Abstract.
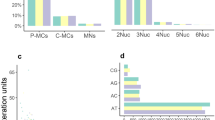
Size homoplasy was analyzed at microsatellite loci by sequencing electromorphs, that is, variants of the same size (base pairs). This study was conducted using five interrupted and/or compound loci in three invertebrate species, the honey bee Apis mellifera, the bumble bee Bombus terrestris, and the freshwater snail Bulinus truncatus. The 15 electromorphs sequenced turned out to hide 31 alleles (i.e., variants identical in sequence). Variation in the amount of size homoplasy was detected among electromorphs and loci. From one to seven alleles were detected per electromorph, and one locus did not show any size homoplasy in both bee species. The amount of size homoplasy was related to the sequencing effort, since the number of alleles was correlated with the number of copies of electromorphs sequenced, but also with the molecular structure of the core sequence at each locus. Size homoplasy within populations was detected only three times, meaning that size homoplasy was detected mostly among populations. We analyzed population structure, estimating F st and a genetic distance, based on either electromorphs or alleles. Whereas little difference was found in A. mellifera, uncovering size homoplasy led to a more marked population structure in B. terrestris and B. truncatus. We also showed in A. mellifera that the detection of size homoplasy may alter phylogenetic reconstructions.
Similar content being viewed by others
Author information
Authors and Affiliations
Additional information
Received: 21 July 1997 / Accepted: 29 January 1998
Rights and permissions
About this article
Cite this article
Viard, F., Franck, P., Dubois, MP. et al. Variation of Microsatellite Size Homoplasy Across Electromorphs, Loci, and Populations in Three Invertebrate Species. J Mol Evol 47, 42–51 (1998). https://doi.org/10.1007/PL00006361
Issue Date:
DOI: https://doi.org/10.1007/PL00006361