Abstract
This paper reports brief systematization of the current knowledge of the biology of mycoviruses, viral morphology, and genetics in particular as well as characteristics of the virus transmission and infection symptoms in fungal cells. The mechanisms involved in antiviral defense in the members of different classes of fungi are discussed. Insights into the role of hypovirulent mycoviruses in the biotechnological control of phytopathogenic fungi are provided.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Mycoviruses are viruses that infect fungi and are able to replicate only within the cells of filamentous fungi, yeasts, and oomycetes [1]. The viruses of fungi were discovered much later than the viruses of plants and animals. They remain the least studied viruses, with few exceptions. This may be explained by their limited economic significance [1, 2], by latency of the majority of mycoviral infections, and relatively small number of laboratories focused on the research of the kingdom of fungi [3].
Nevertheless, the interest in mycoviruses has increased in recent decades, and considerable acceleration of their study is, therefore, observed. First of all, this is associated with the fact that many mycoviruses cause phenotypic changes in their host fungi, and this characteristic may be valuable enough for both fundamental research and practical use. The interest in these pathogens is also stimulated by the technical achievements in the field of molecular biology of mycoviruses. At the same time, viruses of fungi are successfully used as tools to study the mechanisms for interaction and control of the host.
The first evidence of the existence of lysogenic viruses was presented for yeasts in 1936 [2], while the first viral disease of the higher fungi (Agaricus bisporus basidiomycetes) was first revealed in 1950 [3]. The fruiting bodies of the infected fungi were deformed; they had brown spots, grew slowly, and ripened early, which resulted in severe crop loss. The disease was named “la France disease,” and its viral origin was revealed in 1962 [4]. La France isometric virus (LFIV) is periodically observed on different continents under the conditions of industrial champignon cultivation [5–7]. Mycoviruses causing diseases of other commercially significant basidiomycetes Flammulina velutipes [8] and Lentinula edodes [9], cultivated primarily in Japan and China, were isolated.
In 1958, phytopathogenic Olpidium brassicae chytridiomycete was shown to be a vector of the virus causing the lettuce big-vein disease; nevertheless, it was not yet proven at that time that it was a fungus that maintained reproduction of the virus [10, 11]. Penicillium chrysogenum virus (PcV) was one of the first mycoviruses that were studied in detail at the biochemical, biophysical, and ultrastructural levels. The host of this virus is a well-known producer of penicillin [12].
Fifty-five different mycoviruses had been identified by 1972 [13]. Today, mycoviruses are known to be widely spread in all major taxonomic groups of fungi [1, 14, 15]. Over 300 genomes of mycoviruses were sequenced and registered in the databases of the National Center for Biotechnology Information (NCBI). This number increases with every year since mycoviruses are widely used as a tool for biological control of the fungal pathogens, which infect economically significant plants and, what is especially important, for the treatment of invasive human mycoses [15, 16].
Symptomatology of the viral infection of fungi. Mycoviruses are most commonly latent in fungi and rarely cause visible symptoms [17]. This indicates an extremely good adaptation of mycoviruses to coexistence with their hosts during a long evolutionary period; such a biotic union is often beneficial for both mycoviruses and fungi [18, 19]. In contrast to the latent symptomless infection, some mycoviruses cause irregular growth, abnormal pigmentation, and the change in reproduction of their hosts [14, 17, 20, 21].
In some cases, viruses cause no symptoms in their hosts but can significantly alter their biology. Thus, the killer antagonism (the killer phenomenon of sensitive cells under the effect of species-specific toxins) is widely distributed among cultural (primarily, wine) Saccharomyces cerevisiae yeasts. The phenomenon of the killer activity in the yeast cells was first described in 1963 [22]. Some strains of S. cerevisiae suppressed the growth of sensitive strains of the same species. The antagonistic activity of the strains was subsequently shown to result from the protein-containing components, which were released by the killer yeasts into the environment and were toxic for the closely related cultures [23, 24]. Much later, this phenomenon was revealed to be associated with the simultaneous presence of double-stranded RNA of viruses (which resemble dsRNA of the viruses of mammals), their satellites (single-stranded RNA viruses of naked RNA), or yeast prions (virus-like DNA elements) [25]. To date, the killer strains have been proven to contain two cytoplasmically inherited viruses: ScV-M and ScV-L-A. The genes present in ScV-M code for the synthesis of the specific toxin (K1, K2, or K28). The stability and replication of ScV-M depends on the presence of the ScV-L-A virus [26]. Cytoplasmic viruses are present in the whole generation of the infected yeasts due to non-Mendelevian inheritance laws. Thus, the generation acquires the ability to kill sensitive virus-free cells, while the killer yeasts synthesizing the toxin are resistant to the latter. Hence, infected hosts possess a selective advantage in comparison with the uninfected ones [27].
Interestingly, the majority of the symptoms associated with mycoviral infections can be useful for their hosts and can often affect the interaction between fungi and other organisms. Thus, the presence of mycoviruses in endophytic fungi is associated with adaptation of plants to extreme conditions, including increased temperatures. For instance, thermal resistance of the grass Dichanthelium lanuginosum growing on the geothermal soil is caused by the presence of the mycovirus Curvularia thermal tolerance virus in the Curvularia protuberate endophytic fungus [28, 29]. Moreover, mycoviral infection may affect the adaptation of endophytes to plant hosts. Thus, entomopathogenic fungi, the life cycle of which includes the natural endophytic phase in the wild grass Festuca rubra and Holcus lanatus, were shown to acquire the ability to colonize other plants after being infected with mycovirus [30].
An important aspect of the effect of the viral infection on fungi infecting plants is a decrease in their virulence: hypovirulence. This characteristic was revealed in viruses of phytopathogenic fungi of the genera Cryphonectria (CHV1) and Fusarium graminearum (FgV1). FgV1 can be transmitted to Cryphonectria parasitica and other fungal species of the genus Fusarium and may cause a more pronounced hypovirulence in C. parasitica than CHV1 [31]. This issue will be considered in detail further.
Mycovirus transmission ways. Similarly to the viruses of animals and plants, mycoviruses require live cells of other organisms for replication. At the same time, apart from some common properties with these viruses, mycoviruses possess some unique characteristics as well: (1) extracellular viral particles are not able to infect fungi; (2) mycoviruses are transferred intercellularly only via cell differentiation, sporulation, and cell fusion; and (3) mycoviruses lack the transport protein vital for the life cycle of the viruses of animals and plants [32].
Mycoviral infections have a number of specific characteristics that differentiate them from the viruses of plants and animals. The most essential difference is that the life cycle of mycoviruses is not interrupted by the extracellular phase of transmission, i.e., mycoviruses are noninfectious in the classical understanding of this word. Thus, hyphae of the virus-free fungal organism cannot be infected when the hyphae are placed in the cell extract obtained from the infected strains. In contrast to the majority of viruses of plants and animals, natural vectors are unknown for mycoviruses [14, 33].
The transmission of mycoviruses is possible only between vegetatively compatible strains of the fungus via formation of the anastomosis of the hyphae or heterokaryosis (horizontal transfer). The viruses can also distribute with spores acquired in a sexual or an asexual way (vertical transfer) [14, 34]. Mycoviruses are suggested to have only an intracellular phase, and the spread of infection in the environment occurs only during sporulation when spores containing viruses are formed. Moreover, fungi have a rigid cell wall, and mycoviruses are considered unable to leave the cell. However, the recent data indicate that this suggestion is not absolutely correct, since some mycovirus particles can infect virus-free fungi. Thus, the researches isolated SsHADV-1 mycovirus and infected a virus-free host strain (Sclerotinia sclerotiorum) of the fungus that can infect and destroy a wide spectrum of agriculturally significant cultures [35]. The authors indicated that the distribution of the viruses in mycelial hyphae through the pores in septa occurs in a passive way: viral particles (similarly to cell organelles) can easily migrate between the neighbor cells.
According to the aforementioned findings, the transfer of the virus between different strains is limited to the vegetative incompatibility, which limits the use of hypovirulent mycoviruses as the agents of biological control. Recent studies showed that seven vic genes associated with five of six vic loci in Cryphonectria parasitica cause such incompatibility and affect the transmission of the virus [36].
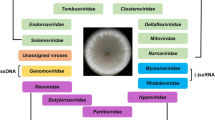
Genetics and morphology of mycoviruses. Viruses that infect fungi are primarily composed of isometric (icosahedral) or spherical particles with a diameter of 25–80 nm (in the case of Mycoreovirus (Reoviridae)) and have a segmented double-stranded RNA (dsRNA) or linear single-stranded RNA (ss(+)RNA) genome and are rarely packed into the capsid [1, 17, 37]. Unclassified mycoviruses with linear (–)RNA and viruses with circular ssDNA are differentiated as well [20, 38–40]. Simple mycoviruses with the unencapsidated naked dsRNA genome are known only in the members of the Endornaviridae and Narnaviridae families [17, 41, 42]. Unencapsidated genomes are in pleomorphic vesicles and, as a rule, are rarely observed among mycoviruses (they are known in the Hypoviridae family only) [17] (Table 1).
Kanhayuwa et al. [43] revealed a unique mycovirus in 2015. This virus infects Aspergillus fumigatus, which is one of the causative agents of human aspergillosis, the disease primarily causing mortality in the patients with a low immune status. The mycovirus named Aspergillus fumigatus tetramycovirus–1 (AfuTmV–1) contains four dsRNA segments (dsRNA 1 (~2.4 kbp), dsRNA 2 (~2.2 kbp), dsRNA 3 (~1.9 kbp), and dsRNA 4 (~1.1 kbp)) and has a unique organization. In contrast to other viruses, its genome is unencapsidated but has a coat consisting of virus-encoded proteins. The authors showed that both the purified AfuTmV-1 and dsRNA infect protoplasts of fungi. The scientists suggested that this will enable one to change the genome of the virus by the methods of gene engineering and to develop the silencing vectors as well as to use it as a tool of switching off the genes of fungi, which may result in the efficient struggle with human aspergilloses.
The viruses with the single-stranded linear (+)RNA genome are classified into seven families: Alphaflexiviridae, Barnaviride, Gammaflexiviridae, Endornaviridae, Hypoviridae, Narnaviridae, and the recently proposed family Fusariviridae [17, 20, 44, 45].
The simplest genomes are considered (+)RNA genomes of mycoviruses of the family Narnaviridae. The genome (2–3 kb in size) encodes RNA-dependent RNA polymerase (RdRp) necessary for replication. Narnaviruses lack the coat protein (CP), which forms the capsid structure, as well as the transport protein (TP) [46]. Phylogenetically close relatives of narnaviruses are viruses of plants of the genus Ourmiavirus. The genome of the plant Ourmiavirus is represented by a three-segment ssRNA encoding three proteins: RdRp, CP, and TP [47]. Phylogenetic analysis indicates that the genus Ourmiavirus occupies an intermediate position between mycoviruses and viruses of plants [48].
Mycoviruses with the dsRNA genome are widely spread in yeasts and mycelial fungi and, in contrast to the viruses of mammals, lack the extracellular stage and have no pathological effect on their hosts [1, 33], with some exceptions [49]. Double-stranded (ds)RNA of mycoviruses are classified into seven families: Crysoviridae, Partitiviridae, Totiviridae, Reoviridae, Megabirnaviridae, Endornaviridae, and Quadriviridae.
The viruses with dsRNA such as viruses encoding killer toxins in the cells of the Ustilago maydis and Saccharomyces cerevisiae [50–53] are well examined. These viruses belong to the family Totiviridae. The members of this family are characterized by the presence of nonsegmented dsRNA genome and isometric virions (Fig. 1), which are usually called virus-like particles (VLP), since they are not infectious [25].
Morphology of the RnMBV1 particles isolated from the mycelium of Rosellinia necatrix strain W779 [54].
In 2014, Liu et al. [38] proved the existence of mycoviruses with the ssRNA genome of negative polarity (ss(–)RNA). The virus named Sclerotinia sclerotiorum negative-stranded RNA virus 1 (SsNSRV-1) was isolated from the hypovirulent Sclerotinia sclerotiorum strain. Purified preparations of the filamentary virion-like structures with a diameter of 22 nm and the length of 200–2000 nm were obtained (Fig. 2). The virus-free S. sclerotiorum strain was infected with the purified virus and became hypovirulent after infection. The revealed virus belongs to the family Mycomononegaviridae and possesses some unique characteristics in the morphology and specific composition of the genome.
Morphology of the SsNSRV-1 particles isolated from the mycelium of Sclerotinia sclerotiorum [38]. (a) Filamentous, probably encapsidated virions (indicated with arrows) and ribonucleoprotein complexes (RNP); (b) purified densely packed or loose RNP.
It is noteworthy that the well-known viruses dangerous for humans (Ebola virus, respiratory syncytial virus, measles, Nipah virus, and rabies virus) also belong to mononegaviruses. Many mononegaviruses are found in animals, and only some of them infect plants and invertebrates (nematodes). The scientists suggest that the discovery of the SsNSRV-1 mycovirus can provide insights into the global ecology and evolution of viruses, while the S. sclerotiorum–SsNSRV-1 system may become the basis for the study of the fungal factors participating in replication and the life cycle of the (–)RNA viruses. Moreover, this system may become a perfect model for screening the antiviral compounds against dangerous human and animal (–)RNA viruses.
To date, only one mycovirus with single-stranded circular DNA has been described in the literature. The first mycovirus with the ssDNA genome was isolated from the hypovirulent strain DT-8 of the S. sclerotiorum fungus [55] and was named Sclerotinia sclerotiorum hypovirulence-associated DNA virus 1 (SsHADV-1). Phylogenetic analysis showed that SsHADV-1 was related to the viruses of the family Geminiviridae. Therefore, the authors supposed that the mycovirus was closely related to plant geminiviruses but differed from them in the morphology of the viral particles and genome organization (Fig. 3). In 2016, the International Committee on Taxonomy of Viruses (ICTV) approved a novel Genomoviridae family for classification of the mycoviruses with ssDNA [56].
Viral particles of SsHADV-1 isolated from the mycelium of Sclerotinia sclerotiorum strain DT-8 [55].
In contrast to geminiviruses, the genomic ssDNA and purified virions retained their infectiousness and could be easily transfered from the DT-8 strain to other strains from vegetatively compatible group. This fact is evidence of the existence of other ways of virus transmission, which are different from the earlier known ones in mycoviruses with RNA genome or geminiviruses. These data suggest that the host range of such mycoviruses is not limited only to S. sclerotiorum. This fact stimulates the search for new infectious DNA mycoviruses in other important phytopathogenic fungi. The study of interactions between SsHADV-1 and S. sclerotiorum is important for understanding the mechanisms of interaction between DNA viruses and fungi as well as for the development and probable application for biological control of fungal diseases of important farm cultures.
At the moment of writing the present paper, the literature lacked information about the existence of mycoviruses with ssDNA and fungal infections caused by them. At the same time, such a virus was isolated and described in hyphochytriomycetes [57], which belonged to aqueous fungi earlier and were subsequently assigned to the kingdom of protists after the development of molecular methods of research used for classification of organisms.
Fungal antiviral defense mechanisms. In the course of evolution, animals and plants have developed complicated mechanisms for defense against viral infection. These mechanisms involve innate and adaptive responses, which include RNA silencing, interferon production, and antibody production. Antiviral defense mechanisms currently identified in fungi include a self/nonself recognition system that represents a barrier to the major mode of mycovirus transmission and an RNA recognition system that targets mycovirus RNA for destruction.
A self/nonself recognition system (vegetative incompatibility) is the ability of fungi to form anastomoses between hyphae of different strains of the certain species. Anastomoses are special cell connections in which cytoplasmic mixing of vegetatively compatible strains occurs, which results in heterokaryon formation. On the one hand, the ability of vegetatively compatible strains to form anastomoses is one of the major ways of their horizontal transfer for distribution of mycoviruses. At the same time, on the other hand, evolutionally developed genetic control of vegetative incompatibility is a strong barrier to their spread, which was mentioned above for the vic genes in Cryphonectria parasitica [36].
Gene silencing is known to be the most conservative mechanism for defense of cellular RNA from the foreign information in the form of nucleic acids of viruses, transposons, or transgenes. RNA interference (RNAi) is one of the mechanisms for realization of silencing and a fundamental mechanism based on the recognition of dsRNA of exogenous or endogenous origin by cellular proteins (Dicer enzyme) and its cleavage into short (21–26 nucleotides) fragments known as small interfering RNA and microRNA [58]. The components essential for RNAi (Dicer, Argonaute, and RdRp) were identified in all kingdoms of the domain (superkingdom) of eukaryotes [59], which is evident of its important contribution to the functioning of the defense and regulatory mechanisms in their common ancestor.
Until recently, the issue on the significance of the RNA silencing in the defense mechanisms of fungi against invasive nucleic acids and viruses has been open. Lately, virus-induced RNA silencing in fungi named quelling was studied in detail for Cryphonectria parasitica and Aspergillus nidulans [60]. Thus, the genes dcl2 (Dicer protein) and agl2 (Argonaute protein), which are essential for induction of the RNA silencing as a mechanism of antiviral defense in C. parasitica, were shown to be induced in response to infection with Cryphonectria hypovirus 1 (CHV1) [61]. Considerable changes in the expression level of some genes associated with RNA silencing (rdr1, dcl1, dcl2, and agl2) were also observed in Fusarium gramineraum infected with mycoviruses [62]. Defects in the genes responsible for RNAi considerably increased sensitivity of fungi to viral infections [63]. Mycoviruses, similarly to other viruses of plants and animals, produce suppressors of RNA silencing in order to block such antiviral defense. For example, the p29 protein of the mycovirus CHV1 and the S10 gene product of Rosellinia necatrix mycoreovirus are able to inhibit the RNAi mechanism [64].
Thus, the results obtained in some experimental studies indicate the important role of RNA silencing in the fungal defense against mycovirus infection. The absence of the components essential for realization of RNAi (e.g., in Candida albicans and Ustilago maydis) indicate the probable existence of different unknown mechanisms.
The use of experimental systems involving fungi and mycoviruses for the study of the mechanisms underlying induction and suppression of the antiviral RNA silencing are promising due to simpler mechanisms of RNAi and evolutionary position of fungi in relation to plants and animals.
Hypovirulence as a unique property of mycoviruses. Biotechnological application of mycoviruses is associated with their ability to considerably decrease the virulence of pathogenic fungi. This phenomenon was named hypovirulence.
The connection between hypovirulence of fungal strains and viral infection was first established in the early 1950s [65]. The chestnuts infected with Cryphonectria parasitica were shown to survive, and the lesions on the stems healed without any external impact. C. parasitica strains isolated from the necrotic segments were poorly pigmented, in contrast to the normal bright-orange strains. Although they infected European chestnuts, they rarely caused a lethal infection; i.e., they possessed a decreased spore-forming ability and insufficient synthesis of laccases. Double-stranded RNA containing two open reading frames (RNA-dependent RNA polymerase and ATP-dependent helicase) were subsequently extracted from these isolates. This resulted in the discovery of two hypoviruses (HAV1 and CHV1-713) and later in the application of hypovirulent strains for biological control of the outbreak of cryphonecrosis of chestnuts, which occurred in the United States and Europe in the beginning of the 20th century and is considered the largest botanical catastrophe ever observed [66].
The biological program with the use of hypovirulent strains of the causative agent of this disease in Europe was first applied in southern France in 1967–1972 and provided for surprisingly positive results. This was the first significant success of the attempts to control the disease [67, 68]. Despite the skeptical attitude of some researchers [69], to date, the use of hypovirulence is still the best method for biological control of the chestnut infections caused by C. parasitica [70–75].
One of the problems of the application of this method is an unpredictable result of the use of hypovirulence in the ecosystem, since the success or failure of the method depends on the interaction between the hypovirus, fungal pathogen, plants, and environmental factors. Despite these difficulties, application of mycoviruses to control cryphonecrosis in the gardens of eastern North America and Europe remains a common practice for over 40 years [14].
Recently, intense research into the application of hypovirulent strains in order to control fungal infections under field conditions is being carried out. Pathosystems in agroecosystems are significantly different from those in forests and gardens, where perennial trees are planted. Due to the high density of the planted cultures and small species diversity, the conditions favorable for growth, reproduction, and pathogen transmission are developed in the sown fields. These conditions facilitate the spread of mycoviruses in sowings.
The use of mycoviruses associated with hypovirulence has some advantages. First of all, this is a fast effect of the mycovirus infection: the viruses are transferred to the virulent fungal strains and rapidly inhibit the growth of fungal lesions. Taking into account the fact that fungal diseases often damage or kill plants during the vegetation period, this property may play a crucial role for successful biological control [76]. Moreover, hypovirulent strains produce pathogen-associated molecular patterns and/or effectors, which are recognized by their hosts. In this case, a defense response is induced in plants, which is specifically directed against the virulent strain [15]. Thus, the DNA virus (SsHADV-1) associated with the fungal hypovirulence is used to struggle the rot of colza caused by Sclerotinia sclerotiorum. This hypovirulent strain suppresses the disease both in the form of suspension of the hyphal fragments infected with the virus or viral particles [15, 21]. It is noteworthy that SsHADV-1 possesses pronounced infectivity, and its horizontal transfer is independent of vegetative compatibility of different strains of the pathogen during the formation of anastomoses between their hyphae. Purified SsHADV-1 viral particles may infect virus-free hyphae of S. sclerotiorum and, therefore, suppress the development of the fungal infection and reduce the pathological effect of the causative agent. According to the authors’ opinion [35], mycoviruses capable of extracellular distribution may be used as natural fungicides.
Rosellinia necatrix ascomycete is a phytopathogenic fungus causing white root blight of the apple tree, vine, pear, plum, poplar, and walnut [77]. Fungal infection may result in wilting, chlorosis, and defoliation as well as fast death of infected trees under the certain environmental conditions. Probable prospects of virocontrol (biological control using viruses) resulted in the necessity to study mycoviruses infecting R. necatrix [1]. In 2001, dsRNA of different types was revealed in 65 of 298 isolates of R. necatrix [78]. Molecular characteristic of these viruses showed that R. necatrix is a natural host of mycoviruses of at least five families, including Partitiviridae [79, 80], Quadriviridae [81, 82], Reoviridae [83], Totiviridae [64], and Megabirnaviridae [54]. The virus Rosellinia necatrix megabirnavirus 1 (RnMBV1) (the family Megabirnaviridae), which causes a pronounced decrease in both the mycelium growth of R. necatrix and its virulence, independently of the pathogen strain, can be promising for the biological control of the white root blight infection [84].
Botrytis cinerea is also a phytopatogenic fungus infecting vegetables, ornamental plants, and economically significant cultures, such as grapes, strawberry, raspberry, kiwi, and pears [85]. The disease associated with the infection by B. cinerea is gray mold, which causes vast economical losses worldwide. The struggle against B. cinerea often turns out to be unsuccessful due to the emergence of the isolates resistant to fungicides and to different levels of pathogen virulence caused by its genetic variability and the presence of mycoviral infection. The ability to use mycoviruses for biocontrol and as tools for the investigation of the interactions between plants and a pathogen promoted their study in B. cinerea [86]. Similarly to other phytopathogenic fungi, mycoviruses are widely spread in the population of fungi of the genus Botrytis [87–89], although the main barrier to their application as agents of biological control in the struggle with the fungal diseases of plants is vegetative incompatibility of the strains of this species. This characteristic limits the transfer of the mycoviruses to virus-free strains via hyphal anastomoses.
Recently, a new RNA-bearing mycovirus was revealed in the hypovirulent strain BerBc-1. It can be transferred to other strains of B. cinerea vertically via macroconidia or horizontally through hyphal contact [90]. The presence of the mycovirus Botrytis cinerea RNA virus 1 (BcRV1) proved to correlate positively with the hypovirulence of B. cinerea. At the same time, the growth of mycelium and pathogenicity of the fungus decreased significantly with accumulation of BcRV1. Thus, the existence of mycoviruses, which are able to overcome vegetative incompatibility during transmission, is a prerequisite for the biological control of B. cinerea.
The fungi of the genus Fusarium are phytopathogens widely spread all over the world. Some of them are not only responsible for crop loss and a decrease in quality of the grain cultures but also produce mycotoxins in the grain cultures that are able to affect the human and animal health. Nowadays, mycoviruses have been revealed in F. graminearum, F. poae, F. solani, F. oxysporum, F. boothii, and F. virguliforme [91, 92]. Despite the fact that eleven mycoviruses were revealed in the fungi of the genus Fusarium, only one of them (in the maize phytopathogen F. graminearum 1-DK21 (FgV1)) induced hypovirulence in the host as well as a decrease in the level of biosynthesis of trichothecene mycotoxin [91, 93, 94]. At the same time, it should be noted that absolutely all strains of F. poae capable of synthesizing trichothecene mycotoxin contained mycoviruses with dsRNA in the mycelium cells [95].
Thus, many examples of the successful application of mycoviruses in the struggle against fungal diseases of plants exist. In each of the cases, the result was determined by the vegetative compatibility of the fungal isolates and characteristics of hypoviruses [36]. Therefore, first of all, the problem of the vegetative incompatibility should be solved and efficient transfer of the mycovirus from a hypovirulent strain to the target one should be provided, when mycoviruses are used as the agents of biotechnological control [32]. The following strategies can be used to achieve this goal: screening of mycoviruses with strong infectivity, the search for the universal donor of mycoviruses (developed under both the conditions of the laboratory and isolated from the natural environment), and a decrease in the response to vegetative incompatibility of the host via affecting with the chemical compounds. The transfer of the mycoviruses by the protoplast fusion can be promising in this field [96]. This approach was successfully used in the case of phytopathogenic fungi, including fungi of the genera Aspergillus [97], Fusarium [96], Trichoderma [98], and Rosellinia [99].
Since mycoviruses do not affect their hosts, they can be used as gene vectors. The suitable candidates may be ssRNA mycoviruses of the family Flexiviridae: Botrytis virus X (BVX), Botrytis cinerea virus F (BCVF), and Sclerotinia sclerotiorum debilitation-associated RNA virus (SsDRV). The use of mycoviruses for expression of the antigenic epitopes of humans and animals for vaccine development (similarly to the capsid proteins of some viruses of plants) should not be excluded. Moreover, mycoviruses are promising tools for fundamental research as well as for diverse bio- and nanotechnologies [100].
REFERENCES
Ghabrial, S.A. and Suzuki, N., Viruses of plant pathogenic fungi, Annu. Rev. Phytopathol., 2009, vol. 47, pp. 353–384. doi 10.1146/annurev-phyto-080508-081932
Wiebols, G.L.W. and Wieringa, K.T., Bacteriophagie een algemeen voorkomend verschijnsel, Fonds Landbouw Export Bureau, 1936, no. 16, pp. 1916–1918.
Sinden, J.W. and Hauser, E., Report on two new mushroom diseases, Mushroom Sci., 1950, vol. 1, pp. 96–100.
Hollings, M., Viruses associated with a die-back disease of cultivated mushroom, Nature, 1962, vol. 196, pp. 962–965.
Romaine, C.P. and Schlagnhaufer, B., PCR Analysis of the viral complex associated with La France disease of Agaricus bisporus, Appl. Envirom. Microbiol., 1995, vol. 61, no. 6, pp. 2322–5.
Revill, P.A. and Wright, P.J., RT-PCR detection of dsRNAs associated with La France disease of the cultivated mushroom Agaricus bisporus (Lange) Imbach, J. Virol. Methods, 1997, vol. 63, nos. 1–2, pp. 17–26.
Borodynko, N., Hasiyw-Jaroszewska, B., Rymelska, N., and Pospieszny, H., La France disease of the cultivated mushroom Agaricus bisporus in Poland, Acta Virol., 2010, vol. 54, no. 3, pp. 217–219.
Magae, Y. and Sunagawa, M., Characterization of a mycovirus associated with the brown discoloration of edible mushroom, Flammulina velutipes, Virol. J., 2010, vol. 7, p. 342.
Magae, Y., Molecular characterization of a novel mycovirus in the cultivated mushroom, Lentinula edodes, Virol. J., 2012, vol. 6, no. 9, p. 60. doi 10.1186/ 1743-422X-9-60
Grogan, R.G. and Campbell, R.N., Fungi as vectors and hosts of viruses, Ann. Rev. Phytopathol., 1966, vol. 4, pp. 29–52. doi.org/10.1146/annurev.py. 04.090166.000333
Hollings, M. and Stone, O.M., Viruses in fungi, Sci. Progr., 1969, vol. 57, no. 227, p. 371.
Jiang, D. and Ghabrial, S.A., Molecular characterization of Penicillium chrysogenum virus: reconsideration of the taxonomy of the genus Chrysovirus, J. Gen. Virol., 2004, vol. 85, pp. 2111–2121. doi 10.1099/vir.0.79842-0
Bozarth, R.F., Mycoviruses: a new dimension in microbiology, Environ. Health Perspect., 1972, vol. 2, pp. 23–39.
Nuss, D.L., Hypovirulence: mycoviruses at the fungal–plant interface, Nat. Rev. Microbiol., 2005, vol. 3, pp. 632–642. doi 10.1038/nrmicro1206
Xie, J. and Jiang, D., New insights into mycoviruses and exploration for the biological control of crop fungal diseases, Annu. Rev. Phytopathol., 2014, vol. 52, pp. 45–68. doi 10.1146/annurev-phyto-102313-050222
van de Sande, W.W.J., Lo-Ten-Foe, J.R., van Belkum, A., Netea, M.G., Kullberg, B.J., and Vonk, A.G., Mycoviruses: future therapeutic agents of invasive fungal infections in humans?, Eur. J. Clin. Microbiol. Infect. Dis., 2010, vol. 29, no. 7, pp. 755–763. doi 10.1007/s10096-010-0946-7
Pearson, M.N., Beever, R.E., Boine, B., and Arthur, K., Mycoviruses of filamentous fungi and their relevance to plant pathology, Mol. Plant Pathol., 2009, vol. 10, no. 1, pp. 115–128. doi 10.1111/j.1364-3703.2008.00503.x
Roossinck, M.J., The good viruses: viral mutualistic symbioses, Nat. Rev. Microbiol., 2011, vol. 9, no. 2, pp. 99–108. doi 10.1038/nrmicro2491
Márquez, L.M. and Roossinck, M.J., Do persistent RNA viruses fit the trade-off hypothesis of virulence evolution?, Curr. Opin. Virol., 2012, vol. 2, no. 5, pp. 556–60.
Ghabrial, S.A., Castyn, J.R., Jiang, D., Nibert, M.L., and Suzuki, N., 50-Plus years of fungal viruses, Virology, 2015, vols. 479–480, pp. 356–368. doi 10.1016/ j.virol.2015.02.034
Jiang, D., Fu, Y., Guoqing, L., and Ghabrial, S.A., Mycoviruses: viruses of the plant pathogenic fungus Sclerotinia sclerotiorum, Adv. Virus Res., 2013, vol. 86, pp. 215–248. doi 10.1016/B978-0-12-394315-6.00008-8
Woods, D.R., Ross, I.W., and Hendry, D.A., A new killer factor produced by a killer/sensitive yeast strain, J. Gen. Microbiol., 1974, vol. 81, pp. 285–289.
Woods, D.R. and Bevan, E.A., Studies on the nature of the killer factor produced by Saccharomyces cerevisiae, J. Gen. Microbiol., 1968, vol. 51, pp.115–126.
Bussey, H., Physiology of the killer factor in yeast, Adv. Microbial. Physiol., 1981, vol. 22, pp. 93–122.
Wickner, R.B., Fujimura, T., and Esteban, R., Viruses and prion of Saccharomyces cerevisiae, Adv. Virus Res., 2013, vol. 86, pp. 1–36. doi 10.1016/B978-0-12-394315-6.00001-5
Marquina, D., Santos, A., and Peinado, J., Biology of killer yeasts, Int. Microbiol., 2002, vol. 5, no. 2, pp. 65–71. doi 10.1007/s10123-002-0066-z
Wloch-Salamon, D.M., Sociobiology of the budding yeast, J. Biosci., 2014, vol. 39, no. 2, pp. 225–236.
Márquez, L.M., Redman, R.S., Rodriguez, R.J., and Roossinck, M.J., A virus in a fungus in a plant: three-way symbiosis required for thermal tolerance, Science, 2007, vol. 315, no. 5811, pp. 513–515. doi 10.1126/science.1136237
Herrero, N., Marquez, S.S., and Zabalgogeazcoa, I., Mycoviruses are common among different species of endophytic fungi of grasses, Arch. Virol., 2009, vol. 154, pp. 327–330. doi 10.1007/s00705-008-0293-5
Asencio, N.H., Máquez, S.S., and Zabalgogeazcoa, I., Mycovirus effect on the endophytic establishment of the entomopathogenic fungus Tolypocladium cylindrosporum in tomato and bean plants, BioControl., 2013, vol. 58, no. 2, pp. 225–232.
Lee, K-M., Yu, J., Son, M., Lee, Y.-W., and Kim, K-H., Transmission of Fusarium boothii mycovirus via protoplast fusion causes hypovirulence in other phytopathogenic fungi, PLoS One, 2011, vol. 6, no. 6. e21629. doi 10.1371/journal.pone.0021629
Son, M., Yu, J., and Kim, K.-H., Five questions about mycoviruses, PLoS Pathog., 2015, vol. 11, no. 11. e1005172. doi.org/10.1371/journal.ppat.1005172
Ghabrial, S.A., Origin, adaptation and evolutionary pathways of fungal viruses, Virus Genes, 1998, vol. 16, no. 1, pp. 119–131.
Xie, J., Wei, D., Jiang, D., Fu, Y., Li, G., Ghabrial, S., and Peng, Y., Characterization of debilitation associated mycovirus infecting the plant-pathogenic fungus Sclerotinia sclerotiorum, J. Gen. Virol., 2006, vol. 87, no. 1, pp. 241–249. doi 10.1099/vir.0.81522-0
Yu, X., Li, B., Fu, Y., Xie, J., Cheng, J., Ghabrial, S.A., Li, G., Yi, X., and Jiang, D., Extracellular transmission of a DNA mycovirus and its use as a natural fungicide, Proc. Natl. Acad. Sci. U. S. A., 2013, vol. 110, no. 4, pp. 1452–1457. doi 10.1073/pnas.1213755110
Choi, G.H., Dawe, A.L., Churbanov, A., Smith, M.L., Milgroom, M.G., and Nuss, D.L., Molecular characterization of vegetative incompatibility genes that restrict hypovirus transmission in the chestnut blight fungus Cryphonectria parasitica, Genetics, 2012, vol. 190, no. 1, pp. 113–127. doi 10.1534/genetics.111.133983
Fauquet, C.M., Mayo, M., Maniloff, M.A., Desselberger, U., and Ball, L.A., Virus Taxonomy: VIIIth Report of the International Committee on Taxonomy of Viruses, London: Academic Press, 2005.
Liu, L., Xie, J., Cheng, J., Fu, Y., Li, G., Yi, X., and Jiang, D., Fungal negative-stranded RNA virus that is related to bornaviruses and nyaviruses, Proc. Natl. Acad. Sci. U. S. A., 2014, vol. 111, no. 33, pp. 12205–12210. doi 10.1073/pnas.1401786111
Marzano, S.-Y.L. and Domier, L.L., Novel mycoviruses discovered from metatranscriptomics survey of soybean phyllosphere phytobiomes, Virus Res., 2016, vol. 213, pp. 332–342. doi 10.1016/j.virusres.2015.11.002
Marzano, S.-Y.L., Nelson, B.D., Ajayi-Oyetunde, O., Bradley, C.A., Hughes, T.J., Hartman, G.L., Eastburn, D.M., and Domier, L.L., Identification of diverse mycoviruses through metatranscriptomics characterization of the viromes of five major fungal plant pathogens, J. Virol., 2016, vol. 90, no. 15, pp. 6846–6863. doi 10.1128/JVI.00357-16
Horiuchi, H. and Fukuhara, T., Putative replication intermediates in Endornavirus, a novel genus of plant dsRNA viruses, Virus Genes, 2004, vol. 29, no. 3, pp. 365–375. doi 10.1007/s11262-004-7441-0
Cole, T.E., Hong, Y., Brasier, C.M., and Buck, K.W., Detection of an RNA-dependent RNA polymerase in mitochondria from a mitovirus-infected isolate of the Dutch elm disease fungus, Ophiostoma novo-ulmi, Virology, 2000, vol. 268, no. 2, pp. 239–243. doi.org/ 10.1006/viro.1999.0097
Kanhayuwa, L., Kotta-Loizou, I., Özkan, S., Gunning, A.P., and Coutts, R.H.A., A novel mycovirus from Aspergillus fumigatus contains four unique dsRNAs as its genome and is infectious as dsRNA, Proc. Natl. Acad. Sci. U. S. A., 2015, vol. 112, no. 29, pp. 9100–9105. doi 10.1073/pnas.1419225112
Zhang, R., Liu, S., Chiba, S., Kondo, H., Kanematsu, S., and Suzuki, N., A novel single-stranded RNA virus isolated from a phytopathogenic filamentous fungus, Rosellinia necatrix, with similarity to hypolike viruses, Front. Microbiol., 2014, vol. 5, p. 360. doi 10.3389/fmicb.2014.00360
Wang, L., Zhang, J., Zhang, H., Qiu, D., and Guo, L., Two novel relative double-stranded RNA mycoviruses infecting Fusarium poae strain SX63, Int. J. Mol. Sci., 2016, vol.17, no. 5, p. 641. doi 10.3390/ijms17050641
Hillman, B.I. and Cai, G., The family Narnaviridae: simplest of RNA viruses, Adv. Virus Res., 2013, vol. 86, pp. 149–176. doi 10.1016/B978-0-12-394315-6.00006-4
Rastgou, M., Habibi, M.K., Izadpanah, K., Masenga, V., Milne, R.G., Wolf, Y.I., Koonin, E.V., and Turina, M., Molecular characterization of the plant virus genus Ourmiavirus and evidence of interkingdom reassortment of viral genome segments as its possible route of origin, J. Gen. Virol., 2009, vol. 90, no. 10, pp. 2525–2535. doi 10.1099/vir.0.013086-0
Donaire, L., Rozas, J., and Ayllon, M.A., Molecular characterization of Botrytis ourmia-like virus, a mycovirus close to the plant pathogenic genus Ourmiavirus, Virology, 2016, vol. 489, pp. 158–164. doi 10.1016/ j.virol.2015.11.027
Bhatti, M.F., Jamal, A., Petrou, M.A., Cairns, T.C., Bignell, E.M., and Coutts, R.H., The effects of dsRNA mycoviruses on growth and murine virulence of Aspergillus fumigatus, Fungal. Genet. Biol., 2011, vol. 48, no. 11, pp. 1071–1075. doi 10.1016/j.fgb.2011.07.008
Bruenn, J., The double-stranded RNA viruses of Ustilago maydis and their killer toxins, in dsRNA Genetic Elements, Boca Raton, FL: CRC Press, 2001, pp. 109–124. doi 10.1201/9781420039122.ch4
Kang, J., Wu, J., Bruenn, J.A., and Park, C., The H1 double-stranded RNA genome of Ustilago maydis virus-H1 encodes a polyprotein that contains structural motifs for capsid polypeptide, papain-like protease, and RNA-dependent RNA polymerase, Virus Res., 2001, vol. 76, no. 2, pp. 183–189.
Schmitt, M.J. and Breinig, F., Yeast viral killer toxins: lethality and self-protection, Nat. Rev. Microbiol., 2006, vol. 4, pp. 212–221. doi 10.1038/nrmicro1347
Tipper, D.J. and Schmitt, M.J., Yeast dsRNA viruses: replication and killer phenotypes, Mol. Microbiol., 1991, vol. 5, no. 10, pp. 2331–2338.
Chiba, S., Salaipeth, L., Lin, Y.H., Sasaki, A., Kanematsu, S., and Suzuki, N., A novel bipartite double-stranded RNA mycovirus from the white root rot fungus Rosellinia necatrix: molecular and biological characterization, taxonomic considerations, and potential for biological control., J. Virol., 2009, vol. 83, no. 24, pp. 12801–12812. doi 10.1128/JVI.01830-09
Yu, X., Li, B., Fu, Y., Jiang, D., Ghabrial, S.A., Li, G., Peng, Y., Xie, J., Cheng, J., Huang, J., and Yi, X., A geminivirus-related DNA mycovirus that confers hypovirulence to a plant pathogenic fungus, Proc. Natl. Acad. Sci. U. S. A., 2010, vol. 107, no. 18, pp. 8387–8392. doi 10.1073/pnas.0913535107
Krupovic, M., Ghabrial, S.A., Jiang, D., and Varsani, A., Genomoviridae: a new family of widespread single-stranded DNA viruses, Arch. Virol., 2016, vol. 161, no. 9, pp. 2633–2643. doi 10.1007/s00705-016-2943-3
Dawe, V.H. and Kuhn, C.W., Isolation and characterization of a double-stranded DNA mycovirus infecting the aquatic fungus, Rhizidiomyces, Virology, 1983, vol. 130, no. 1, pp. 21–28. doi 10.1016/0042-6822(83)90114-9
Meng, H., Wang, Z., Wang, Y., Zhu, H., and Huang, B., Dicer and Argonaute genes involved in RNA interference in the entomopathogenic fungus Metarhizium robertsii, Appl. Environ. Microbiol., 2017, vol. 83, no. 7. e03230-16. doi 10.1128/AEM.03230-16
Shabalina, S.A. and Koonin, E.V., Origins and evolution of eukaryotic RNA interference, Trends Ecol. Evol., 2008, vol. 23, no. 10, pp. 578–587. doi 10.1016/j.tree.2008.06.005
Chang, S.S., Zhang, Z., and Liu, Y., RNA interference pathways in fungi: mechanisms and functions, Annu. Rev. Microbiol., 2012, vol. 66, pp. 305–323. doi 10.1146/annurev-micro-092611-150138
Zhnag, D.X., Spiering, M.J., and Nuss, D.L., Characterizing the roles of Cryphonectria parasitica RNA-dependent RNA polymerase-like genes in antiviral defense, viral recombination and transposon transcript accumulation, PLoS One, 2014, vol. 9, no. 9. e108653. doi 10.1371/journal.pone.0108653
Lee, K.-M., Cho, W.K., Yu, J., Son, M., Choi, H., Min, K., Lee, Y.W., and Kim, K.H., A Comparison of transcriptional patterns and mycological phenotypes following infection of Fusarium graminearum by four mycoviruses, PLoS One, 2014, vol. 9, no. 6. e100989. doi 10.1371/journal.pone.0100989
Segers, G.C., Zhang, X., Deng, F., Sun, Q., and Nuss, D.L., Evidence that RNA silencing functions as an antiviral defense mechanism in fungi, Proc. Natl. Acad. Sci. U. S. A., 2007, vol. 104, no. 31, pp. 12902–12906. doi 10.1073/pnas.0702500104
Yaegashi, H., Yoshikawa, N., Ito, T., and Kanematsu, S., A mycoreovirus suppresses RNA silencing in the white root rot fungus, Rosellinia necatrix, Virology, 2013, vol. 444, nos. 1–2, pp. 409–416. doi 10.1016/j.virol.2013.07.010
Biraghi, A., Ulteriori notizie sulla resistenza di Castallea sativa Mill. nei confronti di Endothia parasitica (Murr.) And., Boll. Staz. Patol. Veg., 1954, vol. 9, pp. 149–157.
Anagnostakis, S.L., Biological control of chestnut blight, Science, 1982, vol. 215, no. 4532, pp. 466–471. doi 10.1126/science.215.4532.466
Turchetti, T., Hypovirulence in chestnut blight (Endothia parasitica [Murr.] And.) and some practical aspects in Italy, Eur. J. For. Pathol., 1982, vol. 12, pp. 414–416. doi 10.1111/j.1439-0329.1982.tb01296.x
Rigling, D. and Prospero, S., Cryphonectria parasitica, the causal agent of chestnut blight: invasion history, population biology and disease control, Mol. Plant Pathol., 2018, vol. 19, no. 1, pp. 7–20. doi 10.1111/mpp.12542
Milgroom, M.G. and Cortesi, P., Biological control of chestnut blight with hypovirulence: a critical analysis, Annu. Rev. Phytopathol., 2004, vol. 42, pp. 311–338. doi 10.1146/annurev.phyto.42.040803.140325
Heiniger, U. and Rigling, D., Biological control of chestnut blight in Europe, Annu. Rev. Phytopathol., 1994, vol. 32, pp. 581–599.
Robin, C., Lanz, S., Soutrenon, A., and Rigling, D., Dominance of natural over released biological control agents of the chestnut blight fungus Cryphonectria parasitica in southeastern France is associated with fitness-related traits, Biol. Control, 2010, vol. 53, no. 1, pp. 55–61.
Robin, C., Anziani, C., and Cortesi, P., Relationship between biological control., incidence of hypovirulence, and diversity of vegetative compatibility types of Cryphonectria parasitica in France, Phytopathology, 2000, vol. 90, no. 7, pp. 730–737. doi 10.1094/ PHYTO.2000.90.7.730
Juhásová, G. and Bernadovicová, S., Cryphonectria parasitica (Murr.) Barr and Phytophthora spp. in chestnut (Castanea sativa Mill.) in Slovakia, For. Snow Landsc. Res., 2001, vol. 76, no. 3, pp. 373–377.
Hoegger, P.J., Heiniger, U., Holdenrieder, O., and Rigling, D., Differential transfer and dissemination of hypovirus and nuclear mitochondrial genomes of a hypovirus-infected Cryphonectria parasitica strain after introduction into a natural population, Appl. Environ. Microbiol., 2003, vol. 69, no. 7, pp. 3767–3771. doi 10.1128/AEM.69.7.3767-3771.2003
Krstin, L., Katanić, Z., Ježić, M., Poljak, I., Nuskern, L., Matković, I., Idžojtić, M., and Ćurković-Perica, M., Biological control of chestnut blight in Croatia: an interaction between host sweet chestnut, its pathogen Cryphonectria parasitica and the biocontrol agent Cryphonectria hypovirus 1, Pest Manag. Sci., 2017, vol. 73, no. 3, pp. 582–589. doi 10.1002/ps.4335
Boine, B., Kingston, R.L., and Pearson, M.N., Recombinant expression of the coat protein of Botrytis virus X and development of an immunofluorescence detection method to study its intracellular distribution in Botrytis cinerea, J. Gen. Virol., 2012, vol. 93, no. 11, pp. 2502–2511.
Whalley, A.J.S., The xylariaceous way of life, Mycol. Res., 1996, vol. 100, no. 8, pp. 897–922. doi.org/ 10.1016/S0953-7562(96)80042-6
Kanematsu, S., Arakawa, M., Oikawa, Y., Onoue, M., Osaki, H., Nakamura, H., Ikeda, K., Kuga-Uetake, Y., Nitta, H., Sasaki, A., Suzaki, K., Yoshida, K., and Matsumoto, N., A reovirus causes hypovirulence of Rosellinia necatrix, Phytopathology, 2004, vol. 94, no. 6, pp. 561–568. doi 10.1094/PHYTO.2004.94.6.561
Chiba, S., Kondo, H., Tani, A., Saisho, D., Sakamoto, W., Kanematsu, S., and Suzuki, N., Widespread endogenization of genome sequences of nonretroviral RNA viruses into plant genomes, PLoS Pathog., 2011, vol. 7, no. 7. e1002146. doi 10.1371/journal.ppat.1002146
Sasaki, A., Miyanishi, M., Ozaki, K., Onoue, M., and Yoshida, K., Molecular characterization of a partitivirus from the plant pathogenic ascomycete Rosellinia necatrix, Arch. Virol., 2005, vol. 150, pp. 1069–1083. doi 10.1007/s00705-005-0494-0
Lin, Y.H., Chiba, S., Tani, A., Kondo, H., Sasaki, A., Kanematsu, S., and Suzuki, N., A novel quadripartite dsRNA virus isolated from a phytopathogenic filamentous fungus, Rosellinia necatrix, Virology, 2012, vol. 426, pp. 42–50. doi 10.1016/j.virol.2012.01.013
Lin, Y.H., Hisano, S., Yaegashi, H., Kanematsu, S., and Suzuki, N., A second quadrivirus strain from the phytopathogenic filamentous fungus Rosellinia necatrix, Arch. Virol., 2013, vol. 158, pp. 1093–1098. doi 10.1007/s00705-012-1580-8
Wei, C.Z., Osaki, H., Iwanami, T., Matsumoto, N., and Ohtsu, Y., Complete nucleotide sequences of genome segments 1 and 3 of Rosellinia anti-rot virus in the family Reoviridae, Arch. Virol., 2004, vol. 149, no. 4, pp. 773–777.
Kondo, H., Kanematsu, S., and Suzuki, N., Viruses of the white root rot fungus, Rosellinia necatrix, Adv. Virus Res., 2013, vol. 86, pp. 177–214. doi 10.1016/B978-0-12-394315-6.00007-6
Coley-Smith, J.R., Verhoeff, K., and Jarvis, W.R., The Biology of Botrytis, London: Academic Press, 1980.
Fravel, D.R., Commercialization and implementation of biocontrol, Annu. Rev. Phytopathol., 2005, vol. 43, pp. 337–359. doi 10.1146/annurev.phyto.43.032904.092924
Castro, M., Kramer, K., Valdivia, L., Ortiz, S., and Castillo, A., A double-stranded RNA mycovirus confers hypovirulence-associated traits to Botrytis cinerea, FEMS Microbiol. Lett., 2003, vol. 228, no. 1, pp. 87–91.
Rodríguez-García, C., Medina, V., Alonso, A., and Ayllon, M.A., Mycoviruses of Botrytis cinerea isolates from different hosts, Ann. Appl. Biol., 2013, vol. 164, pp. 46–61. doi 10.1111/aab.12073
Vilches, S. and Castillo, A., A double-stranded RNA mycovirus in Botrytis cinerea, FEMS Microbiol. Lett., 1997, vol. 155, no. 1, pp. 125–130.
Yu, L., Sang, W., Wu, M.D., Zhang, J., Yang, L., Zhou, Y.J., Chen, W.D., and Li, G.Q., Novel hypovirulence-associated RNA mycovirus in the plant-pathogenic fungus Botrytis cinerea: molecular and biological characterization, Appl. Environ. Microbiol., 2015, vol. 81, no. 7, pp. 2299–2310. doi 10.1128/AEM.03992-14
Cho, W.K., Lee, K.M., Yu, J., Son, M., and Kim, K.H., Insight into mycoviruses infecting Fusarium species, Adv. Virus. Res., 2013, vol. 86, pp. 273–288. doi 10.1016/B978-0-12-394315-6.00010-6
Marvelli, R.A., Hobbs, H.A., Li, S., McCoppin, N.K., Domier, L.L., Hartman, G.L., and Eastburn, D.M., Identification of novel double-stranded RNA mycoviruses of Fusarium virguliforme and evidence of their effects on virulence, Arch. Virol., 2014, vol. 159, no. 2, pp. 349–352. doi 10.1007/s00705-013-1760-1
Kwon, S.J., Lim, W.S., Park, S.H., Park, M.R., and Kim, K.H., Molecular characterization of a dsRNA mycovirus, Fusarium graminearum virus-DK21, which is phylogenetically related to hypoviruses but has a genome organization and gene expression strategy resembling those of plant potex-like viruses, Mol. Cells, 2007, vol. 23, no. 3, pp. 304–315.
Wang, S., Kondo, H., Liu, L., Guo, L., and Qiu, D., A novel virus in the family Hypoviridae from the plant pathogenic fungus Fusarium graminearum, Virus Res., 2013, vol. 174, nos. 1–2, pp. 69–77. doi 10.1016/ j.virusres.2013.03.002
Dyakov, Yu.T., Shnyriova, A.V., and Sergeev, A.Yu., Introduction to the Genetics of Fungi, Moscow: Academia, 2005.
Madhosingh, C., Production of intraspecific hybrids of Fusarium oxysporum f.sp. radicis-lycopersici and Fusarium oxysporum f.sp. lycopersici by protoplast fusions, J. Phytopathol., 1994, vol. 142, nos. 3–4, pp. 301–309. doi 10.1111/j.1439-0434.1994.tb00026.x
van Diepeningen, A.D., Debets, A.J.M., and Hoekstra, R.F., Intra- and interspecies virus transfer in Aspergilli via protoplast fusion, Fungal Genet. Biol., 1998, vol. 25, pp. 171–180. doi.org/10.1006/fgbi.1998.1096
Lakhani, H.N., Vakharia, D.N., Makhlouf, A.H., Eissa, R.A., and Hassan, M.M., Influence of protoplast fusion in Trichoderma spp. on controlling some soil borne diseases, J. Plant Pathol. Microbiol., 2016, vol. 7, no. 8, p. 370. doi 10.4172/2157-7471.1000370
Kanematsu, S., Sasaki, A., Onoue, M., Oikawa, Y., and Ito, T., Extending the fungal host range of a partitivirus and a mycoreovirus from Rosellinia necatrix by inoculation of protoplasts with virus particles, Phytopathology, 2010, vol. 100, no. 3, pp. 922–930. doi 10.1094/PHYTO-100-9-0922
Wang, S., Ongena, M., Qiu, D., and Guo, L., Fungal viruses: promising open fundamental research and biological control agents of fungi, SM Virol., 2017, vol. 2, no. 1, p. 1011.
Author information
Authors and Affiliations
Corresponding authors
Additional information
Translated by A. Panyushkina
About this article
Cite this article
Kyrychenko, A.N., Tsyganenko, K.S. & Olishevska, S.V. Hypovirulence of Mycoviruses as a Tool for Biotechnological Control of Phytopathogenic Fungi. Cytol. Genet. 52, 374–384 (2018). https://doi.org/10.3103/S0095452718050043
Received:
Published:
Issue Date:
DOI: https://doi.org/10.3103/S0095452718050043