Abstract
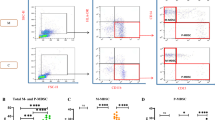
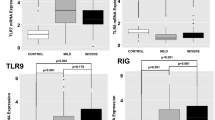
Treatment of severe and critical cases of coronavirus disease 2019 (COVID-19) is still a top priority in public health. Previously, we reported distinct Th1 cytokines related to the pathophysiology of severe COVID-19 condition. In the present study, we investigated the association of Th1 and Th2 cytokine/chemokine endotypes with cell-mediated immunity via multiplex immunophenotyping, single-cell RNA-Seq analysis of peripheral blood mononuclear cells, and analysis of the clinical features of COVID-19 patients. Based on serum cytokine and systemic inflammatory markers, COVID-19 cases were classified into four clusters of increasing (I–IV) severity. Two prominent clusters were of interest and could be used as prognostic reference for a targeted treatment of severe COVID-19 cases. Cluster III reflected severe/critical pathology and was characterized by decreased in CCL17 levels and increase in IL-6, C-reactive protein CXCL9, IL-18, and IL-10 levels. The second cluster (Cluster II) showed mild to moderate pathology and was characterized by predominated CXCL9 and IL-18 levels, levels of IL-6 and CRP were relatively low. Cluster II patients received anti-inflammatory treatment in early-stage, which may have led prevent disease prognosis which is accompanied to IL-6 and CRP induction. In Cluster III, a decrease in the proportion of effector T cells with signs of T cell exhaustion was observed. This study highlights the mechanisms of endotype clustering based on specific inflammatory markers in related the clinical outcome of COVID-19.
Article PDF
Similar content being viewed by others
References
Hasegawa T, Nakagawa A, Suzuki K, et al. Type 1 inflammatory endotype relates to low compliance, lung fibrosis, and severe complications in COVID-19. Cytokine 2021; 148: 155618.
Sun X, Wang T, Cai D, et al. Cytokine storm intervention in the early stages of COVID-19 pneumonia. Cytokine Growth Factor Rev 2020; 53: 38–42.
Gao Y, Li T, Han M, et al. Diagnostic utility of clinical laboratory data determinations for patients with the severe COVID-19. J Med Virol 2020; 92(7):791–6.
Pavel AB, Glickman JW, Michels JR, Kim-Schulze S, Miller RL, Guttman-Yassky E. Th2/Th1 cytokine imbalance is associated with higher COVID-19 risk mortality. Front Genet 2021; 12: 706902.
Wu D, Yang XO. TH17 responses in cytokine storm of COVID-19: an emerging target of JAK2 inhibitor Fedratinib. J Microbiol Immunol Infect 2020; 53(3):368–70.
Chen Z, John Wherry E. T cell responses in patients with COVID-19. Nat Rev Immunol 2020; 20(9):529–36.
Zhou Y, Fu B, Zheng X, et al. Pathogenic T-cells and inflammatory monocytes incite inflammatory storms in severe COVID-19 patients. Natl Sci Rev 2020; 7(6):998–1002.
Sette A, Crotty S. Adaptive immunity to SARS-CoV-2 and COVID-19. Cell 2021; 184(4):861–80.
Spellberg B,, EdwardsJr JE.. Type 1/type 2 immunity in infectious diseases. Clin Infect Dis 2001; 32(1):76–102.
Yamakawa K, Yamamoto R, Terayama T, et al. Japanese rapid/living recommendations on drug management for COVID-19: updated guidelines (September 2021). Acute Med Surg 2021; 8 (1):e706.
Fouladseresht H, Ghamar Talepoor A, Eskandari N, et al. Potential immune indicators for predicting the prognosis of COVID-19 and trauma: similarities and disparities. Front Immunol 2021; 12: 785946.
Vabret N, Britton GJ, Gruber C, et al. Immunology of COVID-19: current state of the science. Immunity 2020; 52(6):910–41.
Kahan SM, Wherry EJ, Zajac AJ. T cell exhaustion during persistent viral infections. Virology 2015;;(479-480):180–93.
Matloubian M, Cyster JG. Th1 cell induction in lymph nodes according to a red-blue chemokine map. Immunity 2012; 37 (6):954–6.
Roncati L, Nasillo V, Lusenti B, Riva G. Signals of Th2 immune response from COVID-19 patients requiring intensive care. Ann Hematol 2020; 99(6):1419–20.
Li CK, Wu H, Yan H, et al. T cell responses to whole SARS coronavirus in humans. J Immunol 2008; 181(8):5490–500.
Huang C, Wang Y, Li X, et al. Clinical features of patients infected with 2019 novel coronavirus in Wuhan, China. Lancet 2020; 395(10223):497–506.
Saeki H, Tamaki K. Thymus and activation regulated chemokine (TARC)/CCL17 and skin diseases. J Dermatol Sci 2006; 43 (2):75–84.
Kakinuma T, Nakamura K, Wakugawa M, et al. Thymus and activation-regulated chemokine in atopic dermatitis: Serum thymus and activation-regulated chemokine level is closely related with disease activity. J Allergy Clin Immunol 2001; 107 (3):535–41.
Beck LA, Thaci D, Hamilton JD, et al. Dupilumab treatment in adults with moderate-to-severe atopic dermatitis. N Engl J Med 2014; 371(2):130–9.
Jameson SC, Masopust D. Understanding subset diversity in T cell memory. Immunity 2018; 48(2):214–26.
Kumar A, Prasoon P, Kumari C, et al. SARS-CoV-2-specific virulence factors in COVID-19. J Med Virol 2021; 93(3): 1343–50.
Schub D, Klemis V, Schneitler S, et al. High levels of SARS-CoV-2-specific T cells with restricted functionality in severe courses of COVID-19. JCI Insight 2020; 5(20):e142167.
Schreiner D, King CG. CD4+ memory T cells at home in the tissue: mechanisms for health and disease. Front Immunol 2018; 9: 2394.
Elsasser J, Janssen MW, Becker F, et al. Antigen-specific CD4 T cells are induced after intravesical BCG-instillation therapy in patients with bladder cancer and show similar cytokine profiles as in active tuberculosis. PLoS One 2013; 8(9):e69892.
Hasegawa T, Okazawa T, Uga H, Kurata H, Mori A. Serum CXCL9 as a potential marker of Type 1 inflammation in the context of eosinophilic asthma. Allergy 2019; 74(12): 2515–8.
Parackova Z, Bloomfield M, Klocperk A, Sediva A. Neutrophils mediate Th17 promotion in COVID-19 patients. J Leukoc Biol 2021; 109(1):73–6.
Bazick HS, Chang D, Mahadevappa K, Gibbons FK, Christopher KB. Red cell distribution width and all-cause mortality in critically ill patients. Crit Care Med 2011; 39 (8):1913–21.
Lin G, Dai C, Xu K, Wu M. Predictive value of neutrophil to lymphocyte ratio and red cell distribution width on death for ST segment elevation myocardial infarction. Sci Rep 2021; 11 (1):11506.
Foy BH, Carlson JCT, Reinertsen E, et al. Association of red blood cell distribution width with mortality risk in hospitalized adults with SARS-CoV-2 infection. JAMA Netw Open 2020; 3 (9):e2022058.
Pascutti MF, Erkelens MN, Nolte MA. Impact of viral infections on hematopoiesis: from beneficial to detrimental effects on bone marrow output. Front Immunol 2016; 7: 364.
Takeuchi A, Saito T. CD4 CTL, a cytotoxic subset of CD4(+) T cells, their differentiation and function. Front Immunol 2017; 8: 194.
Sadanandam A, Bopp T, Dixit S, et al. A blood transcriptome-based analysis of disease progression, immune regulation, and symptoms in coronavirus-infected patients. Cell Death Discov 2020; 6(1):141.
Myasnikov AL, Berns SA, Talyzin PA, Ershov FI. Interferon gamma in the treatment of patients with moderate COVID-19. Vopr Virusol 2021; 66(1):47–54.
Loretelli C, Abdelsalam A, D’Addio F, et al. PD-1 blockade counteracts post-COVID-19 immune abnormalities and stimulates the anti-SARS-CoV-2 immune response. JCI Insight 2021; 6(24):e146701.
Diao B, Wang C, Tan Y, et al. Reduction and functional exhaustion of T cells in patients with coronavirus disease 2019 (COVID-19). Front Immunol 2020; 11: 827.
Vanderbeke L, Van Mol P, Van Herck Y, et al. Monocyte-driven atypical cytokine storm and aberrant neutrophil activation as key mediators of COVID-19 disease severity. Nat Commun 2021; 12(1):4117.
Wherry EJ, Ha SJ, Kaech SM, et al. Molecular signature of CD8 + T cell exhaustion during chronic viral infection. Immunity 2007; 27(4):670–84.
Moskophidis D, Lechner F, Pircher H, Zinkernagel RM. Virus persistence in acutely infected immunocompetent mice by exhaustion of antiviral cytotoxic effector T cells. Nature 1993; 362(6422):758–61.
Utzschneider DT, Legat A, Fuertes Marraco SA, et al. T cells maintain an exhausted phenotype after antigen withdrawal and population reexpansion. Nat Immunol 2013; 14(6):603–10.
Lau CS, Hoo SP, Koh JMJ, Phua SK, Aw TC. Performance of the Roche IL-6 chemiluminescent immunoassay in patients with COVID-like respiratory symptoms. J Virol Methods 2021; 296: 114224.
Acknowledgements
The authors are grateful to Tomohiko Ai for editorial assistance, and Kaori Saito for technical support. We thank Tomoko Sugano and Mao Kuroishi of Central Research Laboratories, Sysmex Corporation for support data acquisition. We thank the Department of Research Support Utilizing Bioresource Bank, and Department of Metabolism & Endocrinology, Juntendo University Graduate School of Medicine for use of their facilities. Simon Young of Oxford Gene Technology for advice on writing this article. Dr Hans Yssel for scientific advice for the manuscript.
Author information
Authors and Affiliations
Contributions
THS established assay system, performed data analysis, and wrote the manuscript. THT, TO, and KI performed data analysis and wrote the manuscript. YM, NI, and KS established assay system and performed data acquisition and analysis. MY established FCM system, performed data acquisition and wrote the manuscript. KY, SY, ET, KN established assay system. AK, FJP, and RO performed data acquisition and wrote the manuscript. TN contributed to sample collection and data interpretation. YT performed data analysis and wrote the manuscript.
Corresponding author
Additional information
Disclosure
Financial support: This work was supported in part by This research was partially supported by AMED under Grant Number JP20fk0108472 to TN. This research was also funded by Sysmex Corporation as an internal company project.
Links of interest: THS, YM, NI, KS, MY, KY, SY, ET, and KN are employed by Sysmex Corporation.
About this article
Cite this article
Hasegawa, T., Hato, T., Okayama, T. et al. Th1 cytokine endotype discriminates and predicts severe complications in COVID-19. Eur Cytokine Netw 33, 1–12 (2022). https://doi.org/10.1684/ecn.2022.0477
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1684/ecn.2022.0477