Abstract
Metabolic reprogramming plays a crucial role in the development of cancer. The aim of this study was to explore the effect of fenofibrate, an agonist of peroxisome proliferator-activated receptor alpha (PPARα), on gene expression profiles of mitochondrial energy metabolism. Our results showed that PPARα expression was negatively correlated with tumor progression in an oral cancer mouse model. Activation of PPARα through fenofibrate suppressed migration of oral cancer cells. Differential protein profiling demonstrated that expressions of genes related to mitochondrial energy metabolism were either up-regulated (Atp5g3, Cyc1, Ndufa5, Ndufa10, and Sdhd) or down-regulated (Cox5b, Ndufa1, Ndufb7, and Uqcrh) through PPARα activation and response. Our results indicate that PPARα exhibits a great potential for anti-oral cancer therapies by modulating cancer cell mitochondrial energy metabolism.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1. Introduction
One of the prominent characteristics of rapidly growing cancer cells is their capacity to sustain high rates of glycolysis for ATP production regardless of whether oxygen is present—a phenomenon known as the Warburg effect [1]. Activators of peroxisome proliferator-activated receptors (PPARs) have been shown to exhibit a great potential for anticancer therapies by modulating cancer cell energy metabolism and signaling pathways [2, [3]. PPARα is known to modulate the expression of genes regulating glucose and lipid metabolism [4, [5]. Fenofibrate is a synthetic agonist of PPARα and a widely used hypolipidemic drug with antiinflammatory and anti-atherosclerotic effects in humans [6, [7]. Fenofibrate has been reported to be involved in several anticancer activities, including induction of apoptosis, reduction of the proliferation rate, attenuation of IGF-1 receptor signaling, inhibition of tumor angiogenesis, and suppression of the inflammatory response and oxidative stress in cancer cells like melanoma, mantle cell lymphoma, medulloblastoma, glioma, and endometrial cancer cells [8-[14]. However, the anticancer activity of fenofibrate in energy homeostasis is not well clarified or understood.
Our previous studies suggest a beneficial role of fenofibrate in anti-oral tumorigenesis in both cell culture and animal models. Fenofibrate inhibits the invasion and migration of CAL 27 oral cancer cells by suppressing the protein expressions of matrix metalloproteinase-1 (MMP-1), MMP-2, MMP-7, and MMP-9 through the AMPK and NF-κB signaling pathway [15]. Fenofibrate causes the reduction in the incidence and size of squamous carcinoma and suppresses the progression of the preneoplastic lesion into squamous cell carcinoma in an oral-specific 4-nitroquinoline 1-oxide/arecoline mouse model [16]. Recently, we demonstrated that fenofibrate provided novel mechanisms for delaying oral tumor development via the reprogramming of metabolic processes by interrupting the binding of hexokinase II to the voltage-dependent anion channel and increasing metabolites in the tricarboxylic acid cycle (unpublished data). It is possible that part of the anticancer mechanisms of fenofibrate might involve regulating the gene expression of mitochondrial energy metabolism. Therefore, this study focused on the changes in genes expression of mitochondrial energy metabolism in oral cancer cells treated with fenofibrate. The protein expression levels of PPARα were also examined in oral tumor progression.
2. Materials and methods
2.1. Administration of 4-NQO and arecoline
Seventy 6-week-old male C57BL/6JNarl mice were purchased from the National Laboratory Animal Center. The mice were handled in accordance with the Animal Care and Use Guidelines of the China Medical University, and the study protocol was approved by the Institutional Animal Care Use Committee. The experiments were controlled as previously described [17]. The carcinogens, 200 μg/ml 4-nitroquinoline 1-oxide (4-NQO; Sigma- Aldrich, St. Louis, MO, USA) and 500 μg/ml arecoline hydrobromide (Tokyo Chemical Industry Co. LTD, Tokyo, Japan), were used to induce oral tumorigenesis in the mice for a period of 8 weeks and then observed for the indicated time including 0, 8, 12, 16, 20, 24, and 28 weeks. Ten mice were euthanized at each indicated time. During the administration, all mice were allowed to access the drinking water and chow diet (Prolab® RMH 2500 PMI Nutrition International, LLC, MO, USA) ad libitum. The experiments were carried out under controlled conditions with a 12-h light/dark cycle.
2.2. Cell culture and wound healing assay
The 28-week mouse oral cancer cells were obtained from primary cultured cells of tongue cancer induced by 4-NQO (200 μg/ml) and arecoline (500 μg/ml) [16]. Cells were grown in DMEM supplemented with 10% fetal bovine serum, 1% antibiotic-antimycotic and 2 mM L-glutamine (Gibco; Life Technology Corporation, NY, USA) at 37°C in a humidified 5% CO2 atmosphere incubator. For migration assay, 5 × 104 cells were seeded in ibidi Culture-Insert (ibidi GmbH, Martinsried, Germany) on a 24 well plate. After appropriate cell attachment, the Culture-Insert was gently removed, and cells were then incubated with or without fenofibrate (50 μM). Next cells were allowed to migrate into the wound area for 0, 16, 24, and 38 h. Cells in the wound area were photographed and measured at an image analysis platform. (http://ibidi.com/applications/wound-healing-and-migration/)
2.3. RNA Extraction and RT2 Profiler PCR Array examination
The 28-week mouse oral cancer cells were incubated with 50 μM fenofibrate for 18 h, and an equivalent volume of 0.1% DMSO was used as control. Total RNA was extracted from the cells with or without fenofibrate treatment by using the Trizol reagent solution (Ambion, Life Techniologies) according to the manufacturer’s recommendations. The quality of RNA samples was determined by Agilent 2100 Bioanalyzer analysis using a RNA 6000 Nano Kit (Agilent Technologies, Inc.). The quantity of RNA samples was determined using a NanoDrop ND-1000 spectrophotometer (Thermo Scientific). c-DNA was converted using a RE3 Reverse Transcriptase Mix (Qiagen Ltd.). Quantitative PCR was performed according to the RT2 Profiler PCR array instructions on a 7300 Real-Time PCR System (Applied Biosystems, CA, USA). The Mouse Mitochondrial Energy Metabolism (PAMM-008Z) RT2 Profiler PCR Array, which profiles a total of 84 genes expression, was purchased from Qiagen Ltd. The PCR cycling condition was set as follows: 1 cycle of 95°C for 10 min; 40 cycles of 95°C for 15 sec and 60°C for 1 min; and 1 cycle of 95°C for 15 sec, 60°C for 30 sec, and 95°C for 15 sec. The results of Ct values were submitted to the Web-based PCR array data analysis software, which is available at www.SABioscience.com/pcrarraydataanalysis.php.
2.4. Protein extraction and Western blotting
To analyze PPARα protein levels in mouse tongue tissue, each sample at the indicated time was homogenized and proteins were extracted as previously described [18]. Total protein contents were determined using a Bio-Rad protein assay reagent (Bio- Rad Laboratories, CA, USA) with bovine serum albumin as a standard. Equal amounts (50 μg) of extracted proteins from each condition were fractionated by 12% SDS-Polyacryamide gels and transferred onto a polyvinylidene difluoride (PVDF) membrane (Millipore, MA, USA). The Western blots were probed with a PPARα primary antibody (Santa Cruz Bioechnology, TX, USA) and then incubated with horseradish peroxide-conjugated secondary antibodies. The immunoreactive protein bands were detected by a SuperSignal Enhanced Chemiluminescence (Millipore, MA, USA). Western blots were quantified by densitometric analysis using KODAK image analysis software (Kodak EDAS290, Eastman Kodak, Rochester, NY, USA).
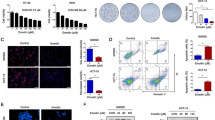
The PPARα protein expression levels in tumor progression. The protein levels of PPARα in each mouse tongue tissue were measured by Western blot analysis at the indicated time (8, 12, 16, 20, 24, and 28 weeks). Data are shown as the fold of control (0 week). Each dot represents a mouse tongue tissue. A statistical analysis of the PPARα expression was performed by Linear regression analysis. The results showed that PPARα expression was negatively correlated with tumor progression (P = 0.003).
2.5. Statistical analysis
All data were analyzed using SPSS 15.0 for Windows. Statistical analyses of the PPARα expression were performed using Linear regression analysis. A P-value of less than 0.05 was considered statistically significant.
3. Results
3.1. PPARα protein expression levels in tongue tissues
First, we determined the PPARα protein levels of tongue tissues in an 4-NQO/arecoline induced mouse model. The PPARα protein levels at each time point are shown in Figure 1. The PPARα protein levels were progressively decreased in a time-dependent manner over a 28-week observation period when compared with the control (0 week). Linear regression analysis showed that PPARα expression was negatively correlated with the advancing of tumor development (P = 0.003).
3.2. Activation of PPARα suppressed migration of oral cancer cells
We next assessed whether activation of PPARα influenced cancer cell migration. The 28-week mouse oral cancer cells were treated with fenofibrate (0 or 50 μM) for 0, 16, 24, and 38 h. We found that fenofibrate decreased cell migration ability up to the observation time of 38 h (Figure 2).
Fenofibrate inhibits the migration of oral cancer cells. The 28-week mouse oral cancer cells were treated with fenofibrate (0 and 50 μM) for 0, 16, 24, and 38 h. Cell migration was examined by wound healing assay as described in the Materials and Methods section. Cells in the wound area were photographed (Magnification, 100×). The migration ability of cells in the wound area was suppressed after treating with fenofibrate.
3.3. mRNA expression levels of genes involved in mitochondrial energy metabolism after fenofibrate treatment
To explore the expression levels of genes involved in mitochondrial energy metabolism through PPARα activation and response, oral cancer cells from the 28-week group were treated with or without 50 μM fenofibrate for 18 h. Table 1 shows the changes in the mRNA expression levels of 9 genes involved in mitochondrial energy metabolism detecting by an RT2 Profiler PCR Array. Fold-change (2-ΔΔCt) is measured as the level of normalized gene expression (2-ΔCt) in the fenofibrate-treated group divided by the level of normalized gene expression (2-ΔCt) in the control group. Fold-change values greater than one indicate a positive- or an upregulation; however, fold-change values less than one indicate a negative or down-regulation. The expression levels of Atp5g3 (ATP synthase, H+ transporting, mitochondrial F0 complex, subunit C3), Cyc1 (Cytochrome c-1), Ndufa5 (NADH dehydrogenase (ubiquinone) 1 alpha subcomplex, 5), Ndufa10, and Sdhd (Succinate dehydrogenase complex, subunit D, integral membrane protein) genes were up-regulated in fenofibrate-treated cells compared with those in control cells. Conversely, expressions of Cox5b (Cytochrome c oxidase, subunit Vb), Ndufa1, Ndufb7 (NADH dehydrogenase (ubiquinone) 1 beta subcomplex, 7), and Uqcrh (Ubiquinol-cytochrome c reductase hinge protein) genes were down-regulated. These findings indicate that PPARα activation and response might modulate mitochondrial function and energy production.
4. Discussion
It is well known that cancer cells undergo significant metabolic adaptations in energy metabolism [1]. Shifting the balance between the glycolytic and mitochondrial processes could be an important point in cancer therapy. In this study, we found that PPARα protein levels were negatively associated with tumor development in an oral cancer mouse model (Figure 1). Activation of PPARα by fenofibrate decreased the migration ability of oral cancer cells in vitro (Figure 2) and suppressed the tumor progression into squamous cell carcinoma in vivo [16]. We thought that these molecular events might be linked to interrupting the Warburg effect through reprogramming ATP production pathway. Therefore, the transcription levels of genes involved in mitochondrial energy metabolism were determined by an RT2 Profiler PCR Array. We found that nine genes’ transcription levels in the respiratory chain, including Ndufa1, Ndufa5, Ndufa10, Ndufb7, Sdhd, Cyc1, Uqcrh, Cox5b, and Atp5g3, were changed in the fenofibrate-treated cells (Table 1).
It has been shown that Ndufa genes play an essential role in the assembly pathway and function of NADH dehydrogenase (Complex I) in mammals [19]. Fenofibrate induced high expression levels of Ndufa5 and Ndufa10, and low expression levels of Ndufa1 (Table 1). Ndufa1 was shown to interact with the subunits encoded by mitochondrial DNA during the Complex I assembly process [20]. Two missense mutations (G8R and R37S) in Ndufa1 have been identified in male patients presenting with neurological syndromes [21]. Ndufa5 is involved in building up the electrochemical potential required to produce ATP. To date, no mutations in Ndufa5 are reported in association with any diseases. Nevertheless, low expression levels are found in the brain regions of those affected by autism [22, [23]. Mutations in Ndufa10 lead to lowering holo-complex I levels with the accumulation of complex I subcomplexes, which indicates a disturbance in the assembly and/or stability of complex I [24]. The Ndufb7 gene is classified in the hydrophobic group with NADH-binding and oxidizing properties [19, [25]. To a point, Ndufa1, Ndufa5, Ndufa10, and Ndufb7 genes are likely to contribute a significant impact to PPARα response. However, function of the above 4 genes in the regulation of mitochondrial energy production and cancer development needs further investigation.
Additionally, fenofibrate up-regulated the expression of Sdhd, Cyc1, and Atp5g3 genes. Succinate dehydrogenase (SDH) or Complex II is part of both the Krebs cycle and the electron transport chain [26]. The SDHD protein is the membrane-anchoring protein that contains one heme and is essential for ubiquinone binding. The Mitochondrial Sdhd gene is required for early embryogenesis [27]. Mutations in the Sdhd gene are associated with paraganglioma [28]. The cytochrome c1 (CYC1) protein, a subunit of respiratory chain complex III, directly interacts with cytochrome c and mediates electron transport from cytochrome b to cytochrome c during oxidative phosphorylation [29]. The Cyc1 gene plays an important role in the development of nasopharyngeal carcinoma and osteosarcoma [30, [31]. The mitochondrial ATP synthase, subunit c, isoform 3 gene (Atp5g3) encodes subunit 9, which catalyzes ATP synthesis during oxidative phosphorylation in mitochondria. A recent report shows that the expression level of Atp5g3 is reduced in autism patients [32]. In the present study, the up-regulated expressions of the Sdhd, Cyc1, and Atp5g3 genes are consistent with metabolic reprogramming, which might switch the Warburg effect to oxidative phosphorylation with slowing energy production rate and inhibiting cancer cells growth in fenofibrate-treated cells. Recently, we have demonstrated that fenofibrate inhibits the invasion and migration of CAL 27 oral cancer cells through the AMPK and NF-κB signaling pathway [15].
Cox5b is an extra-membrane subunit located at the matrix side of the complex IV that facilitates the oxidation of cytochrome c by O2. Cox5b has been shown to play a role in suppressing ROS production [33, [34] and provide a binding site for protein kinase A [35]. Uqcrh is a component of the ubiquinol-cytochrome c reductase complex (complex III), which catalyzes electron transfer from succinate and nicotinamide adenine dinucleotide-linked dehydrogenases to cytochrome c [36]. Uqcrh is a downstream target gene of PGC-1α (Peroxisome proliferator-activated receptor gamma coactivator 1 alpha). The down-regulation of the Uqcrh gene suggests the decrease in mitochondrial oxidative phosphorylation activity, which results in reducing ROS accumulation and creating an antioxidant feedback [37]. Therefore, we suggest that the low expressions of Cox5b and Uqcrh in fenofibrate-treated cells are likely to contribute a significant impact to regulating ROS production involved in oral cancer development. Further investigation is needed to elucidate the association between ROS production and PPARα signaling.
5. Conclusions
The protein levels of PPARα were negatively associated with oral tumor progression. Activation of PPARα inhibited oral cancer cell migration. The molecular mechanism may be linked to modulate the expression of genes involved in mitochondrial energy metabolism.
References
Vander Heiden MG, Cantley LC, Thompson CB. Understanding the Warburg effect: the metabolic requirements of cell proliferation. Science 2009; 324: 1029–33.
Vamecq J, Colet JM, Vanden Eynde JJ, Briand G, Porchet N, Rocchi S. PPARs: interference with Warburg’ effect and clinical anticancer trials. PPAR Res 2012; 304760.
Grabacka M, Pierzchalska M, Reiss K. Peroxisome proliferator activated receptor α ligands as anticancer drugs targeting mitochondrial metabolism. Curr Pharm Biotechnol 2013; 14: 342–56.
Varet J, Vincent L, Mirshahi P, Pille JV, Legrand E, Opolon P, et al. Fenofibrate inhibits angiogenesis in vitro and in vivo. Cell Mol Life Sci. 2003; 60: 810–9.
Finck BN, Kelly DP. Peroxisome proliferator-activated receptor α (PPARα) signaling in the gene regulatory control of energy metabolism in the normal and diseased heart. J Mol Cell Cardiol 2002; 34: 1249–57.
Fruchart JC. Peroxisome proliferator-activated receptor-alpha activation and high-density lipoprotein metabolism. Am J Cardiol 2001; 88: 24–9.
Staels B, Dallongeville J, Auwerx J, Schoonjans K, Leitersdorf E, Fruchard JC. Mechanism of action of fibrates on lipid and lipoprotein metabolism. Circulation 1998; 98: 2088–93.
Saidi SA, Holland CM, Charnock-Jones DS, Smith SK. In vitro and in vivo effects of the PPAR alpha agonists fenofibrate and retinoic acid in endometrial cancer. Mol Cancer 2006; 5: 13.
Zak Z, Gelebart P, Lai R. Fenofibrate induces effective apoptosis in mantle cell lymphoma by inhibiting the TNFalpha/NF-kappaB signaling axis. Leukemia 2010; 24: 1476–86.
Drukala J, Urbanska K, Wilk A, Grabacka M, Wybieralska E, Del Valle L, et al. ROS accumulation and IGF-IR inhibition contribute to fenofibrate/PPARalpha-mediated inhibition of Glioma cell notility in vitro. Mol Cancer 2010; 9: 159.
Urbanska K, Pannizzo P, Grabacka M, Croul S, Del Valle L, Khalili K, et al. Activation of PPARalpha inhibits IGF-I-mediated growth and survival responses in medulloblastoma cell lines. Int J Cancer 2008; 123: 1015–24.
Varet J, Vincent L, Mirshahi P, Pille JV, Legrand E, Opolon P, et al. Fenofibrate inhibits angiogenesis in vitro and in vivo. Cell Mol Life Sci 2003; 60: 810–9.
Panigrahy D, Kaipainen A, Huang S, Butterfield CE, Barnes CM, Fannon M, et al. PPAR alpha agonist fenofibrate suppresses tumor growth through direct and indirect angiogenesis inhibition. Proc Natl Acad Sci USA 2008; 105: 985–90.
Grabacka M, Placha W, Plonka PM, Pajak S, Urbanska K, Laidler P, et al. Inhibition of melanoma metastases by fenofibrate. Arch Dermatol Res 2004; 296: 54–8.
Tsai SC, Tsai MH, Chiu CF, Lu CC, Kuo SC, Chang NW, et al. AMPK-dependent signaling modulates the suppression of invasion and migration by fenofibrate in CAL 27 oral cancer cells through NF-κB pathway. Environ Toxicol 2014; http://dx.doi.org/10.1002/tox.22097.
Chang NW, Tsai MH, Lin C, Hsu HT, Chu PY, Yeh CM, et al. Fenofibrate exhibits a high potential to suppress the formation of squamous cell carcinoma in an oral-specific 4-nitroquinoline 1-oxide/ arecoline mouse model. Biochim Biophys Acta 2011; 1812: 558–64.
Chang NW, Pei RJ, Tseng HC, Yeh KT, Chan HC, Lee MR, et al. Co-treating with arecoline and 4-nitroquinoline 1-oxide to establish a mouse model mimicking oral tumorigenesis. Chem Biol Interact. 2010; 183: 231–7.
Chang NW, Wu CT, Chen DR, Yeh CY, Lin C. High levels of arachidonic acid and peroxisome proliferator-activated receptoralpha in breast cancer tissues are associated with promoting cancer cell proliferation. J Nutr Biochem. 2013; 24: 274–281.
Koene S, Willems PHGM, Roestenberg P, Koopman WJH, Smeitink JAM. Mouse models for nuclear DNA-encoded mitochondrial complex I deficiency. J Inherit Metab Dis 2011; 34: 293–307.
Yadava N, Houchens T, Potluri P, Scheffler IE. Development and characterization of a conditional mitochondrial complex I assembly system. J Biol Chem 2004; 279: 12406–13.
Potluri P, Davila A, Ruiz-Pesini E, Mishmar D, O’Hearn S, Hancock S, et al. A novel NDUFA1 mutation leads to a progressive mitochondrialcomplex I-specific neurodegenerative disease. Mol Genet Metab 2009; 96: 189–95.
Marui T, Funatogawa I, Koishi S, Yamamoto K, Matsumoto H, Hashimoto O, et al. The NADH-ubiquinone oxidoreductase 1 alpha subcomplex 5 (NDUFA5) gene variants are associated with autism. Acta Psychiatr Scand 2011; 123: 118–24.
Peralta S, Torraco A, Wenz T, Garcia S, Diaz F, Moraes CT. Partial complex I deficiency due to the CNS conditional ablation of Ndufa5 results in a mild chronic encephalopathy but no increase in oxidative damage. Hum Mol Genet 2013; 23: 1399–412.
Hoefs SJ, van Spronsen FJ, Lenssen EW, Nijtmans LG, Rodenburg RJ, Smeitink JA, et al. NDUFA10 mutations cause complex I deficiencyin a patient with Leigh disease. Eur J Hum Genet 2011; 19: 270–4.
Mimaki M, Wang X, McKenzie M, Thorburn D, Ryan M. Understanding mitochondrial complex I assembly in health and disease. Biochim Biophys Acta 2012; 1817: 851–62.
Hederstedt L. Complex II is complex too. Science 2003; 299: 671–2.
Piruat JI, Pintado CO, Ortega-Saenz P, Roche M, Lopez-Barneo J. The mitochondrial SDHD gene is required for early embryogenesis, and its partial deficiency results in persistent carotid body glomus cell activation with full responsiveness to hypoxia. Mol Cell Biol 2004; 24: 10933–40.
Baysal BE, Ferrell RE, Willett-Brozick JE, Lawrence EC, Myssiorek D, Bosch A, et al. Mutations in SDHD, a mitochondrial complex II gene in hereditary paraganglioma. Science 2000; 287: 848–51.
Crofts AR. The cytochrome bc1 complex: function in the context of structure. Annu Rev Physiol 2004; 66: 689–733.
Liu Z, Wang HY, Long J, Fang WY. Construction of a lentivirus interfering vector targeting Cyc1 and its interfering efficiency in nasopharyngeal carcinoma cells. Nan Fang Yi Ke Da Xue Xue Bao 2010; 30: 2629–32.
Li G, Fu D, Liang W, Fan L, Chen K, Shan L, et al. CYC1 silencing sensitizes osteosarcoma cells to TRAIL-induced apoptosis. Cell Physiol Biochem 2014; 34: 2070–80.
Anitha A, Nakamura K, Thanseem I, Matsuzaki H, MiyachiT, Tsujii M, et al. Downregulation of the expression of mitochondrial electron transport complex genes in autism brains. Brain Pathol 2013; 23: 294–302.
Galati D, Srinivasan S, Raza H, Prabu SK, Hardy M, Chandran K, et al. Role of nuclear-encoded subunit Vb in the assembly and stability of cytochrome c oxidase complex: implications in mitochondrial dysfunction and ROS production. Biochem J 2009; 420: 439–49.
Campian JL, Gao X, Qian M, Eaton JW. Cytochrome C oxidase activity and oxygen tolerance. J Biol Chem 2007; 282: 12430–8.
Yang WL, Iacono L, Tang WM, Chin KV. Novel function of the regulatory subunit of protein kinase A: regulation of cytochrome c oxidase activity and cytochrome c release. Biochem 1998; 37: 14175–80.
Mitchell P. Possible molecular mechanisms of the protonmotive functionof cytochrome systems. J Theor Biol 1976; 62: 327–67.
Zaza G, Granata S, Masola V, Rugiu C, Fantin F, Gesualdo L, et al. Downregulation of nuclear-encoded genes of oxidative metabolism in dialyzed chronic kidney disease patients. PLoS One 2013; 8: e77847.
Acknowledgements
This study was supported by a research grant from the China Medical University (CMU104-S-13), Taiwan, ROC.
Author information
Authors and Affiliations
Corresponding author
Additional information
aDepartment of Physiology, College of Medicine, China Medical University, Taichung 404, Taiwan
bDepartment of Biochemistry, College of Medicine, China Medical University, Taichung 404, Taiwan
*Corresponding author. Department of Biochemistry, College of Medicine, China Medical University, No. 91, Hsueh-Shih Road, Taichung 404, Taiwan.
E-mail address: nwchang@mail.cmu.edu.tw (N.W. Chang).
Declaration of interest
The authors declare no conflicts of interest for this work.
Open Access This article is distributed under terms of the Creative Commons Attribution License which permits any use, distribution, and reproduction in any medium, provided original author(s) and source are credited.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made.
The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder.
To view a copy of this licence, visit https://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Huang, YP., Chang, N. PPARα modulates gene expression profiles of mitochondrial energy metabolism in oral tumorigenesis. BioMed 6, 3 (2016). https://doi.org/10.7603/s40681-016-0003-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.7603/s40681-016-0003-7