Abstract
Proteases or proteinases are essential constituents for all the existing live forms. They act as important industrial enzymes occupying about 60% of total enzyme market. In the present study proteases extracted from nine medicinally important spices; Carum copticum, Syzygium aromaticum, Cuminum cyminum, Nigella sativa, Cinnamomum verum, Foeniculum vulgare, Zingiber officinale, Cinnamomum tamala and Curcuma longa; used in our food and as household medicines on regular basis, have been investigated. Amongst these spices, the specific activity of the isolated protease enzyme was found to be significantly high in Nigella sativa (204 units/mg) and Curcuma longa (124 units/mg) and therefore, their extract was further partially purified and biochemically characterized. The crude extract of the two spices on being subjected to salt precipitation using (NH4)2SO4 as neutral salt, yielded three fractions at 0-30%, 30-60% and 60-90% saturation level. The specific activity of protease enzyme was found to be highest in 0-30% fraction, obtained from both the sources. The values of specific activity and purification-fold of enzyme from Nigella sativa and Curcuma longa were found to be 409 U/mg, 590 U/mg and 2.0-, 4.8-, respectively. The enzyme showed maximum activity at pH and temperature conditions of 5.0 and 40°C, respectively, at 20 min of incubation time, in both the cases. The pH stability values of protease from Nigella sativa ranged from 5.0 to 9.0 whereas in Curcuma longa values were from 4.0 to 8.0. The enzyme from Nigella sativa and Curcuma longa exhibited resistance against heat treatment upto 60°C and 50°C, respectively, increasing the industrial feasibility. The present work hereby, indicates that Nigella sativa and Curcuma longa may serve as good source of thermostable protease; an enzyme of great relevance in various chemical and bio-industries along with pharma sector.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
I. Introduction
Proteases are biocatalyst that perform a multitude of chemical reaction and are commercially used in detergents, food, pharma, diagnostic and fine chemical industries [1, 2]. They are group of enzymes which catalyse the cleavage of peptide bonds, generally referred to as proteolytic enzymes or proteinases. Proteases are ubiquitous in nature widely distributed in plants, animals and microorganisms.
The significant role of proteases has made them industrially important. They account for 65% of total worldwide enzyme sale. Several proteases have been isolated from latex, fruits and seeds. They have major applications in industrial process such as laundry, silk, pharmaceutical, food, degradation of gelatin on X-ray films [3] and bioremediation process. They are widely used in detergents, leather, waste management and silver recovery [4].
A lot of work has been reported on proteases of microbial origin holding industrial potential due to their biochemical diversity. A variety of microorganisms such as bacteria, fungi, yeast and actinomycetes are known to produce these enzymes [5]. It would be advantageous to utilize a fungal protease as fungal expression systems are capable of producing larger quantities of enzymes than bacterial expression system [6]. Proteases are produced by many species of fungi such as Aspergillus [7], Mucor [8], Fusarium [9], Cephalosporium [10] and Rhizopus [11]. Filamentous fungi, such as Aspergillus have been the organism of choice for large scale production of bulk industrial enzymes [12]. Actinomycetes being a protease producing bacteria have the potential for use industrial purpose, pharmaceutical and cytotoxic agent [13]. The sources of protease are enormous. Bacillus sp. was found to be predominant and rich source of alkaline proteases. Many of the fungi have also been reported to produce extracellular alkaline protease [14]. B. substilis yield maximum protease production and be used efficiently for industrial purposes [15]. Among various proteases, bacterial proteases are more significant compared to with animal and fungal protease [16]. They are present in Bacillus subtilis, B. amyloliquefaciens, Pseudomonas sp., Lysobacter enzymogene, E. coli etc. [17]. B. subtilis produce alkaline protease that can be used in textiles, leather and food industries [18]. Plants are also the rich source of protease enzyme [19] but not been worked upon much. In this study, spices such as Carum copticum, Syzygium aromaticum, Cuminum cyminum, Nigella sativa, Cinnamon verum, Foeniculum vulgare, Zingiber officinale, Cinnamomum tamala and Curcuma longa have been explored as source of protease enzyme.
II. Materials and methods
Spices were procured from Green Earth products Pvt. Ltd., New Delhi. All chemicals were of reagent grade and obtained from standard commercial firms.
A. Screening of various spices for Protease enzyme
Screening of various spices viz; Ajwain (Carum copticum), Cloves (Syzygium aromaticum), Cumin (Cuminum cyminum), Black cumin (Nigella sativa), Dal chini (Cinnamon verum), Fennel (Foeniculum vulgare), Tej patta (Cinnamomum tamala), Ginger (Zingiber officinale) and Turmeric (Curcuma longa) was done for protease activity by using standard protocols.
B. Extraction of Protease enzyme
The pre-weighed samples were crushed in sodium acetate buffer (pH 5.0, 0.05 M), filtered through Whatman filter paper and centrifuged for 20 minutes at 10,000 rpm at 4°C. The pellet was discarded, supernatant collected and subjected to further purification.
C. Determination of specific activity
Specific activity was determined by using the following relationship:
Specific activity= Total enzyme units /Total protein (mg)
-
Protein determination: Protein content of the enzyme extract was determined by Lowry method [20] using BSA as standard.
-
Protease Assay: Protease activity was assayed using Folin-Ciocalteau method [21] The reaction mixture containing casein (1%), enzyme, 0.05 M sodium acetate buffer (pH 5.0) was incubated for 30 min at 30°C followed by addition of 0.5 M NaOH and folins reagent. 1 International Unit of protease enzyme is defined as 1μg of tyrosine released per minute per ml under standard assay conditions. The amount of protease produced was measured with the help of a tyrosine standard graph [22]. The activity was reported as mean of three determinations.
D. Partial purification of protease enzyme
-
Ammonium sulphate fractionation: The crude extract of the screened spices were subjected to precipitation using salting out process. Ammonium sulphate fractionation (0-30%) was done by adding salt in the extract according to the required saturation level, slowly while keeping on ice [23]. The ice-cold saturated solution of the protein was stirred continuously and kept at 0-4°C for at least one hour followed by centrifugation at 10,000 rpm for 10-15 min. Pellets were collected and dissolved in minimal amount of sodium acetate buffer (0.05 M, pH 5.0) and used as 0-30% fraction, the supernatant being subjected to next fractionation steps and further two fractions (30-60% and 60-90%) were obtained in the similar manner.
E. Biochemical characterization of Protease enzyme
-
Time course: To determine the time course, the reaction mixture was incubated at different time intervals ranging from 10 min to 90 min and the protease activity was determined using standard conditions.
-
Temperature Optima: The reaction mixture containing enzyme and substrate was incubated at 30°C for time period ranging between 10-90 minutes and the product released estimated by Folin’s method.
-
pH optima: To determine the pH optima, suitable buffers of different pH values ranging from 3.0 to 9.0 were used. The reaction mixture was incubated for optimum time period and activity determined using standard assay.
-
pH stability: The enzyme alone was incubated for 2 hours with suitable buffers of pH ranging from 3.0 to 9.0 and further assayed using optimum assay conditions and residual protease activity, determined.
-
Heat Resistance: The enzyme was incubated alone at different temperature ranging from 10°C to 90°C for 2h followed by standard assay.
III. Results and discussion
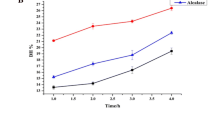
Protease plays an essential role in various pathological processes. Arthritis, tumor invasion and metastasis, infections and number of degenerative disease have been linked with proteolytic enzymes which have widespread applications in bioremediation processes [24]. This research work was undertaken to study the various biochemical aspects of high specific activity protease enzyme isolated from spices. Out of the nine spices studied, the extract of Nigella sativa (204 units/mg of protein) and Curcuma longa (124 units/mg) was found to contain protease with maximum specific as shown in Table I, and was further characterized. A better understanding of the functions of enzymes could be determined by purification of enzyme [25]. Figure 1 shows the specific activity of fractions of protease isolated from Nigella sativa and Curcuma longa at three different saturation levels of (NH4)2SO4. It can be seen from Table II that 0-30% fraction has highest specific activity with 2- fold and 4.8- fold purification level in N. sativa (409 U/mg) and C. longa (590 U/mg), respectively. Therefore, 0-30% enzyme fraction from both the sources was used for further characterization studies. The time course of enzyme catalyzed reaction for protease isolated from N. sativa and C. longa (figure 2). It shows that the enzyme substrate reaction reached to maximum value in 20 min and 30 min for N. sativa and C. longa, respectively, later no change in activity was observed. Figure 3 illustrates temperature optima of protease isolated from N. sativa and C. longa, showing maximum activity at 40°C, in both the cases. Increase in temperature above optimum level affects important factors like protein denaturation, protein ionization state and solubility of species in solution reducing enzyme activity [26]. Enzymes, being proteinaceous in nature, have properties that are quite pH sensitive. pH can affect activity by changing the charges on an amino acid residue which is functional in substrate binding or catalysis [27]. The pH optima of protease from N. sativa and C. longa were found to be (Figure 4) 5.0, in both the cases. Similar observation was reported in metalloprotease production in P. fluorescens [28] and Adhatoda vasica [29] with an optimum activity at pH 5.0. The proteases produced by isolates with enzymatic activity optima at pH 5.0 could be used to coagulate milk proteins for the dairy industry, as debittering agents in cheese and in peptide synthesis [30]. Figure 5 shows the pH stability curve of protease from N. sativa and C. longa. The protease from N. sativa was found to retain its activity in the pH range of 5.0 to 9.0 whereas respective values were from 4.0 to 8.0 in C. longa. The most significant result is the heat resistant property of the enzymes obtained from the two species, the temperature values being 60°C and 50°C for Nigella sativa and Curcuma longa (figure 6), respectively. Proteases from various sources with temperature stability upto 50°C have been reported by various workers [31-36]. Thus, it is imperative to state that spices may serve as good source of thermostable proteases which further have great importance in therapeutics and many other industries.
IV. Conclusion
In the present work out of the nine spices studied, Nigella sativa and Curcuma longa were found to be good source of protease enzyme. Very limited work has been done on enzymes from these spices which call for a more detailed research in future. Proteases obtained from spices may serve as potential candidate for targeting many diseases. From the results obtained, it may be concluded that spices which are of great medicinal and culinary importance can be explored for heat resistant protease; an enzyme of immense value in therapeutics, chemical, leather, food and other bio-industries.
Acknowledgment
We are grateful to Director, Amity Institute of Biotechnology, Amity University Uttar Pradesh, Noida, Uttar Pradesh, for his constant support and encouragement during this study.
References
[1] M. B. Rao, A. M. Tanksale, M. S. Ghatge, and V. V. Deshpande, “Molecular and biotechnological aspects of microbial proteases”, Microbiol. Mol. biol. Rev., 1998, pp. 597–600.
[2] N. Jabalia, P. C. Mishra, and N. Chaudhary, “Applications, Challenges and Future Prospects of Proteases: An Overview”, Journal of Agroecology and Natural Resource Management, 2014, pp. 179–183.
[3] Ishikawa, K. Ishimi, M. Sugiura, A. Sowa, and N. Fujiwara, “Kinetics and mechanism of enzymatic hydrolysis of gelatin layers of X-ray film and release of silver particles”, J. Ferm. Bioeng., 1993, pp. 300–305.
[4] Sumantha, C. Sandhya, G. Szakacs, C. R. Soccol, and A. Pandey, “Production and partial purification of a neutral metalloprotease by fungal mixed substrate fermentation.”, Food Technology and Biotechnology, 2005, pp. 313–319.
[5] M. Madan, S. Dhillon, and R. Singh, “Production of alkaline protease by a UV mutant of Bacillus polymyxa”, Indian J. Microbiol., 2002, pp. 155–159.
[6] D. Anandan, W. Marmer, and R. L. Dudley, “Isolation, characterization and optimization of culture parameters for production of an alkaline protease isolated from Aspergillus tamarii”, J. Ind. Microbiol. Biotechnol., 2007, pp. 339–47.
[7] R. Chakraborty, M. S. Srinivasan, and S. K. Raghwan, “Production of acid proteases by a new Aspergillus niger during solid substrate fermentation”, J. Microbiol., Biotechnol., 1995, pp. 17–30.
[8] M. S. Thakur, N.G. Karant, and K. Nand, “Production of fungal rennet by Mucor Miehi using solid state fermentation”, Appl., Microbiol. Biotechnol., 1990, pp. 409–413.
[9] M. R. Khan J. A. Blain, and J. D. E. Patterson, “Intracellular protease of Mucor Pusillus. ” J. Biochem., 1979, pp. 719–724.
[10] K. Tuschiya, T. Arai, K. Seki, And T. Kimeua, “Purification and some properties of alkaline proteinases from Cephalosporium sp. KM 388”, J. Argic. Biol. Chem., 1987, pp. 2959–2965.
[11] L. Ikasari, and D. A. Mitchell, “Protease production by Rhizopus oligosposus in solid state fermentation”, Microbiol. Biotechnol., 1994, 10, pp. 320–324.
[12] P.L. Bergquiest, Jr. V. S Te'O, M. D. Gibbs, A. C. E. Cziferszky, Defarria, M. O. Azevedo, and K. M. H. Nevalainen K. M. H., “Proudction of recombinant bleaching enzymes from thermophilic microorganisms in fungal hosts”, Appl. Biochem. Biotechnol., 2002, pp. 165–176.
[13] Balachandran, V. Duraipandiyan and S. Ignacimuthu, “Purification and characterization of protease enzyme from actinomycetes and its cytotoxic effect on cancer cell line (A549)”, Asian Pacific Journal of Tropical Biomedicine, 2012, pp. 392–400.
[14] H. S. Joo, and C. S. Chang, “Production of an oxidant and SDS-stable alkaline protease from an alkaophilic Bacillus clausii I-52 by submerged fermentation: Feasibility as a laundry detergent additive”, Enzyme microbial, 2006, pp. 176–183.
[15] N. S. Nisha, and J. Divakaran, “Optimization of alkaline protease production from Bacillus substilis NS isolated from sea water”, African Journal of Biotechnology, 2014, pp. 1707–1713.
[16] Tunga, B. Shrivastava, and R. Banerjee, “Purification and characterization of a protease from solid state cultures of Aspergillus parasiticus,” Process Biochemistry, 2003, pp. 1553–1558.
[17] F. J. Ustariz, A. Laca, L. A. Garcia, and M. Diaz, “Fermentation of individual proteins for protease production by Serratia marcescens”, Biochem. Engr. J., 2004, pp. 147–153.
[18] N. Vanitha, S. Rajan, and A. G. Murugesan, “Optimization and production of alkaline protease enzyme from Bacillus substilis 168 isolated from food industry waste”, International Journal of Current Microbiology and Applied Sciences, 2014, pp. 36–44.
[19] J. S. Sharmila, R. L. Jeyanthi, M. P. Das, and Md. “Saduzzaman, “Isolation and partial purification of Protease from plant leaves”, Journal of Chemical and Pharmaceutical Research, 2012, pp. 3808–3812.
[20] O. H. Lowry, M. J. Rosebrogh, A. L. Farr, and R. J. “Randall, “Protein measurement with folin reagent”, Journal of Biological Chemistry, 1951, pp. 265–275.
[21] F. Abidi, F. Limamm, and M. M. Nejib, “Production of alkaline proteases by Botrytis cinerea using economic raw materials: Assay as biodetergent”, Proc. Biochem., 2008, pp. 1202–12.
[22] H. Takami, T. Alaba, and K. Horikosha, “Production of extremely thermostable alkaline protease from Bacillus Sp. No. AH-101”, Applied Microbiology and Biotechnology, 1989, pp. 120–124.
[23] G. Gomori, “Preparation of buffers for use in enzyme active studies, in methods in Enzymology”, 1955.s
[24] P. D. Brown, “Clinical trials of a low molecular weight matrix metalloproteinase inhibitor in cancer”, Annals of the New York Academy of Sciences, 1994, pp. 217–221.
[25] Sandhya, A. Sumantha, G. Szakacs, and A. Pandey, “Comparative evaluation of neutral protease production by Aspergillus oxyzae in submerged and solid state fermentation”, Process Biochemistry, 2005, pp. 2689–2694.
[26] Zefferen, and P. L. Hall, “Kinetics I and Kinetics II, in the study of enzyme mechanisms”, John Wiley and Son Inc. NewYork, 1973, pp. 53–99.
[27] N. Sharma, and S. Tripathi, “Kinetics study of free and immobilized protease from Aspergillus sp. ”, Journal of Pharmacy and Biological Sciences, 2013, pp. 86–96.
[28] R. Koka, and B.C. Weimer, “Isolation and characterization of a protease from Pseudomonas fluorescens RO98”, Journal of Applied Microbiology, 2000, pp. 280–288.
[29] B. Khurana, A. Mishra, N. Jabalia, and N. Chaudhary, “Various Biochemical Parameters of Protease Isolated From Adhatoda Vasica: A Medicinally Important Plant”, International Journal of Genetic Engineering and Biotechnology, 2014, pp. 1–6.
[30] Sumantha, C. Larroche, and A. Pandey “Microbiology and industrial biotechnology of food-grade proteases: a perspective”, Food Technology and Biotechnology, 2006, pp. 211–220.
[31] Durham, D.R., D.B. Stewart, and E.J. Stellwag, “Novel alkaline and heat stable serine proteases from Bacillus sp. Strain GX 6638”, J. Bacteriolgy, 1987, 169, pp. 2762–2768.
[32] Takil, Y., N. Kuriyama, and Y. Suzuki, 1990. “Alkaline serine protease produced from citric acid by Bacillus alkalophilus sub sp. halodurans Kp 1239”, Appl. Microbiol. Biotechnol., 1990, pp. 57–62.
[33] Kobayashi, T., A. Ogasawara, S. Ito, and M. Saitoh, “Purification and some properties of alkaline proteinase produced by Pseudomonas maltophilia”, Agric. Biol. Chem., 1995, pp. 693–698.
[34] Kobayashi, T., Y. Hakamada, J. Hitomi, K. Koike, and S. Ito, “Purification of alkaline proteases from a Bacillus strain and their possible interrelationship”, Appl Microbio. Biotechnol., 1996, pp. 63–71.
[35] Ferrero, M. A., G. R. Abate, C. M. Baigori, and F. Sineriz, “Thermostable alkaline protease of Bacillus licheniformis MIR 29: isolation, production and characterization”, Appl. Microbiol. Biotechnol., 1996, pp. 327–332.
[36] G. Lakshmi, and N. N. Prasad, “Purification and Characterization of Alkaline Protease from a Mutant Bacillus licheniformis Bl8”, Advances in Biological Research, 2015, pp. 15–23.
Author information
Authors and Affiliations
Additional information
Authors’ profile

Dr. Nidhee Chaudhary became a member of GSTF in 2012 and Program Committee Member in 2013. She became Reviewer-in-charge of GSTF Journal of biosciences in 2013. She is a the Head of Centre of Biotechnology and Biochemical Engineering at Amity Institute of Biotechnology, Amity University (AUUP), Noida, India. Previously she was the Faculty and Director, Department of Biotechnology, D. A. V. (PG) College, Muzaffarnagar. She has been in academia for over 18 years at various positions. She has more than 18 years of research experience and has supervised several Ph. D. (theses), M. Phil., M. Sc., M. Tech. and B. Tech. dissertations in the field of Enzymology and Bioprocess Technology. She has many international and national publications in journals and books of repute.
Dr. Chaudhary is member of various scientific groups like: The Protein Society, USA, American Society of Microbiology, Society of Biological Chemists, Society of Agricultural Biochemists, OMICS group, USA, Biotech Research Society, India (BRSI), Biotech Society of India (BSI), World Academy of Science, Engineering and Technology (WASET), USA, and Asian Federation of Biotechnology (AFOB). She has served as reviewer for several reputed Journals like; Protein Science, Journal of Biotechnology and Applied Biochemistry, Journal of Preparative Biochemistry and Biotechnology, Advances in Bioscience and Biotechnology, Journal of Biotechnology, published by Elsevier, Taylor & Francis, Wiley-Blackwell and other well established groups. Recently, she has been granted with Finn Wold Travel Award by The Protein Society, USA.

Ms. Neetu Jabalia is currently an Assistant Professor in the Centre for Bioinformatics at Amity University Uttar Pradesh, Noida, India. She obtained her B.Sc. in Biotechnology from Guru NanaK Dev University, Amritsar, India in 2005. She received her M.Sc. on Bioinformatics from Panjab University, Chandigarh, India in 2007. She is currently pursuing her Ph.D in Bioinformatics.
This article is distributed under the terms of the Creative Commons Attribution License which permits any use, distribution, and reproduction in any medium, provided the original author(s) and the source are credited.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made.
The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder.
To view a copy of this licence, visit https://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Jabalia, N., Chaudhary, N. Heat Resistant Proteases Isolated from Medicinally Significant Spices: Partial Purification and Biochemical Characterization. GSTF J Biosci 3, 5 (2015). https://doi.org/10.7603/s40835-014-0005-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.7603/s40835-014-0005-8