Abstract
Aims/hypothesis
Non-invasive imaging of the pancreatic beta cell mass (BCM) requires the identification of novel and specific beta cell biomarkers. We have developed a systems biology approach to the identification of promising beta cell markers.
Methods
We followed a functional genomics strategy based on massive parallel signal sequencing (MPSS) and microarray data obtained in human islets, purified primary rat beta cells, non-beta cells and INS-1E cells to identify promising beta cell markers. Candidate biomarkers were validated and screened using established human and macaque (Macacus cynomolgus) tissue microarrays.
Results
After a series of filtering steps, 12 beta cell-specific membrane proteins were identified. For four of the proteins we selected or produced antibodies targeting specifically the human proteins and their splice variants; all four candidates were confirmed as islet-specific in human pancreas. Two splice variants of FXYD domain containing ion transport regulator 2 (FXYD2), a regulating subunit of the Na+–K+-ATPase, were identified as preferentially present in human pancreatic islets. The presence of FXYD2γa was restricted to pancreatic islets and selectively detected in pancreatic beta cells. Analysis of human fetal pancreas samples showed the presence of FXYD2γa at an early stage (15 weeks). Histological examination of pancreatic sections from individuals with type 1 diabetes or sections from pancreases of streptozotocin-treated Macacus cynomolgus monkeys indicated a close correlation between loss of FXYD2γa and loss of insulin-positive cells.
Conclusions/interpretation
We propose human FXYD2γa as a novel beta cell-specific biomarker.
Similar content being viewed by others
Introduction
Both type 1 diabetes and type 2 diabetes are characterised by a reduced functional beta cell mass (BCM) [1–3]. Reduction of the BCM in type 1 diabetes is the consequence of a sustained autoimmune attack, while decreased beta cell mass in type 2 diabetes is less pronounced and probably secondary to prolonged metabolic stress [1–3]. The ability to measure the BCM prospectively in diabetic patients would be an important asset in understanding the natural history and pathogenesis of diabetes and would accelerate the development of new therapies for the disease. Unfortunately, there are no available tools for reliable and non-invasive in vivo BCM imaging [4]. The major difficulties in imaging the BCM are that islets are small (50–300 µm), constitute only 1–2% of the total pancreatic mass and are scattered in the exocrine tissue [5]. Pancreatic islets are composed of different cell types and beta cells have a common origin with other islet endocrine cells, acinar cells and the pancreatic ductal cells, making it difficult to find beta cell-specific biomarkers.
In vivo imaging of the BCM will require a combination of high sensitivity and specificity. Positron emission tomography (PET) and single photon emission computed tomography have high sensitivity and do not require ex vivo labelling. Imaging is achieved using either islet-specific binding compounds or by the use of radioactively labelled compounds specifically taken up by transporters in the beta cells. Ideally, two basic criteria should be met for adequate beta cell imaging [6]: (1) beta cells should retain the tracer 100- to 1,000-fold more than the exocrine cells; and (2) agents that label beta cell surface proteins should do this at high enough concentrations to overcome imaging signals arising from unbound tracer retained in the extracellular and vascular spaces, and from tracer bound to the other tissues surrounding the pancreas. Agents being tested currently for BCM imaging include glibenclamide [6], glucagon-like peptide 1 receptor [7, 8], sulfonylurea receptor [6], vesicular monoamine transporter 2 (VMAT2) [9–13] and gangliosides [14]. Unfortunately, the uptake/binding of these agents to beta cells, compared with exocrine pancreas and non-beta cells, is insufficient to allow reliable imaging of the BCM [6, 15]. For instance, the most promising beta cell marker, VMAT2, is produced in beta cells and monoaminergic neurons [10] but recent studies indicate that VMAT2 is also present in other islet cells besides beta cells [13]. Promising preliminary data were obtained with 11C-labelled dihydrotetrabenazine (the PET ligand binding specifically to VMAT2) in streptozotocin (STZ)-induced diabetic Lewis rats [11] and in the BB-DP rat [12], but the data obtained in baboons [15] and in patients with longstanding type 1 diabetes [16] are not conclusive. A recent study using an 18F-labelled epoxide derivative of tetrabenazine showed good pancreas specificity and significantly reduced uptake in rat liver [9], but it remains unclear whether substituting isotopes will produce better biodistribution results in larger mammals. As a whole, these observations emphasise the need for the identification of new beta cell biomarkers.
Here we used a systems biology approach on human islet massive parallel signal sequencing (MPSS) data sets and Affymetrix microarray data sets obtained in purified primary rat islet beta cells and non-beta cells. These studies identified FXYD domain containing ion transport regulator 2 (FXYD2)γa, a regulatory subunit of the Na+–K+-ATPase [17–19], as a novel and specific beta cell membrane protein.
Methods
Tissue samples
Human tissue samples were obtained with institutional review board approval from the Erasme Hospital (Université Libre de Bruxelles, Belgium), the University Hospital Dijkzigt (University Medical Center Rotterdam, The Netherlands) and the Churchill Hospital (Oxford, UK). Pancreas tissue sections from three patients with type 1 diabetes (time since diagnosis of type 1 diabetes, 31 years, 5 years and 3 days respectively; age at time of death, 62, 22 and 7 years) were obtained through the Oxford Centre for Diabetes (Churchill Hospital). Human fetal pancreas tissue samples (15, 18, 19, 24 and 25 weeks) were kindly provided by R. de Krijger (University Medical Center Rotterdam) to the Cell Differentiation Unit, Diabetes Research Center (Vrije Universiteit Brussel, Brussels, Belgium).
Pancreas sections from adult macaques (Macacus cynomolgus) treated or untreated with the diabetogenic drug STZ were obtained from the Laboratory of Experimental Surgery (Université Catholique de Louvain, Louvain-la-Neuve, Belgium). To induce diabetes (defined as a fasting blood glucose concentration >8.3 mmol/l), a low dose of filter-sterilised STZ (50 mg/kg body weight; Sigma, Bornem, Belgium) was administered intravenously over 5 min through the jugular vein [20]. One month after STZ injection, STZ-treated animals had significantly higher values for fasting blood glucose concentration (12.0 ± 2.7 vs 3.3 ± 0.5 mmol/l, p < 0.005), polyuria (609 ± 138 vs 90 ± 18 ml/24 h, p < 0.005), polydipsia (1,167 ± 147 vs 342 ± 17 ml/24 h, p < 0.005) and glycosuria (10 g/l vs no glucose) compared with animals not treated with STZ [20].
Animal care and welfare
All animal experiments were performed in accordance with accepted principles of good laboratory animal care and according to protocols approved by the Université Libre de Bruxelles (rodent experiments) and Université Catholique de Louvain (primate studies) Animal Care and Use Committees.
Screening of beta cell-specific biomarkers
Massively parallel signature sequencing (MPSS) data sets of two human islet preparations from healthy donors [21] were compared with a published data set of 32 normal human tissues [22] (complete data set available from Ludwig Institute for Cancer Research, http://mpss.licr.org). Mean signature counts for specific genes in islet samples were compared with counts for the same genes in other tissues: signatures that differed between islets and the other tissues by at least 4 SD of the mean and with a minimum count of 50 transcripts per million (tpm) were retained as ‘pancreatic islet-enriched genes’. This list of 883 genes enriched in pancreatic islets was introduced into TMHMM [23] and Ingenuity Pathway Analysis (IPA) 5.0 [24, 25] to select genes encoding membrane proteins. In parallel, Affymetrix (Santa Clara, CA, USA) microarray data obtained in primary FACS-purified rat islet beta cells and non-beta cells [21, 26–28] were compared to obtain selected beta cell-specific genes encoding membrane proteins.
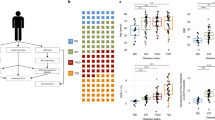
Cytokines produced locally in the course of insulitis may influence the production of beta cell proteins [3, 29] and thus induce a misleading evaluation of BCM. Therefore we compared the MPSS and microarray data with previous microarray data sets posted in the Beta Cell Gene Bank [30], including comparisons between cytokine-treated and control human islets [31], primary rat beta cells [26, 27] and INS-1E cells [32]. Based on these data, and following the selection procedure described in Fig. 1, we identified 12 beta cell-specific membrane proteins that are not significantly modified by cytokine treatment.
Selection procedure for screening beta cell-specific membrane biomarkers. MPSS data sets obtained in control and cytokine (IL1-β + IFN-γ)-exposed human islets [21] were compared with LICR MPSS data sets obtained in 32 human tissues [22], including total pancreas. Microarray data (included in the Beta Cell Gene Bank [30]) obtained in INS-1E cells, primary rat beta cells or human islets either in the control condition or exposed to cytokines were compared with data sets available on the Internet (Symatlas, GEO Omnibus). After this step, 950 islet-specific genes were selected and were further tested for membrane localisation using the TMHMM and IPA programs; this resulted in the selection of 114 plasma membrane proteins. By comparing microarray data sets obtained on rat primary beta cells and non-beta cells, we finally selected 44 beta cell-specific membrane proteins. Genes whose expression was modified by cytokines (>50%) were removed, resulting in a final list of 12 beta cell-specific membrane proteins. BGGB, Beta Cell Gene Bank ; Ctrl, control; CYT, cytokines ; LICR, Ludwig Institute for Cancer Research
FXYD2 splice variants produced in pancreatic islets
Poly(A) + RNA was isolated from mouse and human pancreatic islets using oligo(dT)-coated polystyrene Dynabeads (Dynal, Oslo, Norway), and reverse-transcribed as previously described [32, 33]. All primers [34] are described in the Electronic supplementary material (ESM).
Monoclonal antibody and development of polyclonal antibodies directed against tissue-restricted human splice variants
A monoclonal antibody directed against human FXYD2γb (1C3-B3) was purchased from Abnova (Taipei, Taiwan). Rabbit polyclonal antibodies were produced against the two islet-specific human membrane proteins of FXYD2 following the Super Speedy protocol (Eurogentec, Seraing Belgium). For this purpose, the following splice variant-specific peptides in the N-terminal extracellular domain of human FXYD2 were chosen: EP070936 (MTGLSMDGGGS) for FXYD2γa and EP070937 (MDRWYLGGS) for FXYD2γb. Two rabbits were injected per peptide and rabbit polyclonal antibodies were affinity-purified against the respective peptides. Using this approach, novel rabbit polyclonal antibodies directed against human FXYD2γa (SPY393) and FXYD2γb (SPY394), were developed.
Splice variant specificity was confirmed by ELISA and by western blotting, including testing with the respective blocking peptides (ESM Fig. 1b).
Western blotting
Total protein extracts were prepared from rat kidney and pancreatic islets, resolved on Tricine–SDS gels (15% polyacrylamide) and transferred to polyvinylidene fluoride membranes. SPY393 polyclonal antibody (1:1,000; Eurogentec) was used to detect FXYD2γa. The secondary antibody used was a horseradish peroxidase-conjugated donkey anti-rabbit antibody (1:5,000; Amersham Life Sciences, Amersham, UK). Membranes were incubated with the detection system of enhanced chemiluminescence (ECL; Supersignal West Femto Maximum Sensitivity Substrate; Pierce, Rockford, IL, USA). The protein-specific signals were identified using Biomax 1D image analysis software (Eastman Kodak, Rochester, NY, USA).
Immunohistochemistry
Paraffin-embedded human adult tissues (3 µm) were analysed with either 0.5 µg/ml monoclonal 1C3-B3 anti-FXYD2 or 7.5 µg/ml polyclonal SPY393 antibodies after antigen retrieval in citrate buffer, as described in Supplementary methods in the ESM.
Tissue microarray (human and macaque)
A human tissue microarray (TMA) was constructed with tissues from 36 organs with normal histology (with an average of three cases per organ and four spots per case). A normal pancreas TMA was constructed with tissue from 27 patients, each with ten spots (160 of these spots contained pancreatic islets).
Ten tissues were used for the construction of the macaque TMA.
Quantification and statistical analysis
Sections of pancreas from non-diabetic patients were analysed by immunofluorescence. Positive staining for insulin, glucagon and FXYD2γa was quantified by three observers unaware of sample identity and colocalisation between insulin and FXYD2γa and between glucagon and FXYD2γa was scored.
Sections of pancreas from non-diabetic and STZ-treated macaques were stained for IHC with SPY393, insulin or glucagon antibodies. After identification with the avidin–biotin–peroxidase complex method, six islets per case were quantified. Quantification was done with Spot Browser V2e software (Alphelys, Plaisir, France). Three observers, unaware of sample identity, scored for the presence of insulin, glucagon and FXYD2γa.
Results
A systems biology approach identifies 12 beta cell-specific membrane markers
To identify novel beta cell biomarkers, we used a step-by-step approach (Fig. 1) based on unique functional genomics data sets obtained by our group using either MPSS in human islets [21] or Affymetrix microarray analysis comparing purified primary rat beta cells against islet non-beta cells [21, 26]. Potential biomarkers were selected using the following criteria:
-
1.
Islet specificity criteria based on comparison of quantitative MPSS data from human islets against MPSS data from 32 human tissues. We used as the cut-off point >50-fold enrichment of expression in islets vs whole pancreas.
-
2.
Beta cell specificity criteria, i.e. higher gene expression in islet beta than in alpha cells.
-
3.
Expression levels should be higher than or comparable to glucokinase, an enzyme produced in beta cells at a ratio of 9 tpm in the MPSS data set.
-
4.
Selected biomarkers should be located in the plasma membrane. We first used the TMHMM program to detect transmembrane regions and then selected plasma membrane proteins using the Ingenuity Pathway Analysis program, which was followed by a thorough literature search.
-
5.
Expression levels of the selected markers should not be modified to any major extent by inflammation, as evaluated by our microarray data sets obtained in purified primary rat beta cells [26, 28, 30, 32] and in human islets [31].
Using this selection procedure (Fig. 1), we identified 12 beta cell-specific membrane proteins that were not significantly modified by cytokine treatment. These candidate biomarkers were further evaluated as described below. Nine of the 12 genes were not detected in total pancreas MPSS data, while all of them have expression levels higher than 88 tpm in human islets. Two genes (FXYD2, GNAS/XLAS and GNAS/ALEX—the last two genes are both annotated to GNAS loci in the MPSS data) showed 500-fold and 1,280-fold enrichment, respectively, in islet vs total pancreas MPSS data. After this step, four candidates (ESM Table 1) were selected for further validation at the protein level regarding islet specificity in the human pancreas; this was done by immunohistochemistry and by analysis of human TMA (data not shown for XLAS, ALEX and annexin 7 [ANXA7]). FXYD2 was selected as the best candidate and only results regarding FXYD2 are presented in this paper.
Validation of the candidate biomarker in a human tissue microarray
A monoclonal antibody directed against the full-length recombinant human FXYD2 (1C3-B3; Abnova) detected two bands with molecular mass 20 and 6.9 kDa respectively in rat pancreatic islets and in kidney (data not shown). In the human TMA, FXYD2 was present in the pancreas as well as faintly in the kidney tubules (ESM Table 2). Similar results were found when analysing a macaque TMA (data not shown). Analysis of the normal pancreas sections showed the presence of FXYD2 specifically in the islets (Fig. 2a, b). After this initial selection, subsequent work was focused on FXYD2, which was considered the most promising candidate beta cell biomarker.
FXYD2γa and FXYD2γb were present in human pancreatic islets but not in insulinomas. Normal human pancreas sections were analysed by immunohistochemistry with a monoclonal antibody directed against FXYD2 (Abnova) (magnification 200× in a and 400× in b) and with the polyclonal anti-FXYD2γa (SPY393) (magnification 200× in c and 400× in d). Consecutive sections of insulinoma (magnification 100×) were analysed with anti-insulin antibody (e) and SPY393 (f)
Human FXYD2γa is identified as a beta cell-specific protein
Three splice variants of FXYD2 have been reported in mouse kidney, namely FXYD2γa, -γb and -γc, differing only in the aminoterminal extracellular sequence. Using mouse Fxyd2 splice-specific PCR primers [34], we identified two of the three splice variants in mouse islets; namely, a fragment of 223 bp (Fxyd2γa) and a fragment of 211 bp (Fxyd2γb), while no fragment of 343 bp (Fxyd2γc) was detected (ESM Fig. 1a). Similar data were obtained using splice-specific PCR primers for the human FXYD2 in three different human islet preparations; namely, a fragment of 198 bp (FXYD2γa) and a fragment of 192 bp (FXYD2γb), but no fragment of 325 bp (FXYD2γc) (ESM Fig. 2a).
To further characterise the presence of these spliced variants, specific antibodies directed against the first 11 amino acids from the aminoterminal extracellular domain of FXYD2γa (SPY393; Eurogentec) and FXYD2γb (SPY394; Eurogentec) were produced. Both isoforms were detected as membrane and intracellular proteins in INS-1E cells and primary rat beta cells (data not shown). In western blotting, the antibody directed against the γa isoform (SPY393) detected a 20 kDa band in rat pancreatic islets (ESM Fig. 1b, lane 5) and kidney (ESM Fig. 1b, lanes 1, 2 and 6). The human FXYD2γa peptide, but not the FXYD2γb peptide, specifically blocked detection of this band by SPY393 (ESM Fig. 1b, lanes 3 and 4 and ESM Fig. 1b, lanes 5 and 6). In human TMA, FXYD2γa was restricted to the pancreas, while FXYD2γb was expressed both in the pancreas and in the kidney tubules (ESM Table 2). Therefore, further validation was performed only with the SPY393 antibody, which recognises the γa isoform.
Exposure of human islets or rat dispersed islet cells for 24 h to cytokines (IL1β + IFNγ) did not change the expression of FXYD2γa at either the mRNA or the protein level (ESM Fig. 2).
The aminoterminal sequence of FXYD2γa is identical in the human and other primates, but differs in four amino acids in the rat protein, which may explain the weak staining for FXYD2γa in rat islets compared with primate and human islets (Fig. 3). Histological examination of consecutive serial sections of human, macaque and rat pancreas with antibodies directed against insulin, glucagon and FXYD2γa demonstrated the presence of FXYD2γa in beta cells but not alpha cells (Fig. 3). Next, we performed immunofluorescence on human pancreas sections and made a quantitative analysis, observing the colocalisation of FXYD2γa with most insulin-positive beta cells; 86% of the beta cells showed the presence of FXYD2γa (Fig. 4a, b). On the other hand, fewer than 2% of the alpha cells were positive for FXYD2γa (Fig. 4c, d). Triple immunofluorescence labelling showed FXYD2γa on the cell membrane and in the cytoplasm of pancreatic beta cells (Fig. 4e, h).
FXYD2γa was specifically present in beta cells in human, macaque (Macacus cynomolgus) and rat pancreas. Consecutive paraffin sections of human (left column), macaque (middle column) and rat (right column) pancreas were analysed by immunohistochemistry with polyclonal rabbit anti-FXYD2γa (SPY393) (upper row), anti-insulin antibody (middle row) and anti-glucagon antibody (lower row). Magnification 400×
FXYD2γa was present in human pancreatic beta cells but not in alpha cells. Consecutive paraffin sections of normal adult human pancreas were analysed by immunofluorescence. a, b Colocalisation of FXYD2γa (red) with insulin (green) gives a yellow/orange colour (magnification 400×), (c, d) Lack of colocalisation between FXYD2γa (red) and glucagon (green) (magnification 400×). e–h Detection of FXYD2γa (e, green), insulin (f, red), (g) glucagon (g, blue) and triple immunofluorescence labelling showing that FXYD2γa was present on the membrane and in the cytoplasm of beta cells (h). Magnification 400×. Nuclei were stained with Hoechst
Six cases of human insulinomas were also analysed by immunohistochemistry. While FXYD2γa was present in the pancreatic islets of the remaining normal pancreas, its abundance was decreased or absent in the insulinomas (Fig. 2f and data not shown). To gain information on the presence of FXYD2γa during human pancreas development, samples from fetal human pancreas (15, 18, 19, 24 and 25 weeks of gestation) were analysed by triple immunofluorescence labelling (Fig. 5). FXYD2γa was present in beta cells from the earliest time point (15 weeks) onwards, first in single beta cells and clusters of beta cells and at the later time points (18–25 weeks) in islets. Labelling was also detected in cells that were not insulin-positive, suggesting that FXYD2γa production may precede insulin production (Fig. 5, 15 weeks, upper panel). Histological examination of consecutive serial sections of human fetal pancreas (19 and 25 weeks) with antibodies directed against somatostatin, pancreatic polypeptide and FXYD2γa showed no colocalisation with cells containing somatostatin and pancreatic polypeptide (Fig. 6).
Early presence of FXYD2 in human endocrine pancreas. Consecutive paraffin sections of fetal human pancreas (15, 18 and 25 weeks) were analysed by immunofluorescence. Green, FXYD2γa; red, insulin; blue, glucagon (blue); yellow/orange, colocalisation of FXYD2γa and insulin. There was no colocalisation of FXYD2γa with glucagon. Magnification 400×
FXYD2γa was present in human pancreatic beta cells but not in delta or pancreatic polypeptide cells. Consecutive paraffin sections of human pancreas were analysed by immunohistochemistry with anti-pancreatic polypeptide antibody (a) and polyclonal rabbit anti-FXYD2γa (SPY393) (b) or with anti-somatostatin antibody (c) and polyclonal rabbit anti-FXYD2γa (SPY393) (d). Magnification 1,000× in a and b, 400× in c and d
Decreased FXYD2γa in islets from patients with type 1 diabetes
Consecutive serial sections were analysed for the presence of cells containing glucagon, insulin and FXYD2γa in pancreas material obtained from three individuals with type 1 diabetes with different durations of the disease (Fig. 7). In an individual with 31 years of type 1 diabetes (patient 1, aged 62 years) the few remaining insulin-positive cells scattered over the pancreas were also FXYD2γa-positive, while in another individual with a 5 year history of type 1 diabetes (patient 2, aged 22 years) the absence of insulin-containing cells correlated with the absence of FXYD2γa. In a case of type 1 diabetes diagnosed 3 days before death (patient 3, aged 7 years) no insulin-positive cells were detected, but faint membrane-located FXYD2γa staining was still present in a few scattered cells, suggesting that FXYD2γa may still be detectable in degranulated beta cells. No FXYD2γa production was detected in the glucagon-producing cells present in the three patients (Fig. 7).
Decreased presence of FXYD2γa correlated with beta cell loss in three patients with type 1 diabetes. Consecutive sections of pancreas from an individual control (Ctrl) and three individual patients with type 1 diabetes. These patients had different durations of diabetes (3 years, 25 years and 3 days). The samples were analysed with polyclonal anti-FXYD2γa antibody (SPY393) (left column), anti-insulin antibody (middle column) or anti-glucagon antibody (right column)
FXYD2γa and insulin decreased in parallel in macaques treated with streptozotocin
The presence of FXYD2γa was analysed in ten macaque tissues: pancreas, kidney, spleen, lymph node, liver, lung, aorta, heart, muscle and fat. FXYD2γa was restricted to pneumocytes and to the pancreatic islets; weak positive staining was also detected in the kidney tubules (data not shown). Pancreas sections of six macaques treated with STZ to partially ablate BCM were evaluated for the presence of FXYD2γa, insulin and glucagon (quantification shown in Table 1). The loss of beta cells in the STZ-treated animals correlated well with the decreased presence of the FXYD2γa marker (ESM Figs 3 and 4; Table 1), while quantification of FXYD2γa in a non-STZ-treated control showed a close correlation between the percentages of insulin- and FXYD2γa-positive cells (55 ± 13 and 59 ± 9 respectively). In all STZ-treated animals there was a small increase in glucagon-positive cells compared with the control, but no FXYD2γa staining was detected in these glucagon-positive cells (ESM Figs 3 and 4; Table 1).
Discussion
We have developed a functional genomics-based strategy to identify beta cell-specific biomarkers with potential for non-invasive imaging of the BCM. For this purpose, we compared a unique human islet MPSS data set [21] against MPSS data sets obtained in 32 human tissues [22]. This allowed the identification of 114 islet-specific membrane genes with a minimum count of 50 tpm. The quantitative MPSS data used in the present study enabled us to calculate enrichment in human islets vs total pancreas. The mRNAs with the highest level of enrichment identified in this study (FXYD2, GNAS/XLAS, GNAS/ALEX and ANXA7; ESM Table 1 and data not shown) were selected for validation in human pancreas by immunohistochemistry and tested in a human tissue microarray containing 36 human tissues.
The best candidate beta cell biomarker identified, FXYD2, is a regulating subunit of the Na+–K+-ATPase. It was found to be 500-fold enriched in pancreatic islets compared with total pancreas. This gene has not been reported previously as being expressed in pancreatic islets. Using PCR, we confirmed that two out of the three known splice variants of FXYD2 [34] are expressed in human pancreatic islets, and then produced polyclonal antibodies targeting specifically these human splice variant proteins for subsequent studies. Histological examination of human pancreas confirmed the islet-specific localisation of FXYD2γa and FXYD2γb. These two splice variants were previously reported as being produced in kidney proximal tubules and in distal convoluted tubules respectively [17, 18]. As the present data indicate, the FXYD2γb subunit is present in both kidney and pancreatic islets, while FXYD2γa is only present in human pancreatic islets. Importantly, FXYD2γa is mostly produced in beta cells, but not in alpha, delta or pancreatic polypeptide cells. Analysis of human fetal samples indicates that FXYD2γa expression is an early embryonic event, which may precede insulin expression. On the other hand, tumoral transformation of beta cells in human insulinomas is paralleled by loss of FXYD2γa expression.
A set of 35 human islet membrane-associated proteins were previously identified in an elegant study by Maffei et al. [35] using Affymetrix oligonucleotide chips to compare expression profiles of human islets against exocrine pancreas, liver and kidney. Most of the membrane genes identified in that study were also present in our list. Of note, this previous study [35] detected neither FXYD2, the most promising marker so far identified, nor GNAS/ALEX or GNAS/XLAS. The probable reason, at least for FXYD2, is that the probe sets for this mRNA in the Affymetrix microarrays do not provide consistent results (D. Flamez and D. L. Eizirik, unpublished data).
In order to quantify the remaining BCM in type 1 diabetes, markers selected for imaging should not be present in the infiltrating mononuclear cells and there should be a good correlation between beta cell loss and disappearance of the marker. FXYD2 expression was not detected in lymphocytes or other immune cells in the Symatlas data set [36] and FXYD2γa expression was not significantly modified by cytokines at either the mRNA or the protein level. Furthermore, there were clear and parallel decreases in FXYD2γa and insulin staining in pancreas both from individuals with type 1 diabetes and from macaques treated with STZ.
As mentioned above, FXYD2γa is a regulating subunit of the ubiquitous protein Na+–K+-ATPase. Na+-K+-ATPase has two obligatory subunits, a catalytic α subunit and a glycoprotein β subunit. Regulation of this pump occurs at several levels: gene expression, recruitment and internalisation of active pump subunits to/from the plasma membrane, and modulation of ATPase activity. The intrinsic properties of the pump are regulated by association with tissue-specific small single-span membrane proteins belonging to the FXYD family. A human FXYD2 G41R mutation is responsible for renal hypomagnesaemia [37], while silencing and overexpression of the γ subunit of Na+-K+-ATPase affects survival of mouse kidney (IMCD3) cells exposed to hypertonic stress [38].
A recent study shows the association of hepatocyte nuclear factor 1-β (HNF1β) with renal hypomagnesaemia and suggests that HNF1β regulates the transcription of FXYD2 [39]. Little is known about the function of FXYD2γa and FXYD2γb in beta cells and future experiments, involving overexpression and knockdown of FXYD2γa and FXYD2γb in beta cells, should be performed to clarify their role in cell function and survival.
In conclusion, we have developed a systems biology approach, starting from MPSS and microarray data, for the identification of novel beta cell biomarkers. This, coupled with the systematic use of human tissue microarrays, allowed the identification of 12 proteins that are enriched in beta cells and have the potential to serve as biomarkers. Of special interest, the splice variant FXYD2γa was identified as beta cell-specific in both the human and a non-human primate model. This protein is not present in other islet cells, and its production decreases in parallel with beta cell loss in both human type 1 diabetes and in macaques rendered diabetic by injection of STZ. We propose that this newly identified beta cell protein, FXYD2γa, may be a useful biomarker for future non-invasive imaging and quantification of the pancreatic beta cell mass.
Abbreviations
- BCM:
-
Beta cell mass
- FXYD2:
-
FXYD domain containing ion transport regulator 2
- MPSS:
-
Massive parallel signal sequencing
- PET:
-
Positron emission tomography
- STZ:
-
Streptozotocin
- TMA:
-
Tissue microarray
- tpm:
-
Transcripts per million
- VMAT2:
-
Vesicular monoamine transporter 2
- LICR:
-
Ludwig Institute for Cancer Research
- BCGB:
-
Beta Cell Gene Bank
References
Butler AE, Janson J, Bonner-Weir S, Ritzel R, Rizza RA, Butler PC (2003) Beta-cell deficit and increased beta-cell apoptosis in humans with type 2 diabetes. Diabetes 52:102–110
Rahier J, Guiot Y, Goebbels RM, Sempoux C, Henquin JC (2008) Pancreatic beta-cell mass in European subjects with type 2 diabetes. Diabetes Obes Metab 10:32–42
Cnop M, Welsh N, Jonas JC, Jörns A, Lenzen S, Eizirik DL (2005) Mechanisms of pancreatic beta-cell death in type 1 and type 2 diabetes: many differences, few similarities. Diabetes 54:S97–S107
Martinic MM, von Herrath MG (2008) Real-time imaging of the pancreas during development of diabetes. Immunol Rev 221:200–213
Sweet IR, Cook DL, Lernmark A, Greenbaum CJ, Krohn KA (2004) Non-invasive imaging of beta cell mass: a quantitative analysis. Diabetes Technol Ther 6:652–659
Sweet IR, Cook DL, Lernmark A et al (2004) Systematic screening of potential beta-cell imaging agents. Biochem Biophys Res Commun 314:976–983
Gotthardt M, Lalyko G, van Eerd-Vismale J et al (2006) A new technique for in vivo imaging of specific GLP-1 binding sites: first results in small rodents. Regul Pept 137:162–167
Wild D, Béhé M, Wicki A et al (2006) [Lys40(Ahx-DTPA-111In)NH2]exendin-4, a very promising ligand for glucagon-like peptide-1 (GLP-1) receptor targeting. J Nucl Med 47:2025–2033
Kung MP, Hou C, Lieberman BP et al (2008) In vivo imaging of beta-cell mass in rats using 18F-FP-(+)-DTBZ: a potential PET ligand for studying diabetes mellitus. J Nucl Med 49:1171–1176
Harris PE, Ferrara C, Barba P, Polito T, Freeby M, Maffei A (2008) VMAT2 gene expression and function as it applies to imaging beta-cell mass. J Mol Med 86:5–16
Simpson NR, Souza F, Witkowski P et al (2006) Visualizing pancreatic beta-cell mass with [11C]DTBZ. Nucl Med Biol 33:855–864
Souza F, Simpson N, Raffo A et al (2006) Longitudinal noninvasive PET-based beta cell mass estimates in a spontaneous diabetes rat model. J Clin Invest 116:1506–1513
Saisho Y, Harris PE, Butler AE et al (2008) Relationship between pancreatic vesicular monoamine transporter 2 (VMAT2) and insulin expression in human pancreas. J Mol Histol 39:543–551
Ladrière L, Malaisse-Lagae F, Alejandro R, Malaisse WJ (2001) Pancreatic fate of a (125)I-labelled mouse monoclonal antibody directed against pancreatic B cell surface ganglioside(s) in control and diabetic rats. Cell Biochem Funct 19:107–115
Murthy R, Harris P, Simpson N et al (2008) Whole body [11C]-dihydrotetrabenazine imaging of baboons: biodistribution and human radiation dosimetry estimates. Eur J Nucl Med Mol Imaging 35:790–797
Goland R, Freeby M, Parsey R et al (2009) 11C-Dihydrotetrabenazine PET of the pancreas in subjects with long-standing type 1 diabetes and in healthy controls. J Nucl Med 50:382–389
Sweadner KJ, Arystarkhova E, Donnet C, Wetzel RK (2003) FXYD proteins as regulators of the Na, K-ATPase in the kidney. Ann N Y Acad Sci 986:382–387
Li C, Grosdidier A, Crambert G, Horisberger JD, Michielin O, Geering K (2004) Structural and functional interaction sites between Na, K-ATPase and FXYD proteins. J Biol Chem 279:38895–38902
Wetzel RK, Pascoa JL, Arystarkhova E (2004) Stress-induced expression of the gamma subunit (FXYD2) modulates Na, K-ATPase activity and cell growth. J Biol Chem 279:41750–41757
Dufrane D, van Steenberghe M, Guiot Y et al (2006) Streptozotocin-induced diabetes in large animals (pigs/primates): role of GLUT2 transporter and beta-cell plasticity. Transplantation 81:36–45
Kutlu B, Burdick D, Baxter D et al (2009) Detailed transcriptome atlas of the pancreatic beta cell. BMC Med Genomics 2:3
Jongeneel CV, Delorenzi M, Iseli C et al (2005) An atlas of human gene expression from massively parallel signature sequencing (MPSS). Genome Res 15:1007–1014
Krogh A, Larsson B, von Heijne G, Sonnhammer ELL (2001) Predicting transmembrane protein topology with a hidden Markov model: application to complete genomes. J Mol Biol 305:567–580
Gusev Y (2008) Computational methods for analysis of cellular functions and pathways collectively targeted by differentially expressed microRNA. Methods 44:61–72
Eizirik DL, Moore F, Flamez D, Ortis F (2008) Use of a systems biology approach to understand pancreatic beta-cell death in type 1 diabetes. Biochem Soc Trans 36:321–327
Cardozo AK, Kruhoffer M, Leeman R, Orntoft T, Eizirik DL (2001) Identification of novel cytokine-induced genes in pancreatic beta-cells by high-density oligonucleotide arrays. Diabetes 50:909–920
Cardozo AK, Heimberg H, Heremans Y et al (2001) A comprehensive analysis of cytokine-induced and nuclear factor-κ B-dependent genes in primary rat pancreatic beta-cells. J Biol Chem 276:48879–48886
Rasschaert J, Liu D, Kutlu B et al (2003) Global profiling of double stranded RNA- and IFN-γ-induced genes in rat pancreatic beta cells. Diabetologia 46:1641–1657
D’Hertog W, Overbergh L, Lage K et al (2007) Proteomic analysis of cytokine-induced dysfunction and death in insulin-producing INS-1E cells: new insights into the pathways involved. Mol Cell Proteomics 6:2180–2199
Smink LJ, Helton EM, Healy BC et al (2005) T1DBase, a community web-based resource for type 1 diabetes research. Nucleic Acids Res 33:D544–D549
Ylipaasto P, Kutlu B, Rasilainen S et al (2005) Global profiling of coxsackievirus- and cytokine-induced gene expression in human pancreatic islets. Diabetologia 48:1510–1522
Kutlu B, Cardozo AK, Darville MI et al (2003) Discovery of gene networks regulating cytokine-induced dysfunction and apoptosis in insulin-producing INS-1 cells. Diabetes 52:2701–2719
Rasschaert J, Ladrière L, Urbain M et al (2005) Toll-like receptor 3 and STAT-1 contribute to double-stranded RNA + interferon-g-induced apoptosis in primary pancreatic beta-cells. J Biol Chem 280:33984–33991
Jones DH, Golding MC, Barr KJ, Fong GH, Kidder GM (2001) The mouse Na+-K + -ATPase gamma-subunit gene (Fxyd2) encodes three developmentally regulated transcripts. Physiol Genomics 6:129–135
Maffei A, Liu Z, Witkowski P et al (2004) Identification of tissue-restricted transcripts in human islets. Endocrinology 145:4513–4521
Gurkan C, Lapp H, Hogenesch JB, Balch WE (2005) Exploring trafficking GTPase function by mRNA expression profiling: use of the SymAtlas web-application and the Membrome datasets. Methods Enzymol 403:1–10
Sha Q, Pearson W, Burcea LC et al (2008) Human FXYD2 G41R mutation responsible for renal hypomagnesemia behaves as an inward-rectifying cation channel. Am J Physiol Renal Physiol 295:F91–F98
Capasso JM, Rivard CJ, Berl T (2006) Silencing and overexpression of the gamma-subunit of Na-K-ATPase directly affect survival of IMCD3 cells in response to hypertonic stress. Am J Physiol Renal Physiol 291:F1142–F1147
Adalat S, Woolf AS, Johnstone KA et al (2009) HNF1B mutations associate with hypomagnesemia and renal magnesium wasting. J Am Soc Nephrol 20:1123–1131
Acknowledgements
We thank M. A. Neef, M. Urbain and J. Schoonheydt of the Laboratory of Experimental Medicine (ULB) for excellent technical support. We also thank A. Verrellen and S. Kalista (Laboratory of Pathology, Erasme Hospital) for their assistance in the quantification analysis of the immunohistochemistry data. This work was supported by grants from the Pôles de compétitivité de la région Wallonne as part of the KeyMarker–BioWin programme (BioWin Convention No. 5437), European Union (BetaImage, Health-F2-2008-222980; in the Framework Programme 7 of the European Community), Fonds National de la Recherche Scientifique (FNRS) Belgium, the Belgian Program on Interuniversity Poles of Attraction initiated by the Belgian State (IUAP P6/40), Juvenile Diabetes Research Foundation (JDRF) programme project (4-2001-438) and JDRF Bridge Support for Systems Biology of the Beta Cell (4-2008-376). B.K. is a recipient of a JDRF Postdoctoral Fellowship (3-2006-306).
Duality of interest
The biomarker described in this article is the subject of a patent deposited by Université Libre de Bruxelles (ULB), the Institute for Systems Biology (ISB) and Eurogentec SA.
Author information
Authors and Affiliations
Corresponding author
Additional information
D. Flamez and I. Roland contributed equally to this study.
Electronic supplementary materials
Below is the link to the electronic supplementary material.
ESM Table 1
Selection of four beta cell-specific biomarkers (PDF 13 kb)
ESM Table 2
FXYD2a and FXYD2b are preferentially expressed in human pancreas (PDF 87 kb)
ESM Fig. 1
Expression of Fxyd2γa and -γb isoforms in rodent pancreatic islets. a Fxyd2γa and -γb splice variants were detected in mouse pancreatic islets by RT-PCR. Primers were used to detect Fxyd2γa isoform (lane 1; 223 bp), Fxyd2γb isoform (lane 2; 211 bp) and Fxyd2γc isoform (lane 3; 343 bp); marker (M; lane 4). b Polyclonal anti-FXYD2γa antibody (SPY393) recognised only the FXYD2γa isoform in western blotting. Rat dispersed pancreatic islet cells (lanes 3 and 5) and rat kidney (lanes 1, 2, 4 and 6) were analysed. Lane 1, SPY393 with FXYD2γb blocking peptide; lane 2, SPY393 with a non-specific blocking peptide; lanes 3 and 4, SPY393 with FXYD2γa blocking peptide; lanes 5 and 6, SPY393 without blocking peptide (PDF 54 kb)
ESM Fig. 2
Fxyd2 γa expression was not modified by 24h of treatment with cytokines IL1β + IFNγ. a An Fxyd2γa splice variant (198 bp) was detected in human pancreatic islets by RT-PCR. Lanes 1–3, human islet preparations under control condition; lanes 4–6, human islets treated with cytokines for 24h. β-Actin primers were used as internal control. b Polyclonal anti-FXYD2γa antibody (SPY393) recognised only the FXYD2γa isoform in western blotting. Dispersed rat pancreatic islet cells (lane 1, control condition); lane 2 cytokine treated (IL1β + IFNγ, 24h) were analysed (PDF 97 kb)
ESM Fig. 3
Decreased FXYD2γa correlates with beta cell loss in STZ-treated macaques. Consecutive sections of pancreas from control (a–c, CTRL) and STZ-treated macaques(d–f, animal 1; g–i, animal 2) were analysed with SPY393 polyclonal anti-FXYD2γa (a, d, g), anti-insulin antibody (b, e, h) and anti-glucagon antibody (c, f, i). Magnification 400 (PDF 322 kb)
ESM Fig. 4
Significant decrease in insulin and FXYD2γa expression in the pancreas of STZ-induced diabetic macaques. Pancreas sections from control (CTRL) and STZ-induced diabetic macaques (P1 -6) were analysed. Six islets per animal were counted by three observers unaware of sample identity. Cells positive for glucagon, insulin and FXYD2γa were calculated as relative percentages per islet. Values for the control were considered as 100% (PDF 25 kb)
ESM 1
(PDF 59 kb)
Rights and permissions
About this article
Cite this article
Flamez, D., Roland, I., Berton, A. et al. A genomic-based approach identifies FXYD domain containing ion transport regulator 2 (FXYD2)γa as a pancreatic beta cell-specific biomarker. Diabetologia 53, 1372–1383 (2010). https://doi.org/10.1007/s00125-010-1714-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00125-010-1714-z