Abstract
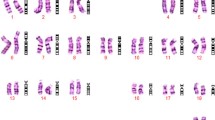
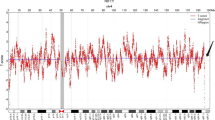
Complex chromosomal rearrangements are rarely observed prenatally. Genetic counceling of CCR carriers is complicated, especially in cases of de novo origin of the rearrangement. Here we present a new case of a de novo CCR involving four chromosomes observed in amniotic fluid cells of the fetus at 17 weeks of gestation. The rearrangement was characterized as an apparently balanced four-way trans-location t(1;11;7;13)(~p21;~q13.5;~q32;~q22)dn by conventional cytogenetic studies. However, array-based comparative genomic hybridization revealed 5 submicroscopic heterozygous interstitial deletions on chromosome 1, 11, 7, 13 with a total loss of 21.1 Mb of genetic material in regions close to those, designated as breakpoints by conventional cytogenetic analysis. The described case clearly illustrates that high-resolution molecular genetic analysis should be combined with conventional cytogenetic techniques to exclude subtle chromosomal abnormalities in CCR cases detected prenatally.
Similar content being viewed by others
References
Pellestor, F., Anahory, T., Lefort, G., Puechberty, J., Liehr, T., Hédon, B., and Sarda, P., Complex chromosomal rearrangements: origin and meiotic behavior, Hum. Reprod. Update, 2011, vol. 17, no. 4, pp. 476–494.
De Gregori, M., Ciccone, R., Magini, P., Pramparo, T., Gimelli, S., et al., Cryptic deletions are a common finding in “balanced” reciprocal and complex chromosome rearrangements: a study of 59 patients, J. Med. Genet., 2007, vol. 44, no. 12, pp. 750–762.
Feenstra, I., Hanemaaijer, N., Sikkema-Raddatz, B., Yntema, H., Dijkhuizen, T., et al., Balanced into array: genome-wide array analysis in 54 patients with an apparently balanced de novo chromosome rearrangement and a meta-analysis, Eur. J. Hum. Genet., 2011, vol. 19, no. 11, pp. 1152–1160.
Miron, P.M., Preparation, culture and analysis of amniotic fluid samples, Curr. Protoc. Hum. Genet., 2012. doi 10.1002/0471142905.hg0804s74
Database of Genomic Variants, http://dgv.tcag.ca/dgv/ app/home.
Giardino, D., Corti, C., Ballarati, L., Colombo, D., Sala, E., et al., De novo balanced chromosome rearrangements in prenatal diagnosis, Prenat. Diagn., 2009, vol. 29, no. 3, pp. 257–265.
Madan, K., Nieuwint, A.W., and van Bever, Y., Recombination in a balanced complex translocation of a mother leading to a balanced reciprocal translocation in the child. Review of 60 cases of balanced complex translocations, Hum. Genet., 1997, vol. 99, no. 6, pp. 806–815.
Gajecka, M., Glotzhach, C.D., Jarmuz, M., Ballif, B.C., and Shaffer, L.G., Identification of cryptic imbalance in phenotypically normal and abnormal translocation carriers, Eur. J. Hum. Genet., 2006, vol. 14, no. 12, pp. 1255–1262.
Kirchhoff, M., Rose, H., and Lundteen, C., High resolution comparative genomic hybridization in clinical cytogenetics, J. Med. Genet., 2001, vol. 38, no. 11, pp. 740–744.
Schluth-Bolard, C., Delobel, B., Sanlaville, D., Boute, O., Cuisset, J.M., et al., Cryptic genomic imbalances in de novo and inherited apparently balanced chromosomal rearrangements: array CGH study of 47 unrelated cases, Eur. J. Med. Genet., 2009, vol. 52, no. 5, pp. 291–296.
Tripputi, P., Bianchi, P., Fermo, E., Bignotto, M., and Zanella, A., Chromosome 7q31.1 deletion in myeloid neoplasms, Hum. Pathol., 2014, vol. 45, no. 2, pp. 368–371.
Weckselblatt, B., Hermetz, K.E., and Rudd, M.K., Unbalanced translocations arise from diverse mutational mechanisms including chromothripsis, Genome Res., 2015, vol. 25, no. 7, pp. 937–947.
Nazaryan, L., Stefanou, E.G., Hansen, C., Kosyakova, N., Bak, M., et al., The strength of combined cytogenetic and mate-pair sequencing techniques illustrated by a germline chromothripsis rearrangement involving FOXP2, Eur. J. Hum. Genet., 2014, vol. 22, no. 3, pp. 338–343.
Genesio, R., Ronga, V., Castelluccio, P., Fioretti, G., Mormile, A., Leone, G. Conti, A., and Cavaliere, M.L., Pure 16q21q22.1 deletion in a complex rearrangement possibly caused by a chromothripsis event, Mol. Cytogenet., 2013, vol. 6, no. 1, p. 1.
Lu, W., Quintero-Rivera, F., Fan, Y., Alkuraya, F.S., Donovan, D.J., et al., NFIA haploinsufficiency is associated with a CNS malformation syndrome and urinary tract defects, PLoS Genet., 2007, vol. 3, no. 5, e80.
Koehler, U., Holinski-Feder, E., Ertl-Wagner, B., Kunz, J., von Moers, A., von Voss, H., and Schell-Apacik, C., A novel 1p31.3p32.2 deletion involving the NFIA gene detected by array CGH in a patient with macrocephaly and hypoplasia of the corpus callosum, Eur. J. Pediatr., 2001, vol. 169, no. 4, pp. 463–468.
Rao, A., O’Donnell, S., Bain, N., Meldrum, C., Shorter, D., and Goel, H., An intragenic deletion of the NFIA gene in a patient with a hypoplastic corpus callosum, craniofacial abnormalities and urinary tract defects, Eur. J. Med. Genet., 2014, vol. 57, nos. 2–3, pp. 65–70.
Author information
Authors and Affiliations
Corresponding author
Additional information
The article is published in the original.
About this article
Cite this article
Pylyp, L.Y., Mykytenko, D.O., Spinenko, L.O. et al. A case of prenatal detection of a de novo unbalanced complex chromosomal rearrangement involving four chromosomes. Cytol. Genet. 50, 339–342 (2016). https://doi.org/10.3103/S009545271605011X
Received:
Published:
Issue Date:
DOI: https://doi.org/10.3103/S009545271605011X