Abstract
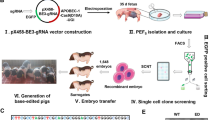
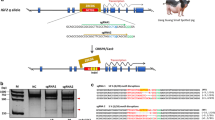
Insulin-like growth factor 2 (IGF2) is an important growth factor, which promotes growth and development in mammals during fetal and postnatal stages. Using CRISPR–Cas9 system, we generated multiple founder pigs containing 12 different mutant alleles around a regulatory element within the intron 3 of IGF2 gene. Crossing two male founders passed four mutant alleles onto F1 generation, and these mutations abolished repressor ZBED6 binding and rendered this regulatory element nonfunctional. Both founders and F1 animals showed significantly faster growth, without affecting meat quality. These results indicated that editing IGF2 intron 3–3072 site using CRISPR–Cas9 technology improved meat production in Bama pigs. This is the first demonstration that editing non-coding region can improve economic traits in livestock.




Similar content being viewed by others
Abbreviations
- Cas9:
-
CRISPR-associated protein 9
- CRISPR:
-
Clustered, regularly interspaced, short-palindromic repeats
- EMSA:
-
Electrophoretic mobility shift assay
- GM:
-
Genetically modified
- IGF2:
-
Insulin-like growth factor 2
- QTL:
-
Trait locus
- QTN:
-
Quantitative trait nucleotide
- sgRNA:
-
Single-guide RNA
- WT:
-
Wild type
- ZBED6:
-
BED-type containing 6
References
Spychalla JP, Kinney AJ, Browse J (1997) Identification of an animal omega-3 fatty acid desaturase by heterologous expression in Arabidopsis. Proc Natl Acad Sci USA 94:1142–1147
Grobet L, Martin LJ, Poncelet D, Pirottin D, Brouwers B, Riquet J, Schoeberlein A, Dunner S, Menissier F, Massabanda J et al (1997) A deletion in the bovine myostatin gene causes the double-muscled phenotype in cattle. Nat Genet 17:71–74
Jeon JT, Carlborg O, Tornsten A, Giuffra E, Amarger V, Chardon P, Andersson-Eklund L, Andersson K, Hansson I, Lundstrom K et al (1999) A paternally expressed QTL affecting skeletal and cardiac muscle mass in pigs maps to the IGF2 locus. Nat Genet 21:157–158
Nezer C, Moreau L, Brouwers B, Coppieters W, Detilleux J, Hanset R, Karim L, Kvasz A, Leroy P, Georges M (1999) An imprinted QTL with major effect on muscle mass and fat deposition maps to the IGF2 locus in pigs. Nat Genet 21:155–156
Laible G, Wei J, Wagner S (2015) Improving livestock for agriculture—technological progress from random transgenesis to precision genome editing heralds a new era. Biotechnol J 10:109–120
Hammer RE, Pursel VG, Rexroad CE Jr, Wall RJ, Bolt DJ, Ebert KM, Palmiter RD, Brinster RL (1985) Production of transgenic rabbits, sheep and pigs by microinjection. Nature 315:680–683
McCreath KJ, Howcroft J, Campbell KH, Colman A, Schnieke AE, Kind AJ (2000) Production of gene-targeted sheep by nuclear transfer from cultured somatic cells. Nature 405:1066–1069
Ohlsen SM, Lugenbeel KA, Wong EA (1994) Characterization of the linked ovine insulin and insulin-like growth factor-II genes. DNA Cell Biol 13:377–388
DeChiara TM, Efstratiadis A, Robertson EJ (1990) A growth-deficiency phenotype in heterozygous mice carrying an insulin-like growth factor II gene disrupted by targeting. Nature 345:78–80
Sun FL, Dean WL, Kelsey G, Allen ND, Reik W (1997) Transactivation of Igf2 in a mouse model of Beckwith–Wiedemann syndrome. Nature 389:809–815
Van Laere AS, Nguyen M, Braunschweig M, Nezer C, Collette C, Moreau L, Archibald AL, Haley CS, Buys N, Tally M et al (2003) A regulatory mutation in IGF2 causes a major QTL effect on muscle growth in the pig. Nature 425:832–836
Markljung E, Jiang L, Jaffe JD, Mikkelsen TS, Wallerman O, Larhammar M, Zhang X, Wang L, Saenz-Vash V, Gnirke A et al (2009) ZBED6, a novel transcription factor derived from a domesticated DNA transposon regulates IGF2 expression and muscle growth. PLoS Biol 7:e1000256
Hu LL, Lu YQ, Xu HY, Yang XG, Lu SS, Lu KH (2015) Production of hGFAP-DsRed transgenic Guangxi Bama mini-pigs via somatic cell nuclear transfer. Genet Mol Res 14:16285–16296
Hai T, Teng F, Guo R, Li W, Zhou Q (2014) One-step generation of knockout pigs by zygote injection of CRISPR/Cas system. Cell Res 24:372–375
Niu Y, Shen B, Cui Y, Chen Y, Wang J, Wang L, Kang Y, Zhao X, Si W, Li W et al (2014) Generation of gene-modified cynomolgus monkey via Cas9/RNA-mediated gene targeting in one-cell embryos. Cell 156:836–843
Wang H, Yang H, Shivalila CS, Dawlaty MM, Cheng AW, Zhang F, Jaenisch R (2013) One-step generation of mice carrying mutations in multiple genes by CRISPR/Cas-mediated genome engineering. Cell 153:910–918
Peng J, Wang Y, Jiang J, Zhou X, Song L, Wang L, Ding C, Qin J, Liu L, Wang W et al (2015) Production of human albumin in pigs through CRISPR/Cas9-mediated knockin of human cDNA into swine albumin locus in the zygotes. Sci Rep 5:16705
Yang Y, Wang K, Wu H, Jin Q, Ruan D, Ouyang Z, Zhao B, Liu Z, Zhao Y, Zhang Q et al (2016) Genetically humanized pigs exclusively expressing human insulin are generated through custom endonuclease-mediated seamless engineering. J Mol Cell Biol 8:174–177
Yan S, Tu Z, Liu Z, Fan N, Yang H, Yang S, Yang W, Zhao Y, Ouyang Z, Lai C et al (2018) A Huntingtin knockin pig model recapitulates features of selective neurodegeneration in Huntington’s disease. Cell 173(989–1002):e1013
Yang L, Guell M, Niu D, George H, Lesha E, Grishin D, Aach J, Shrock E, Xu W, Poci J et al (2015) Genome-wide inactivation of porcine endogenous retroviruses (PERVs). Science 350:1101–1104
Qin W, Kutny PM, Maser RS, Dion SL, Lamont JD, Zhang Y, Perry GA, Wang H (2016) Generating mouse models using CRISPR–Cas9-mediated genome editing. Curr Protoc Mouse Biol 6:39–66
Schmittgen TD, Livak KJ (2008) Analyzing real-time PCR data by the comparative C(T) method. Nat Protoc 3:1101–1108
Dyer BW, Ferrer FA, Klinedinst DK, Rodriguez R (2000) A noncommercial dual luciferase enzyme assay system for reporter gene analysis. Anal Biochem 282:158–161
Yang SL, Wang ZG, Liu B, Zhang GX, Zhao SH, Yu M, Fan B, Li MH, Xiong TA, Li K (2003) Genetic variation and relationships of eighteen Chinese indigenous pig breeds. Genet Sel Evol 35:657–671
Ran FA, Hsu PD, Lin CY, Gootenberg JS, Konermann S, Trevino AE, Scott DA, Inoue A, Matoba S, Zhang Y et al (2013) Double nicking by RNA-guided CRISPR Cas9 for enhanced genome editing specificity. Cell 154:1380–1389
McPherron AC, Lee SJ (1997) Double muscling in cattle due to mutations in the myostatin gene. Proc Natl Acad Sci USA 94:12457–12461
Zheng Q, Lin J, Huang J, Zhang H, Zhang R, Zhang X, Cao C, Hambly C, Qin G, Yao J et al (2017) Reconstitution of UCP1 using CRISPR/Cas9 in the white adipose tissue of pigs decreases fat deposition and improves thermogenic capacity. Proc Natl Acad Sci USA 114:E9474–E9482
Gao CX (2015) Genome editing in crops: from bench to field. Natl Sci Rev 2:13–15
Waltz E (2016) Gene-edited CRISPR mushroom escapes US regulation. Nature 532:293
Sharma A, Lee JS, Dang CG, Sudrajad P, Kim HC, Yeon SH, Kang HS, Lee SH (2015) Stories and challenges of genome wide association studies in livestock—a review. Asian Australas J Anim Sci 28:1371–1379
Freking BA, Murphy SK, Wylie AA, Rhodes SJ, Keele JW, Leymaster KA, Jirtle RL, Smith TP (2002) Identification of the single base change causing the callipyge muscle hypertrophy phenotype, the only known example of polar overdominance in mammals. Genome Res 12:1496–1506
Acknowledgements
We thank H.W and Q.Z laboratory members for helpful discussions and comments on the manuscript. H.W was supported by Strategic Priority Research Program of Chinese Academy of Sciences (no. XDA16010205), the National Natural Science Foundation of China (nos. 31471215 and 31722036), and National Key Research and Development Program of China (no. 2016YFA0101402). Q.Z was supported by National Natural Science Foundation of China Grants 31422038, 31471395, and 31501188; National High Technology Research and Development Program Grants 2015AA020307 and 2014BAI02B01; National Basic Research Program of China Grants 2014CB964900 and 2014CB964800; National Special Key Project for Transgenic Breeding 2016ZX08010001.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
Guanghai Xiang, Jilong Ren, Tang Hai, Rui Fu, Dawei Yu, Jing Wang, Wei Li, Haoyi Wang, and Qi Zhou declare that they have no conflict of interest.
Ethical approval
All experiments related to animal work described in this study were performed strictly in accordance with the guidelines for the Care and Use of Laboratory Animals, and approved by Animal Welfare and Research Ethics Committee of Institute of Zoology, Chinese Academy of Sciences (Animal Ethics Investigation AEI-12-04-2014). All surgical procedures were performed under anesthesia, and great efforts were made to minimize animal suffering.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Xiang, G., Ren, J., Hai, T. et al. Editing porcine IGF2 regulatory element improved meat production in Chinese Bama pigs. Cell. Mol. Life Sci. 75, 4619–4628 (2018). https://doi.org/10.1007/s00018-018-2917-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00018-018-2917-6