Abstract
Key message
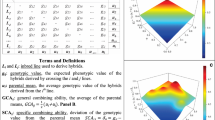
A new genomic model that incorporates genotype × environment interaction gave increased prediction accuracy of untested hybrid response for traits such as percent starch content, percent dry matter content and silage yield of maize hybrids.
Abstract
The prediction of hybrid performance (HP) is very important in agricultural breeding programs. In plant breeding, multi-environment trials play an important role in the selection of important traits, such as stability across environments, grain yield and pest resistance. Environmental conditions modulate gene expression causing genotype × environment interaction (G × E), such that the estimated genetic correlations of the performance of individual lines across environments summarize the joint action of genes and environmental conditions. This article proposes a genomic statistical model that incorporates G × E for general and specific combining ability for predicting the performance of hybrids in environments. The proposed model can also be applied to any other hybrid species with distinct parental pools. In this study, we evaluated the predictive ability of two HP prediction models using a cross-validation approach applied in extensive maize hybrid data, comprising 2724 hybrids derived from 507 dent lines and 24 flint lines, which were evaluated for three traits in 58 environments over 12 years; analyses were performed for each year. On average, genomic models that include the interaction of general and specific combining ability with environments have greater predictive ability than genomic models without interaction with environments (ranging from 12 to 22%, depending on the trait). We concluded that including G × E in the prediction of untested maize hybrids increases the accuracy of genomic models.



Similar content being viewed by others
References
Bernardo R (1994) Prediction of maize single-cross performance using RFLPs and information from related hybrids. Crop Sci 34:20–25
Bernardo R (1996a) Best linear unbiased prediction of maize single-cross performance. Crop Sci 36:50–56. doi:10.2135/cropsci1996.00111183X003600010009x
Bernardo R (1996b) Best linear unbiased prediction of the performance of crosses in maize. Crop Sci 41:68–71. doi:10.2135/cropsci2001.41168x
Bernardo R (1999) Marker-assisted best linear unbiased prediction of single-cross performance. Crop Sci 39:1277–1282
Bernardo R (2002) Breeding for quantitative traits in plants. Stemma Press, Woodbury
Browning BL, Browning SR (2009) A unified approach to genotype imputation and haplotype-phase inference for large data sets of trios and unrelated individuals. Am J Hum Genet 84(2):210–223. doi:10.1016/j.ajhg.2009.01.005
Burgueño J, de los Campos G, Weigel K, Crossa J (2012) Genomic prediction of breeding values when modeling genotype × environment interaction using pedigree and dense molecular markers. Crop Sci 52:707–719
Covarrubias-Pazaran G (2016) Genome-assisted prediction of quantitative traits using the R Package Sommer. Plos One 11(6):e0156744. doi:10.1371/journal.pone.0156744
Crossa J, Pérez-Rodríguez P, de los Campos G, Mahuku G, Dreisigacker D, Magorokosho C (2011) Genomic selection and prediction in plant breeding. J Crop Improv 25:239–261
Dawson JC, Endelman JB, Heslot N, Crossa J, Poland J et al (2013) The use of unbalanced historical data for genomic selection in an international wheat breeding program. Field Crops Res 154:12–22
de los Campos G, Pérez-Rodríguez P (2016) BGLR: Bayesian generalized linear regression. R package v. 1.0.5
de los Campos G, Hickey JM, Pong-Wong R, Daetwyler HD, Calus MPL (2013) Whole genome regression and prediction methods applied to plant and animal breeding. Genetics 193:327–345
Duvick DN, Smith JSC, Cooper M (2004) Long-term selection in a commercial hybrid maize breeding program. Plant Breed Rev Part 2(24):109–152
Federer WT, Raghavarao D (1975) On augmented designs. Biometrics 31:29–35
Geman S, Geman D (1984) Stochastic relaxation, Gibbs distributions and the Bayesian restoration of images. IEEE Trans Pattern Anal Mach Intell 6(6):721–741
Golob GH, Van Loan CF (1996) Matrix computations, 3rd edn. Johns Hopkins University Press, Baltimore
Hastie T, Tibshirani R, Friedman J (2009) The elements of statistical learning: data mining, inference and prediction, 2nd edn. Springer, New York
Heslot N, Akdemir D, Sorrells ME, Jannink JL (2014) Integrating environmental covariates and crop modeling into the genomic selection framework to predict genotype by environment interactions. Theor App Genet 127(2):463–480
Jarquín D, Crossa J, Lacaze X, Cheyron PD, Daucourt J et al (2014) A reaction norm model for genomic selection using high-dimensional genomic and environmental data. Theor Appl Genet 127:595–607
Kadam DC, Potts SM, Bohn MO, Lipka A, Lorenz AJ (2016) Genomic prediction of single crosses in the early stages of a maize hybrid breeding pipeline. G3 Genes Genom Genet 6:3443–3453. doi:10.1534/g3.116.031286
Lehermeier C, Krämer N, Bauer E, Bauland C, Camisan C et al (2014) Usefulness of multiparental populations of maize (Zea mays L.) for genome-based prediction. Genetics 198:3–16
López-Cruz M, Crossa J, Bonnet D, Dreisigacker S, Poland J, Jannink L-L, Singh RP, Autrique E, de los Campos G (2015) Increased prediction accuracy in wheat breeding trials using a marker × environment interaction genomic selection model. G3 Genes Genom Genet. doi:10.1534/g3.114.016097
Massman J, Gordillo A, Lorenzana RE, Bernardo R (2013) Genome-wide predictions from maize single-cross data. Theor Appl Genet 126:13–22. doi:10.1007/s00122-012-1955
Meuwissen, THE, Hayes BJ, Goddard ME (2001) Prediction of total genetic value using genome-wide dense marker maps. Genetics 157:1819–1829
Pérez-Rodríguez P, de los Campos G (2014) Genome-wide regression & prediction with the BGLR statistical package. Genetics 198:483–495
Pérez-Rodríguez P, Crossa J, Bondalapati K, De Meyer G, Pita F, de los Campos G (2015) A pedigree-based reaction norm model for prediction of cotton yield in multi-environment trials. Crop Sci 55(3):1143–1151
Piepho HP (2009) Ridge regression and extensions for genome-wide selection in maize. Crop Sci 49(4):1165–1176
R Core Team (2016) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna. http://www.R-project.org. Accessed 4 Apr 2017
Rodríguez-Ramilo ST, García-Cortés LA, Rodríguez de Cara MA (2015) Artificial selection with traditional of genomic relationships: consequences in coancestry and genetic diversity. Front Genet 6:127. doi:10.3389/fgene.2015.00127
Schrag TA, Möhring J, Melchinger AE, Kusterer B, Dhillon BS, Piepho HP, Frisch M (2010) Prediction of hybrid performance in maize using molecular markers and joint analyses of hybrids and parental inbreds. Theor Appl Genet 120(2):451–461
Technow F, Melchinger AE (2013) Genomic prediction of dichotomous traits with Bayesian logistic models. Theor Appl Genet 125(6):1133–1143
Technow F, Riedelsheimer E, Schrag TA, Melchinger AE (2012) Genomic prediction of hybrid performance in maize with models incorporating dominance and population specific marker effects. Theor Appl Genet 125(6):1181–1194
Technow F, Schrag A, Schipprack W, Bauer E, Simianer H, Melchinger AE (2014) Genome properties and prospects of genomic prediction of hybrid performance in a breeding program of maize. Genetics 197:1343–1355
van Eeuwijk FA, Boer MP et al (2010) Mixed model approaches for the identification of QTLs within a maize hybrid breeding program. Theor Appl Genet 120:429–440
VanRaden PM (2008) Efficient methods to compute genomic predictions. J Dairy Sci 91:4414–4423
Xu S, Zhu D, Zhang Q (2014) Predicting hybrid performance in rice using genomic best linear unbiased prediction. Proc Natl Acad Sci 11(34):12456–12461. doi:10.1073/pnas.1413750111
Zhao Y, Zeng J, Fernando R, Reif J (2013) Genomic prediction of hybrid wheat performance. Crop Sci 53:1–9. doi:10.2135/cropsci2012.08.0463
Acknowledgements
The authors appreciate the positive and detailed comments from two anonymous reviewers and the time invested by the Associated Editor handling the manuscript. These contributions significantly improved the quality and clarity of the article.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Additional information
Communicated by Matthias Frisch.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Acosta-Pech, R., Crossa, J., de los Campos, G. et al. Genomic models with genotype × environment interaction for predicting hybrid performance: an application in maize hybrids. Theor Appl Genet 130, 1431–1440 (2017). https://doi.org/10.1007/s00122-017-2898-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-017-2898-0