age and sex adjusted or adjusted for age, sex, BMI, glucose tolerance status, insulin sensitivity index (if relevant) as potential confounders. The Bonferroni corrected threshold was p ≤ 5.8 × 10−5 (i.e. 858 tests, 143 metabolites × six phenotypes). All possible pairwise metabolite ratios were calculated (log[metab1/metab2]) [12] and analysed as described above for single metabolites. The Bonferroni corrected threshold for the metabolite ratios was p ≤ 9.2 × 10−7 (54,270 tests, 9045 ratios × six phenotypes). In addition, the p gain for each of the metabolite ratios and p gain threshold was calculated (see ESM Methods for details) [12]. A p gain above the threshold value suggests that the association of the metabolite ratio is stronger than that of the two individual metabolites alone.
Validation phase
To allow comparisons across cohorts and to facilitate meta-analysis, metabolite level data were log-transformed followed by z-scaling before analysis. Associations between OGTT-derived measures, prevalent diabetes and metabolite ratios were investigated using either linear or logistic regression models with adjustment for age, sex, BMI, use of lipid lowering medication, study-specific covariates and fasting status (where appropriate) as covariates. Only complete cases with no missing data were analysed. A fixed-effects meta-analysis was performed using the R package Meta v4.3-2 [25] (https://cran.r-project.org/web/packages/meta/index.html).
For the associations between the metabolite ratios and incident diabetes, we performed a Cox proportional hazards regression analysis with covariates as described by Wang-Sattler et al [26] and Floegel et al [7]. See ESM Table 10 for details on the covariates included. The above described base models, to which the ratio of valine and phosphatidylcholine acyl-alkyl (PC ae) C32:2 was added, reflect established prediction models which have been validated in several independent cohort studies [27,28,29]. We used several procedures to evaluate the accuracy of the models as described in the ESM Methods.
Results
Discovery phase
Metabolite dynamics after glucose, GLP-1 and arginine stimulation
There were many significant dynamic metabolite responses observed during the hyperglycaemic clamp procedure. Within group responses were, in general, very similar (i.e. the acylcarnitines, amino acids, etc.; ESM Fig. 3). After glucose stimulation (t = 30 or 120 min vs t = 0), we noted significant reductions (p ≤ 3.5 × 10−4) in the levels of most of the acylcarnitines (10/12), amino acids (21/21), phosphatidylcholines (68/69; except PC ae C42:0), biogenic amines (8/8) and sphingolipids (13/13). However, only a few of the lysophosphatidylcholines (4/11) changed significantly. About one-third of the metabolites that had reduced levels upon stimulation with glucose showed a further reduction after stimulation with GLP-1 (t = 180 vs t = 120). These metabolites belong to the acylcarnitines (10/12), amino acids (21/21), biogenic amines (5/8) and phosphatidylcholines (9/69). Of the metabolites that were unaffected by glucose stimulation only the acylcarnitine C0 decreased significantly after GLP-1 stimulation. After additional stimulation with arginine (t = 190 vs t = 180) about half of the metabolites showed a further significant change. These include acylcarnitines (4/12), amino acids (16/21), phosphatidylcholines (37/69), lysophosphatidylcholines (8/11), biogenic amines (2/8) and sphingolipids (11/13). Only four metabolites, the lysophosphatidylcholines containing myristic acid (C14:0), palmitic acid (16:0), palmitoleic acid (C16:1) and arachidonic acid (C20:4), responded exclusively to arginine stimulation, suggesting that they are specific to arginine. Remarkably, we also observed a large significant increase of phosphatidylcholine acyl-acyl (PC aa) C42:1 after arginine stimulation.
Fasting metabolite levels and insulin secretion (hyperglycaemic clamp)
In the remainder of the discovery study we focused on associations of baseline fasting metabolite levels and pairwise metabolite ratios with the insulin responses after stimulation with the various stimuli. Three baseline metabolites, PC aa C32:1, PC aa C34:4 and PC aa C38:5, showed a significant negative association with second-phase glucose-stimulated insulin secretion (GSIS) or GLP-1-stimulated insulin secretion (SIS) after correction for multiple testing (p < 5.8 × 10−5; Table 1). PC aa C34:4 was associated with both second-phase GSIS and GLP-1-SIS (Table 1). These associations were independent of the effects of age, sex, BMI, glucose tolerance status and insulin sensitivity. PC aa C34:4 and several other metabolites showed suggestive evidence for an association with the other phenotypes as well (p < 1.0 × 10−3; ESM Table 11).
Eighteen fasting pairwise metabolite ratios showed associations that were significantly stronger than the individual metabolites (Table 2), i.e. having a p gain above the threshold. The ratio between alanine and glycine showed the strongest association (with the insulin sensitivity index; β − 0.970 (0.145), p = 2.0 × 10−11, p gain = 2.8 × 108). PC aa C34:4 was the only metabolite that was significant in the single metabolite and the pairwise metabolite ratio analyses (Tables 1, 2; the results from the crude models are shown in ESM Tables 12, 13).
Validation phase
Since it was not possible to replicate our findings in cohorts with similar hyperglycaemic clamp data, we use existing metabolomics data from OGTTs to validate our findings. OGTTs are used to study insulin sensitivity and beta cell responses after stimulation with glucose. Since our main associations were with second-phase GSIS we assumed that similar associations could be found between fasting metabolite levels and insulin secretion measures as derived from OGTTs. We attempted to further validate the observed associations in various epidemiological cohort studies with type 2 diabetes as the endpoint. Most of these existing cohorts used the Biocrates AbsoluteIDQ p150 Kit measuring fewer metabolites. Therefore, a maximum of ten out of the 18 ratios could be used in the meta-analyses (ESM Table 9).
Fasting pairwise metabolite ratios and insulin secretion (OGTT)
In two studies, the LLS and POGO, a total of 340 participants underwent an OGTT. We focused our analyses on six commonly used OGTT-derived measures of insulin secretion and insulin resistance that were available. Analysis of the previously identified fasting metabolite ratios that could also be calculated in these cohorts showed several significant associations (ESM Tables 14, 15). After meta-analysis of the data from both OGTT studies the most significant associations were observed with the ratios of valine to PC ae C32:2, PC aa C32:3 to PC ae C34:3 and valine to lysophosphatidylcholine acyl C18:1 and target variables AUCglucose, AUCinsulin, AUCglucose/AUCinsulin and/or HOMA-IR (all p < 5.4 × 10−3; Table 3), but no associations were found with the insulinogenic index or corrected insulin response. These findings were independent of potential confounders (results from the crude models are shown in ESM Table 16). Additional adjustment for insulin sensitivity, as calculated by HOMA-IR, led to slightly weaker associations with some of the variables (ESM Table 17). However, further adjustment for fasting glucose levels did not essentially affect our results.
Fasting pairwise metabolite ratios and prevalent type 2 diabetes
Next we tested if the pairwise metabolite ratios were associated with prevalent diabetes in three independent epidemiological studies (306 diabetic and 4619 control participants). In a fixed-effects meta-analysis of fully adjusted models, we showed that nine out of the ten tested ratios were significantly associated with prevalent type 2 diabetes (Table 4, all p ≤ 6.4 × 10−5; the results for crude models are shown in ESM Table 18). Only the ratio of valine to PC ae C32:2, showing the strongest association with prevalent type 2 diabetes (ORVal_PC ae C32:2 2.64 [β 0.97 ± 0.09], p = 1.0 × 10−27), showed a p gain above the threshold, i.e. the effect was much stronger than that of the two individual metabolites (Table 4, ESM Table 19; both p ≥ 2.2 × 10−16, p gain = 2.2 × 1011).
Fasting pairwise metabolite ratios at baseline and incident type 2 diabetes
Meta-analysis of the Cox regression results in two independent prospective studies (910 individuals with incident type 2 diabetes and 3367 control participants), with adjustment as shown in ESM Table 10, shows a highly significant association between the ratio of valine to PC ae C32:2 and type 2 diabetes susceptibility (Table 5; HRVal_PC ae C32:2 1.57 [β 0.45 ± 0.06], p = 1.3 × 10−15; the results for the crude models are shown in ESM Table 20). Again, this association was significantly stronger than that observed for the individual metabolites (Table 5, ESM Table 21; both p ≥ 9.2 × 10−9, p gain = 1.3 × 106). Adding glucose levels at baseline to the model only marginally affected the results and the association remained highly significant (HRVal_PC ae C32:2 1.45 [β 0.37 ± 0.06], p = 1.4 × 10−9).
When the valine to PC ae C32:2 ratio was added to the existing baseline prediction model comprising all established traditional risk factors (TRF+glucose) as shown in ESM Table 10, the AUC estimated from the time-dependent receiver operating characteristics improved from 0.780 to 0.801 in the KORA S4_to_F4 study (p = 3.2 × 10−2 for the ratio, ESM Table 22), which was larger than the effect of adding the two single metabolites to the model (AUC 0.793). This is also in line with the results of the net reclassification index.
In the EPIC-Potsdam study we obtained similar results for models with TRF+glucose and TRF+glucose+Val_PC ae C32:2 (0.862 and 0.865, respectively, p = 1.20 × 10−8 for the metabolite ratio). The results were largely similar for the cross-validated performance, suggesting little overfitting in the present situation with a large sample size and few added covariates (ESM Table 22).
Discussion
In the discovery phase, we used the hyperglycaemic clamp, the gold standard for the measurement of insulin secretion [30], to study the association between baseline fasting metabolite levels, pairwise metabolite ratios and insulin response after consecutive stimulation with three different insulin secretagogues [8]. In the validation phase, we tested whether metabolite ratios identified in our clamp study were associated with insulin responses measured using OGTT data from two independent cohorts. Finally, we investigated the associations of the metabolite ratios with prevalent and incident type 2 diabetes in four independent cohorts from the Netherlands and Germany. We observed numerous dynamic metabolite responses during the clamp study reflecting the switch from beta oxidation of fatty acids and gluconeogenesis from amino acids during the overnight fast to a state of glucose oxidation during the hyperglycaemic clamp. We have shown that the ratio of valine to PC ae C32:2 is significantly positively associated with second-phase GSIS, OGTT-derived measures including HOMA-IR, and both prevalent and incident type 2 diabetes.
One limitation of this study is the relatively small sample size in the hyperglycaemic clamp part of the discovery phase, which impacts on power and reproducibility. However, we applied stringent statistical significance criteria in order to correct for multiple testing and have therefore compromised statistical power but enhanced reproducibility. Furthermore, our discovery results are corroborated in the validation phase for which we used at least two independent cohorts per phenotype studied. As described in ESM Methods the Biocrates kit used to detect the metabolites does not allow a detailed analysis of the exact lipid composition of metabolites such as PC ae C32:2. This is a limitation to the interpretation of our results (see ESM Methods for further details). Another limitation is the use of different covariates for adjusting the Cox proportional hazards regression models in the KORA S4_to_F4 and EPIC-Potsdam studies (ESM Table 10). However, both were established sets of risk factors used previously in similar metabolomic studies [7, 26] that have also been validated in external cohorts [27,28,29]. Furthermore, it was the aim of this study to test if metabolite ratios have an added value to these established risk factors and not to find the optimal set of predictors. Since not all covariates are available in both studies the possibilities for harmonisation of the models were limited. Despite these differences both studies yield highly comparable results, which shows the reliability of the findings. In addition, we used a cross-validation approach, which enabled us to assess the accuracy of the predictive model.
It has been shown that metabolite ratios can reveal perturbations in pathways relevant for a certain phenotype and may thus reveal stronger and more meaningful associations [31, 32], even if the mechanism is not clear. Therefore, pairwise ratios may serve as good biomarkers with predictive ability beyond that of the single constituents because noise can be reduced, increasing statistical power [12]. Valine is a BCAA, which are among the most commonly observed metabolites to be increased in type 2 diabetes and are not only responsive to glucose stimulation but also to the glucose-lowering drugs glipizide and metformin [3, 33]. Furthermore, BCAAs are associated with insulin sensitivity [34, 35] and the development of diabetes [4]. A recent Mendelian randomisation study suggested that a causal relationship exists between increased BCAA levels and type 2 diabetes risk [36]; however, it remains to be shown that PC ae C32:2 or the ratio of valine to PC ae C32:2 are also causally related to the disease, but at present there are no genetic instruments available for the latter (see ‘GWAS look-up’ in ESM Methods).
Phosphatidylcholine species, including PC ae C32:2, have been found to be associated with type 2 diabetes. However, since the phosphatidylcholines are not detected on all metabolomics platforms, replication is less frequent compared with the BCAAs [4, 6, 7, 26]. PC ae C32:2 has been shown to be associated with prevalent [6] and incident type 2 diabetes [7] and to respond to glucose stimulation during OGTT and IVGTT [37]. It is clear from our observations that the opposing effects of valine and PC ae C32:2 on insulin secretion are not simply additive, as reflected by the much stronger association of the metabolite ratio compared with the individual metabolites. According to the Human Metabolome database, PC ae C32:2 is composed of either the fatty acids C16:1/C16:1, C18:1/C14:1 or C18:2/C14:0 (www.HMDB.ca, accessed 1 October 2016) [38]. Recently, it has been shown that BCAA catabolism and lipogenesis are linked in adipose tissue [39,40,41]. These studies have shown that catabolism of the BCAAs (leucine, isoleucine and valine) contributes to the synthesis of odd-chain and even-chain fatty acids, such as C14, C16 and C18 chains (i.e. the constituents of PC ae C32:2). It was also shown that BCAA-derived metabolites up or downstream of the branched-chain-alpha-ketoacid dehydrogenase (BCKD) complex, being a rate-limiting step in BCAA catabolism, were associated oppositely with the risk of type 2 diabetes [36]. Further research is necessary to investigate possible functional relationships between valine and PC ae C32:2, and whether or not there is a direct causal relationship with the observed associations with GSIS and the risk of developing diabetes.
In addition to the ratio of valine to PC ae C32:2, we also note several other significant associations in our hyperglycaemic clamp experiments. For example PC aa C32:1 was associated with reduced second-phase GSIS. In previous studies by Floegel et al and Wang-Sattler et al this metabolite has been associated with an increased risk of impaired glucose tolerance and incident type 2 diabetes [7, 26]. Thus, reduced second-phase GSIS provides a potential mechanism for these previous observations. Furthermore, two other phosphatidylcholines, PC aa C34:4 and PC aa C38:5, were previously identified to be reduced in individuals with type 2 diabetes [42] or pregnant women with gestational diabetes mellitus [43]. Interestingly, these metabolites were also found to be influenced by the obesity associated variant in the FTO gene during OGTTs [37]. As such, our data substantiate these previous findings. We also note a significant increase in PC aa C42:1 after arginine stimulation (ESM Fig. 3). This metabolite was previously found to be decreased in individuals with type 2 diabetes [6]. Since the samples from different individuals and time points were randomised and the effect was not caused by a few individuals or outliers this seems to be a genuine observation requiring further investigation.
Next to the single metabolite associations and the valine to PC ae C32:2 ratio, the ratio of alanine and glycine strongly associated with insulin sensitivity measured using the hyperglycaemic clamp and incident diabetes in the KORA S4_to_F4 cohort. It is of interest that both amino acids have previously been identified in metabolomics studies in diabetes, indeed displaying opposing effects (reviewed in [4]). Unfortunately, alanine is not measured with the AbsoluteIDQ p150 Kit and thus the ratio could not be calculated in the other studies and as such findings could not be further validated. If validated in other studies this ratio could be of use in prediction of insulin resistance and diabetes risk.
Here we have shown that the addition of the valine to PC ae C32:2 metabolite ratio improved the accuracy of prediction of incident type 2 diabetes in a model containing known risk factors in both the KORA S4_to_F4 and EPIC-Potsdam cohorts, corroborating results from previous studies that only investigated associations with individual metabolites [7, 26]. We have also shown associations with augmented second-phase GSIS and AUCinsulin independent of measures of insulin resistance and other covariates (ESM Table 23). In addition, we found a positive correlation with HOMA-IR. Therefore, we speculate that the increased diabetes risk is attributable to increases in insulin resistance rather than insulin secretion, as has been suggested previously for valine and other BCAAs [34, 35]. Furthermore, our insulin secretion studies are mainly from healthy individuals and it may be that associations with augmented insulin secretion are dependent on the level of glycaemia as we have previously shown for a genetic variant of G6PC2 [44].
It is important to note that in all of our analyses the effect of the ratio is larger than that observed with the individual metabolites suggesting that the use of ratios may improve prediction above that of the single metabolites. Large prospective studies aiming to identify the best set of predictors (including traditional risk factors and metabolites) are needed to fully elucidate the clinical applicability of using metabolite ratios in the identification of individuals at risk of developing type 2 diabetes. Since metabolomics measurements are simple and relatively non-invasive and alterations in metabolite profiles can be detected years before overt disease develops, the analysis of metabolite ratios may prove to be a useful instrument in personalising prevention and treatment strategies for type 2 diabetes.
In conclusion, we have shown that the ratio of valine to PC ae C32:2 in blood is positively associated with insulin secretion, HOMA-IR and prevalent type 2 diabetes. Furthermore, it predicts incident type 2 diabetes independent of known risk factors, suggesting that it could be useful as an early biomarker for identification of individuals at increased risk for type 2 diabetes.
Abbreviations
- BCAA:
-
Branched-chain amino acids
- BCKD:
-
Branched-chain-alpha-ketoacid dehydrogenase
- EPIC-Potsdam:
-
European Prospective Investigation into Cancer and Nutrition-Potsdam
- GEE:
-
Generalised estimating equations
- GLP-1:
-
Glucagon-like peptide-1
- GSIS:
-
Glucose-stimulated insulin secretion
- KORA:
-
Cooperative health research in the region of Augsburg, Germany
- LLS:
-
Leiden Longevity Study
- NTR:
-
Netherlands Twin Register
- PC aa:
-
Phosphatidylcholine acyl-acyl
- PC ae:
-
Phosphatidylcholine acyl-alkyl
- POGO:
-
Postpartum Outcomes in mothers with Gestational diabetes and their Offspring
- SIS:
-
Stimulated insulin secretion
- TRF:
-
Traditional risk factors
References
Sas KM, Karnovsky A, Michailidis G, Pennathur S (2015) Metabolomics and diabetes: analytical and computational approaches. Diabetes 64:718–732
Roberts LD, Koulman A, Griffin JL (2014) Towards metabolic biomarkers of insulin resistance and type 2 diabetes: progress from the metabolome. Lancet Diabetes Endocrinol 2:65–75
Suhre K (2014) Metabolic profiling in diabetes. J Endocrinol 221:R75–R85
Guasch-Ferre M, Hruby A, Toledo E et al (2016) Metabolomics in prediabetes and diabetes: a systematic review and meta-analysis. Diabetes Care 39:833–846
Friedrich N (2012) Metabolomics in diabetes research. J Endocrinol 215:29–42
Knebel B, Strassburger K, Szendroedi J et al (2016) Specific metabolic profiles and their relationship to insulin resistance in recent-onset type-1 and type-2 diabetes. J Clin Endocrinol Metab 101:2130–2140
Floegel A, Stefan N, Yu Z et al (2013) Identification of serum metabolites associated with risk of type 2 diabetes using a targeted metabolomic approach. Diabetes 62:639–648
Fritsche A, Stefan N, Hardt E, Schutzenauer S, Haring H, Stumvoll M (2000) A novel hyperglycaemic clamp for characterization of islet function in humans: assessment of three different secretagogues, maximal insulin response and reproducibility. Eur J Clin Investig 30:411–418
Illig T, Gieger C, Zhai G et al (2010) A genome-wide perspective of genetic variation in human metabolism. Nat Genet 42:137–141
Suhre K, Raffler J, Kastenmuller G (2016) Biochemical insights from population studies with genetics and metabolomics. Arch Biochem Biophys 589:168–176
Boger RH, Sullivan LM, Schwedhelm E et al (2009) Plasma asymmetric dimethylarginine and incidence of cardiovascular disease and death in the community. Circulation 119:1592–1600
Petersen AK, Krumsiek J, Wagele B et al (2012) On the hypothesis-free testing of metabolite ratios in genome-wide and metabolome-wide association studies. BMC Bioinforma 13:120
Simonis-Bik AM, Eekhoff EM, de Moor MH et al (2009) Genetic influences on the insulin response of the beta cell to different secretagogues. Diabetologia 52:2570–2577
Rozing MP, Westendorp RG, de Craen AJ et al (2010) Favorable glucose tolerance and lower prevalence of metabolic syndrome in offspring without diabetes mellitus of nonagenarian siblings: the Leiden longevity study. J Am Geriatr Soc 58:564–569
Hummel S, Much D, Rossbauer M, Ziegler AG, Beyerlein A (2013) Postpartum outcomes in women with gestational diabetes and their offspring: POGO study design and first-year results. Rev Diabet Stud 10:49–57
Westendorp RG, van Heemst D, Rozing MP et al (2009) Nonagenarian siblings and their offspring display lower risk of mortality and morbidity than sporadic nonagenarians: the Leiden Longevity Study. J Am Geriatr Soc 57:1634–1637
Draisma HH, Pool R, Kobl M et al (2015) Genome-wide association study identifies novel genetic variants contributing to variation in blood metabolite levels. Nat Commun 6:7208
Willemsen G, de Geus EJ, Bartels M et al (2010) The Netherlands Twin Register biobank: a resource for genetic epidemiological studies. Twin Res Hum Genet 13:231–245
Holle R, Happich M, Lowel H, Wichmann HE, Group MKS (2005) KORA—a research platform for population based health research. Gesundheitswesen 67(Suppl 1):S19–S25
Rathmann W, Strassburger K, Heier M et al (2009) Incidence of type 2 diabetes in the elderly German population and the effect of clinical and lifestyle risk factors: KORA S4/F4 cohort study. Diabet Med 26:1212–1219
Boeing H, Korfmann A, Bergmann MM (1999) Recruitment procedures of EPIC-Germany. European Investigation into Cancer and Nutrition. Ann Nutr Metab 43:205–215
Römisch-Margl W, Prehn C, Bogumil R, Röhring C, Suhre K, Adamski J (2012) Procedure for tissue sample preparation and metabolite extraction for high-throughput targeted metabolomics. Metabolomics 8:133–142
Zukunft S, Sorgenfrei M, Prehn C, Möller G, Adamski J (2013) Targeted metabolomics of dried blood spot extracts. Chromatographia 76:1295–1305
Højsgaard S, Halekoh U, Yan J (2006) The R package GEEpack for generalized estimating equations. J Stat Softw 15:1–11
Hartung J, Knapp G (2001) On tests of the overall treatment effect in meta-analysis with normally distributed responses. Stat Med 20:1771–1782
Wang-Sattler R, Yu Z, Herder C et al (2012) Novel biomarkers for pre-diabetes identified by metabolomics. Mol Syst Biol 8:615
Abbasi A, Corpeleijn E, Peelen LM et al (2012) External validation of the KORA S4/F4 prediction models for the risk of developing type 2 diabetes in older adults: the PREVEND study. Eur J Epidemiol 27:47–52
Kengne AP, Beulens JW, Peelen LM et al (2014) Non-invasive risk scores for prediction of type 2 diabetes (EPIC-InterAct): a validation of existing models. Lancet Diabetes Endocrinol 2:19–29
Schulze MB, Hoffmann K, Boeing H et al (2007) An accurate risk score based on anthropometric, dietary, and lifestyle factors to predict the development of type 2 diabetes. Diabetes Care 30:510–515
Defronzo RA, Tobin JD, Andres R (1979) Glucose clamp technique: a method for quantifying insulin secretion and resistance. Am J Physiol-Endocrinol Metab 237:E214–E223
Heemskerk MM, van Harmelen VJ, van Dijk KW, van Klinken JB (2016) Reanalysis of mGWAS results and in vitro validation show that lactate dehydrogenase interacts with branched-chain amino acid metabolism. Eur J Hum Genet: EJHG 24:142–145
Wahl S, Yu Z, Kleber M et al (2012) Childhood obesity is associated with changes in the serum metabolite profile. Obes Facts 5:660–670
Walford GA, Davis J, Warner AS et al (2013) Branched chain and aromatic amino acids change acutely following two medical therapies for type 2 diabetes mellitus. Metabolism 62:1772–1778
Wurtz P, Soininen P, Kangas AJ et al (2013) Branched-chain and aromatic amino acids are predictors of insulin resistance in young adults. Diabetes Care 36:648–655
Wurtz P, Makinen VP, Soininen P et al (2012) Metabolic signatures of insulin resistance in 7,098 young adults. Diabetes 61:1372–1380
Lotta LA, Scott RA, Sharp SJ et al (2016) Genetic predisposition to an impaired metabolism of the branched-chain amino acids and risk of type 2 diabetes: a Mendelian randomisation analysis. PLoS Med 13:e1002179
Wahl S, Krug S, Then C et al (2013) Comparative analysis of plasma metabolomics response to metabolic challenge tests in healthy subjects and influence of the FTO obesity risk allele. Metabolomics 10:386–401
Wishart DS, Knox C, Guo AC et al (2009) HMDB: a knowledgebase for the human metabolome. Nucleic Acids Res 37:D603–D610
Green CR, Wallace M, Divakaruni AS et al (2016) Branched-chain amino acid catabolism fuels adipocyte differentiation and lipogenesis. Nat Chem Biol 12:15–21
Crown SB, Marze N, Antoniewicz MR (2015) Catabolism of branched chain amino acids contributes significantly to synthesis of odd-chain and even-chain fatty acids in 3T3-L1 adipocytes. PLoS One 10:e0145850
Halama A, Horsch M, Kastenmuller G et al (2016) Metabolic switch during adipogenesis: from branched chain amino acid catabolism to lipid synthesis. Arch Biochem Biophys 589:93–107
Suhre K, Meisinger C, Doring A et al (2010) Metabolic footprint of diabetes: a multiplatform metabolomics study in an epidemiological setting. PLoS One 5:e13953
Lehmann R, Friedrich T, Krebiehl G et al (2015) Metabolic profiles during an oral glucose tolerance test in pregnant women with and without gestational diabetes. Exp Clin Endocrinol Diabetes 123:483–438
Heni M, Ketterer C, ‘t Hart LM, et al. (2010) The impact of genetic variation in the G6PC2 gene on insulin secretion depends on glycemia. J Clin Endocrinol Metab 95: E479-E484
Acknowledgements
The authors would like to thank all of the volunteers for their participation in this study. We thank W. Römisch-Margl, J. Scarpa and K. Faschinger for metabolomics measurements performed at the Helmholtz Zentrum München, Genome Analysis Center, Metabolomics Core Facility. Some of the data were presented as an abstract at the EASD meeting in Munich in 2016.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Data availability
The data used in this study are available upon request.
Funding
This work was partly funded by the Innovative Medicines Initiative Joint Undertaking under grant agreement no. 115317 (DIRECT), resources of which are composed of financial contributions from the European Union’s Seventh Framework Programme (FP7/2007–2013) and EFPIA companies’ in kind contributions (http://www.direct-diabetes.org/); the Netherlands Organization for Health Research and Development (Priority Medicines Elderly Program 113102006); a grant from the German Federal Ministry of Education and Research (BMBF) to the German Centre Diabetes Research (DZD); a grant from the Helmholtz Initiative Personalized Medicine (iMED); grants from the German Diabetes Association and the Helmholtz International Research Group (Helmholtz HIRG-0018); the German Diabetes Centre is funded by the German Federal Ministry of Health (BMG) and the Ministry of Innovation, Science, Research and Technology (MIWF) of the State North Rhine-Westphalia. KS was supported by ‘Biomedical Research Program’ funds at Weill Cornell Medicine in Qatar, a programme funded by the Qatar Foundation. The NTR is supported by the European Research Council (grant 230374), by BBMRI-NL, a Research Infrastructure financed by the Dutch government (NWO 184.021.007) and the Netherlands Organization for Scientific Research (grant NWO 480-04-004, NWO/SPI 56-464-14192). The LLS was financially supported by the Innovation-Oriented Research Program on Genomics (SenterNovem IGE01014 and IGE05007), the Centre for Medical Systems Biology and the Netherlands Consortium for Healthy Ageing (grant 05040202 and 050-060-810), all in the framework of the Netherlands Genomics Initiative, Netherlands Organization for Scientific Research (NWO), by BBMRI-NL, a Research Infrastructure financed by the Dutch government (NWO 184.021.007), the European Union-funded Network of Excellence Lifespan (FP6 036894) and the European Union’s Seventh Framework Programme (FP7/2007-2011) under grant agreement no. 259679.
Duality of interest
The authors declare that there is no duality of interest associated with this manuscript.
Contribution statement
LMtH, ERP and JA contributed to the conception and design of the study. SM, SW, MH, RP, AF, JD, DM, CP, MB, HHD, NvL, WB and LMtH researched data. All authors contributed to the acquisition or analysis and interpretation of results. SM, SW and LMtH wrote the manuscript. All authors contributed to revising the manuscript for important intellectual content and approved the final version of the manuscript. LMtH is the guarantor of the study and, as such, had full access to all the data in the study and takes responsibility for the integrity of the data and the accuracy of the data analysis.
Electronic supplementary material
ESM
(PDF 1216 kb)
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Molnos, S., Wahl, S., Haid, M. et al. Metabolite ratios as potential biomarkers for type 2 diabetes: a DIRECT study. Diabetologia 61, 117–129 (2018). https://doi.org/10.1007/s00125-017-4436-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00125-017-4436-7