Abstract
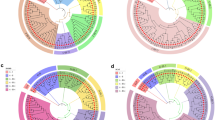
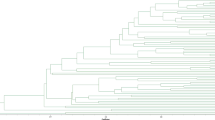
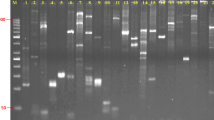
A collaborative international program was initiated to identify and describe the genetic diversity of living germplasm collections of Theobroma cacao genotypes that are maintained in several international collections scattered throughout tropical cacao growing countries of the world. Simple sequence repeat (SSR) DNA analysis was identified as the most appropriate molecular tool for DNA fingerprinting these collections during an international forum representing academic, government and industry scientists in the cacao community. Twenty-five SSR primers, which had been previously described, were evaluated as potential candidates to define an efficient, standardized, molecular fingerprinting protocol for T. cacao accessions. These primers have been evaluated for reliability, widespread distribution across the cacao genome, number of alleles produced by the SSR primers in cacao and their ability to discriminate between cacao accessions. Approximately 690 cacao accessions were used to evaluate the utility of these SSR primers as international molecular standards, and a small number of test samples of T. cacao were sent to two other independent laboratories for verification. DNA fragments were selectively amplified by PCR, using the SSR primers labeled with fluorescent dyes, and separated by capillary electrophoresis. Based on this study, the 15 SSR primers that had the highest reproducibility and consistency within a common genotype, while allowing the differentiation of separate divergent genotypes, were selected as international molecular standards for DNA fingerprinting of T. cacao.

Similar content being viewed by others
References
Bredemeijer GMM, Arens P, Wouters D, Visser D, Vosman B (1998) The use of semi-automated fluorescent microsatellite analysis for tomato cultivar identification. Theor Appl Genet 97:584–590
Charters MY, Wilkinson MJ (2000) The use of self-pollinated progenies as “in-groups” for the genetic characterization of cocoa germplasm. Theor Appl Genet 100:160–166
Chavarriaga-Aguirre P, Maya MM, Bonierbale MW, Kresovich S, Fregene MA, Tohme J, Kochert G (1998) Microsatellites in cassava (Manihot esculenta Crantz): discovery, inheritance and variability. Theor Appl Genet 97:493–501
Christopher Y, Mooleedhar V, Bekele F, Hosein F (1999) Verification of accession in the ICG, T using botanical descriptors and RAPD analysis. In: Annual report 1998. St. Augustine, Trinidad and Tobago: Cocoa Research Unit, The University of the West Indies, pp 15–18
Coe SD, Coe MD (1996) The true history of chocolate. Thames and Hudson, London, pp 16–34
Degani C, Rowland LJ, Saunders JA, Hokanson SC, Ogden EL, Golan-Goldhirsh A, Galletta GJ (2001) A comparison of genetic relationship measures in strawberry (Fragaria ananassaDuch.) based on AFLPs, RAPDs, and pedigree data. Euphytica 117:1–12
Figueira A, Janick J, Levy M, Goldsbrough PB (1994) Re-examining the classification of Theobroma cacao L. using molecular markers. J Am Soc Hort Sci 119:1073–1082
Lanaud C, Risterucci AM, Pieretti I, Falque M, Bouet A, Lagoda PJL (1999) Isolation and characterization of microsatellites inTheobroma cacaoL. Mol Ecol 8:2141–2143
McBride J (2002) Tropical agriculture gets attention. Agric Res 50:10–11
Motamayor JC, Lanaud C (2002) Molecular analysis of the origin and domestication of Theobroma cacao L., In: Engles JMM, Ramanatha RV, Brown AHD, Jackson MT (eds) Managing plant genetic diversity. IPGRI, pp 77–87
Motilal LA, Butler DR, Mooleedhar V (2002) Verification in global cacao germplasm collections. Ingenic Newslett 7:4–8
Risterucci AM, Grivet L, N’Goran JAK, Pierevtti I, Flament MH, Lanaud C (2000) A high density linkage map of Theobroma cacao L. Theor Appl Genet 101:948–955
Saunders JA, Pedroni MJ, Daughtry CS (1999) DNA fingerprinting of marijuana by the AFLP technique. Focus 20:10–11
Saunders JA, Hemeida AA, Mischke S (2001a) USDA DNA fingerprinting programme for identification of Theobroma cacao accessions. In: Proceedings of the international workshop on new technologies and cocoa breeding, 16–17 October 2000, Kota Kinabalu, Malaysia, pp 108–114
Saunders JA, Pedroni MJ, Penrose LDJ, Fist AJ (2001b) AFLP DNA analysis of opium poppy. Crop Sci 41:1596–1601
Swanson J-D, Lee AC, Guiltinan MJ (2003) Ring test: results from Penn State University. Ingenic Newslett 8:22–27
Wright H (1999) Cocoa, its botany, cultivation, chemistry and diseases. Biotech Books, Delhi, pp 1–22
Young AM (1994) The chocolate tree, a natural history of cacao. Smithsonian Institution, Washington, pp 65–79
Acknowledgements
The authors gratefully thank Dr. Ulrike Krauss from CATIE, Costa Rica, Dr. Ricardo Goenaga from the USDA in Puerto Rico, and Dr. David Butler, Dr. Elizabeth Johnson, and Mr. Oliver Sounigo from CRU, Trinidad and Tobago for providing the cacao plant accessions used in this study. They also thank Dr. Claire Lanaud for recommendations on SSR primers and review of the manuscript and Dr. Mark Guiltinan and Dr. Ray Schnell for the independent verification analysis of the DNA primers.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by D.B. Neale
Rights and permissions
About this article
Cite this article
Saunders, J.A., Mischke, S., Leamy, E.A. et al. Selection of international molecular standards for DNA fingerprinting of Theobroma cacao. Theor Appl Genet 110, 41–47 (2004). https://doi.org/10.1007/s00122-004-1762-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-004-1762-1