Abstract
Key message
A major yellow-seed QTL on chromosome A09 significantly increases the oil content and reduces the fiber content of seed in Brassica napus.
Abstract
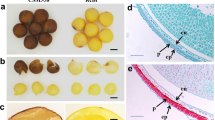
The yellow-seed trait (YST) has always been a main breeding objective for rapeseed because yellow-seeded B. napus generally contains higher oil contents, fewer pigments and polyphenols and lower fiber content than black-seeded B. napus, although the mechanism controlling this correlation remains unclear. In this study, QTL mapping was implemented for YST based on a KN double haploid population derived from the hybridization of yellow-seeded B. napus N53-2 with a high oil content and black-seeded Ken-C8 with a relatively low oil content. Ten QTLs were identified, including four stable QTLs that could be detected in multiple environments. A major QTL, cqSC-A09, on chromosome A09 was identified by both QTL mapping and BSR-Seq technology, and explained more than 41% of the phenotypic variance. The major QTL cqSC-A09 for YST not only controls the seed color but also affects the oil and fiber contents in seeds. More importantly, the advantageous allele could increase the oil content and reduce the pigment and fiber content at the same time. This is the first QTL reported to control seed color, oil content and fiber content simultaneously with a large effect and has great application value for breeding high oil varieties with high seed quality. Important candidate genes, including BnaA09. JAZ1, BnaA09. GH3.3 and BnaA09. LOX3, were identified for cqSC-A09 by combining sequence variation annotation, expression differences and an interaction network, which lays a foundation for further cloning and breeding applications in the future.






Similar content being viewed by others
Data availability
The datasets generated and/or analyzed during the current study are available from the corresponding author on reasonable request.
References
Abbadi A, Leckband G (2011) Rapeseed breeding for oil content, quality, and sustainability. Eur J Lipid Sci Tech 113:1198–1206
Akhov L, Ashe P, Tan YF, Datla R, Selvaraj G (2009) Proanthocyanidin biosynthesis in the seed coat of yellow-seeded, canola quality Brassica napus YN01-429 is constrained at the committed step catalyzed by dihydroflavonol 4-reductase. Botany 87:616–625
Arcade A, Labourdette A, Falque M, Mangin B, Chardon F, Charcosset A, Joets J (2004) BioMercator: integrating genetic maps and QTL towards discovery of candidate genes. Bioinformatics 20:2324–2326
Auger B, Marnet N, Gautier V, Maia-Grondard A, Leprince F, Renard M, Guyot S, Nesi N, Routaboul JM (2010) A detailed survey of seed coat flavonoids in developing seeds of Brassica napus L. J Agr Food Chem 58:6246–6256
Badani AG, Snowdon RJ, Wittkop B, Lipsa FD, Baetzel R, Horn R, De Haro A, Font R, Luhs W, Friedt W (2006) Colocalization of a partially dominant gene for yellow seed colour with a major QTL influencing acid detergent fibre (ADF) content in different crosses of oilseed rape (Brassica napus). Genome 49:1499–1509
Behnke N, Suprianto E, Mollers C (2018) A major QTL on chromosome C05 significantly reduces acid detergent lignin (ADL) content and increases seed oil and protein content in oilseed rape (Brassica napus L.). Theor Appl Genet 131:2477–2492
Buer CS, Djordjevic MA (2009) Architectural phenotypes in the transparent testa mutants of Arabidopsis thaliana. J Exp Bot 60:751–763
Cai GQ, Yang QY, Yi B, Fan CC, Edwards D, Batley J, Zhou YM (2014) A complex recombination pattern in the genome of allotetraploid Brassica napus as revealed by a high-density genetic map. PLoS ONE 9:e109910
Chalhoub B, Denoeud F, Liu S, Parkin IA, Tang H, Wang X, Chiquet J, Belcram H, Tong C, Samans B, Correa M, Da Silva C, Just J, Falentin C, Koh CS, Le Clainche I, Bernard M, Bento P, Noel B, Labadie K, Alberti A, Charles M, Arnaud D, Guo H, Daviaud C, Alamery S, Jabbari K, Zhao M, Edger PP, Chelaifa H, Tack D, Lassalle G, Mestiri I, Schnel N, Le Paslier MC, Fan G, Renault V, Bayer PE, Golicz AA, Manoli S, Lee TH, Thi VH, Chalabi S, Hu Q, Fan C, Tollenaere R, Lu Y, Battail C, Shen J, Sidebottom CH, Wang X, Canaguier A, Chauveau A, Berard A, Deniot G, Guan M, Liu Z, Sun F, Lim YP, Lyons E, Town CD, Bancroft I, Wang X, Meng J, Ma J, Pires JC, King GJ, Brunel D, Delourme R, Renard M, Aury JM, Adams KL, Batley J, Snowdon RJ, Tost J, Edwards D, Zhou Y, Hua W, Sharpe AG, Paterson AH, Guan C, Wincker P (2014) Plant genetics. Early allopolyploid evolution in the post-Neolithic Brassica napus oilseed genome. Science 345:950–953
Chao H, Raboanatahiry N, Wang X, Zhao W, Chen L, Guo L, Li B, Hou D, Pu S, Zhang L, Wang H, Wang B, Li M (2019) Genetic dissection of harvest index and related traits through genome-wide quantitative trait locus mapping in Brassica napus L. Breed Sci 69:104–116
Chao H, Wang H, Wang X, Guo L, Gu J, Zhao W, Li B, Chen D, Raboanatahiry N, Li M (2017) Genetic dissection of seed oil and protein content and identification of networks associated with oil content in Brassica napus. Sci Rep 7:46295
Chen LQ, Lin IWN, Qu XQ, Sosso D, McFarlane HE, Londono A, Samuels AL, Frommer WB (2015) A cascade of sequentially expressed sucrose transporters in the seed coat and endosperm provides nutrition for the Arabidopsis embryo. Plant Cell 27:607–619
Chen MX, Xuan LJ, Wang Z, Zhou LH, Li ZL, Du X, Ali E, Zhang GP, Jiang LX (2014) TRANSPARENT TESTA8 inhibits seed fatty acid accumulation by targeting several seed development regulators in Arabidopsis. Plant Physiol 165:905–916
Cubillos FA, Brice C, Molinet J, Tisne S, Abarca V, Tapia SM, Oporto C, Garcia V, Liti G, Martinez C (2017) Identification of nitrogen consumption genetic variants in yeast through QTL mapping and bulk segregant RNA-Seq analyses. G3 Genes Genom Genet 7:1693–1705
Debeaujon I, Leon-Kloosterziel KM, Koornneef M (2000) Influence of the testa on seed dormancy, germination, and longevity in Arabidopsis. Plant Physiol 122:403–413
Du HW, Zhu JX, Su H, Huang M, Wang HW, Ding SC, Zhang BL, Luo A, Wei SD, Tian XH, Xu YB (2017) Bulked segregant RNA-seq reveals differential expression and SNPs of candidate genes associated with waterlogging tolerance in Maize. Front Plant Sci 8:1022
Fu FY, Liu LZ, Chai YR, Chen L, Yang T, Jin MY, Ma AF, Yan XY, Zhang ZS, Li JN (2007) Localization of QTLs for seed color using recombinant inbred lines of Brassica napus in different environments. Genome 50:840–854
Gacek K, Bayer PE, Anderson R, Severn-Ellis AA, Wolko J, Lopatynska A, Matuszczak M, Bocianowski J, Edwards D, Batley J (2021) QTL genetic mapping study for traits affecting meal quality in winter oilseed rape (Brassica napus L.). Genes (basel) 12:1235
Gu AX, Meng C, Chen YQ, Wei L, Dong H, Lu Y, Wang YH, Chen XP, Zhao JJ, Shen SX (2017) Coupling Seq-BSA and RNA-Seq analyses reveal the molecular pathway and genes associated with heading type in chinese cabbage. Front Genet 8:176
Huang H, Gao H, Liu B, Fan M, Wang JJ, Wang CL, Tian HX, Wang LX, Xie CY, Wu DW, Liu LY, Yan JB, Qi TC, Song SS (2018) bHLH13 regulates jasmonate-mediated defense responses and growth. Evol Bioinform 14:1176934318790265
Jiang CC, Shi JQ, Li RY, Long Y, Wang H, Li DR, Zhao JY, Meng JL (2014) Quantitative trait loci that control the oil content variation of rapeseed (Brassica napus L.). Theor Appl Genet 127:957–968
Jiang JJ, Zhu S, Yuan Y, Wang Y, Zeng L, Batley J, Wang YP (2019) Transcriptomic comparison between developing seeds of yellow- and black-seeded Brassica napus reveals that genes influence seed quality. BMC Plant Biol 19:203
Lepiniec L, Debeaujon I, Routaboul JM, Baudry A, Pourcel L, Nesi N, Caboche M (2006) Genetics and biochemistry of seed flavonoids. Annu Rev Plant Biol 57:405–430
Li B, Zhao W, Li D, Chao H, Zhao X, Ta N, Li Y, Guan Z, Guo L, Zhang L, Li S, Wang H, Li M (2018) Genetic dissection of the mechanism of flowering time based on an environmentally stable and specific QTL in Brassica napus. Plant Sci 277:296–310
Liu CL, Zhou Q, Dong L, Wang H, Liu F, Weng JF, Li XH, Xie CX (2016) Genetic architecture of the maize kernel row number revealed by combining QTL mapping using a high-density genetic map and bulked segregant RNA sequencing. BMC Genom 17:915
Liu L, Stein A, Wittkop B, Sarvari P, Li J, Yan X, Dreyer F, Frauen M, Friedt W, Snowdon RJ (2012a) A knockout mutation in the lignin biosynthesis gene CCR1 explains a major QTL for acid detergent lignin content in Brassica napus seeds. Theor Appl Genet 124:1573–1586
Liu LZ, Qu CM, Wittkop B, Yi B, Xiao Y, He YJ, Snowdon RJ, Li JN (2013) A high-density SNP map for accurate mapping of seed fibre QTL in Brassica napus L. PLoS ONE 8:e83052
Liu SZ, Yeh CT, Tang HM, Nettleton D, Schnable PS (2012b) Gene mapping via bulked segregant RNA-Seq (BSR-Seq). PLoS ONE 7:e36406
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2(T)(-Delta Delta C) method. Methods 25:402–408
Mccouch S, Cho Y, Yano M, Paul E, Blinstrub M, Morishima H, Mccouch S, Cho Y, Paul E, Morishima H (1997) Report on QTL nomenclature. Rice Genet Newsl 14:11
Miao L, Chao H, Chen L, Wang H, Zhao W, Li B, Zhang L, Li H, Wang B, Li M (2019) Stable and novel QTL identification and new insights into the genetic networks affecting seed fiber traits in Brassica napus. Theor Appl Genet 132:1761–1775
Mittasch J, Bottcher C, Frolov A, Strack D, Milkowski C (2013) Reprogramming the phenylpropanoid metabolism in seeds of oilseed rape by suppressing the orthologs of REDUCED EPIDERMAL FLUORESCENCE1. Plant Physiol 161:1656–1669
Niu Y, Wu LM, Li YH, Huang HL, Qian MC, Sun W, Zhu H, Xu YF, Fan YH, Mahmood U, Xu BB, Zhang K, Qu CM, Li JN, Lu K (2020) Deciphering the transcriptional regulatory networks that control size, color, and oil content in Brassica rapa seeds. Biotechnol Biofuels 13:90
Raboanatahiry N, Chao H, Guo L, Gan J, Xiang J, Yan M, Zhang L, Yu L, Li M (2017) Synteny analysis of genes and distribution of loci controlling oil content and fatty acid profile based on QTL alignment map in Brassica napus. BMC Genomics 18:776
Rahman MH, Joersbo M, Poulsen MH (2001) Development of yellow-seeded Brassica napus of double low quality. Plant Breeding 120:473–478
Shannon P, Markiel A, Ozier O, Baliga NS, Wang JT, Ramage D, Amin N, Schwikowski B, Ideker T (2003) Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res 13:2498–2504
Shin J, Park E, Choi G (2007) PIF3 regulates anthocyanin biosynthesis in an HY5-dependent manner with both factors directly binding anthocyanin biosynthetic gene promoters in Arabidopsis. Plant J 49:981–994
Snowdon RJ, Wittkop B, Rezaidad A, Hasan M, Lipsa F, Stein A, Friedt W (2010) Regional association analysis delineates a sequenced chromosome region influencing antinutritive seed meal compounds in oilseed rape. Genome 53:917–928
Stein A, Wittkop B, Liu LZ, Obermeier C, Friedt W, Snowdon RJ (2013) Dissection of a major QTL for seed colour and fibre content in Brassica napus reveals colocalization with candidate genes for phenylpropanoid biosynthesis and flavonoid deposition. Plant Breed 132:382–389
Stone SL, Braybrook SA, Paula SL, Kwong LW, Meuser J, Pelletier J, Hsieh TF, Fischer RL, Goldberg RB, Harada JJ (2008) Arabidopsis LEAFY COTYLEDON2 induces maturation traits and auxin activity: Implications for somatic embryogenesis. P Natl Acad Sci USA 105:3151–3156
Wang B, Wu ZK, Li ZH, Zhang QH, Hu JL, Xiao YJ, Cai DF, Wu JS, King GJ, Li HT, Liu KD (2018) Dissection of the genetic architecture of three seed-quality traits and consequences for breeding in Brassica napus. Plant Biotech J 16:1336–1348
Wang J, Jian HJ, Wei LJ, Qu CM, Xu XF, Lu K, Qian W, Li JN, Li MT, Liu LZ (2015) Genome-wide analysis of seed acid detergent lignin (ADL) and hull content in rapeseed (Brassica napus L.). PLoS ONE 10:e0145045
Wang J, Xian X, Xu X, Qu C, Lu K, Li J, Liu L (2017) Genome-wide association mapping of seed coat color in Brassica napus. J Agric Food Chem 65:5229–5237
Wang X, Wang H, Long Y, Li D, Yin Y, Tian J, Chen L, Liu L, Zhao W, Zhao Y, Yu L, Li M (2013) Identification of QTLs associated with oil content in a high-oil Brassica napus cultivar and construction of a high-density consensus map for QTLs comparison in B napus. PLoS ONE 8:e80569
Wang Z, Chen MX, Chen TL, Xuan LJ, Li ZL, Du X, Zhou LH, Zhang GP, Jiang LX (2014) TRANSPARENT TESTA2 regulates embryonic fatty acid biosynthesis by targeting FUSCA3 during the early developmental stage of Arabidopsis seeds. Plant J 77:757–769
Warwick SI, Simard MJ, Legere A, Beckie HJ, Braun L, Zhu B, Mason P, Seguin-Swartz G, Stewart CN (2003) Hybridization between transgenic Brassica napus L. and its wild relatives: Brassica rapa L., Raphanus raphanistrum L., Sinapis arvensis L., and Erucastrum gallicum (Willd.) OE Schulz. Theor Appl Genet 107:528–539
Wen J, Zhu LX, Qi LP, Ke HM, Yi B, Shen JX, Tu JX, Ma CZ, Fu TD (2012) Characterization of interploid hybrids from crosses between Brassica juncea and B. oleracea and the production of yellow-seeded B. napus. Theor Appl Genet 125:19–32
Western TL, Skinner DJ, Haughn GW (2000) Differentiation of mucilage secretory cells of the Arabidopsis seed coat. Plant Physiol 122:345–355
Xie T, Chen X, Guo TL, Rong H, Chen ZY, Sun QF, Batley J, Jiang JJ, Wang YP (2020) Targeted knockout of BnTT2 homologues for yellow-seeded Brassica napus with reduced flavonoids and improved fatty acid composition. J Agr Food Chem 68:5676–5690
Xuan LJ, Zhang CC, Yan T, Wu DZ, Hussain N, Li ZL, Chen MX, Pan JW, Jiang LX (2018) TRANSPARENT TESTA 4-mediated flavonoids negatively affect embryonic fatty acid biosynthesis in Arabidopsis. Plant Cell Environ 41:2773–2790
Yan Z, Xia L, Wei C, Yi B, Jing W, Shen J, Ma C, Chen B, Tu J, Fu T (2011) Identification of two major QTL for yellow seed color in two crosses of resynthesized Brassica napus line No. 2127–17. Mol Breeding 28:335–342
Zhai Y, Yu K, Cai S, Hu L, Amoo O, Xu L, Yang Y, Ma B, Jiao Y, Zhang C, Khan MHU, Khan SU, Fan C, Zhou Y (2020) Targeted mutagenesis of BnTT8 homologs controls yellow seed coat development for effective oil production in Brassica napus L. Plant Biotechnol J 18:1153–1168
Zhou LH, Li YL, Hussain N, Li ZL, Wu DZ, Jiang LX (2016) Allelic variation of BnaC.TT2.a and its association with seed coat color and fatty acids in rapeseed (Brassica napus L.). PLoS ONE 11:e0146661
Funding
This work was financially supported by the National Natural Science Foundation of China (31871656, 32001583 and 32072098), and the Key Research Plan Project of Shaanxi Province (2020ZDLNY04-01).
Author information
Authors and Affiliations
Contributions
HC carried out QTL mapping and BSR analysis and wrote the manuscript. LG, WZ and HL participated in the field experiment and surveyed and analyzed the phenotypic data. ML designed the overall study and provided guidelines for writing the paper.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest.
Ethical standards
The authors declare that the experiments comply with the current laws of the country in which they were performed.
Additional information
Communicated by Jacqueline Batley.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.


Rights and permissions
About this article
Cite this article
Chao, H., Guo, L., Zhao, W. et al. A major yellow-seed QTL on chromosome A09 significantly increases the oil content and reduces the fiber content of seed in Brassica napus. Theor Appl Genet 135, 1293–1305 (2022). https://doi.org/10.1007/s00122-022-04031-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-022-04031-0