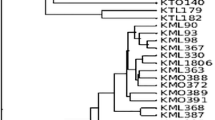
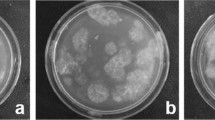
Sequences of the ribosomal DNA (rDNA) internal transcribed spacer (ITS) regions were examined to infer a molecular phylogeny of small-spored Phomopsis isolates, designated W-type (mainly white colony, weakly virulent, bearing both alpha and beta conidia at 25°C on PDA) and G-type (mainly gray colony, highly virulent, bearing only alpha conidia at 25°C on PDA), and P. amygdali from fruit trees. Phomopsis G-type and P. amygdali were a monophyletic group distinct from the W-type. The W-type isolates were divided into two monophyletic groups. Diaporthe citri, D. tanakae, P. asparagi, P. viticola, P. vitimegaspora and D. nomurai, which are morphologically distinguishable from W- and G-types, differed from the W- and G-types in molecular phylogenetic analyses. PCR-RFLP analysis of rDNA ITS regions was useful to distinguish each of the Phomopsis species and groups using three restriction enzymes. In mating tests, W-type isolates from fruit trees were heterothallic and inter-fertile even between isolates belonging to the different monophyletic groups. Isolates of the G-type and P. amygdali collected in Japan were cross-fertile. Some isolates from Lunaria annua, Ulmus glabra and Juglans regia belonged to one of the two monophyletic groups of the W-type and were cross-fertile with W-type isolates from Rosaceous fruit trees.
Similar content being viewed by others
Author information
Authors and Affiliations
Additional information
Received 27 September 1999/ Accepted in revised form 27 January 2000
Rights and permissions
About this article
Cite this article
KANEMATSU, S., MINAKA, N., KOBAYASHI, T. et al. Molecular Phylogenetic Analysis of Ribosomal DNA Internal Transcribed Spacer Regions and Comparison of Fertility in Phomopsis Isolates from Fruit Trees. J Gen Plant Pathol 66, 191–201 (2000). https://doi.org/10.1007/PL00012944
Issue Date:
DOI: https://doi.org/10.1007/PL00012944