Abstract
Background
Sepsis is a life-threatening condition of organ dysfunction caused by a dysregulated host immune response to infection. We performed network analysis of cytokine molecules and compared network structures between a systematic inflammatory response syndrome (SIRS) or normal control (NC) group and a sepsis group.
Results
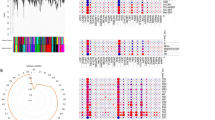
We recruited SIRS (n = 33) and sepsis (n = 89) patients from electronic medical records (EMR) according to whether data on PCT, CRP, interleukin (IL)-1β, IL-2, IL-4, IL-5, IL-6, IL-9, IL-10, IL-12p70, IL-13, IL-17, IL-22, TNF-α, and IFN-γ levels were available. From the public GEO dataset, GSE66099, GSE9960, GSE95233, GSE57065 were downloaded. Genes corresponding to 15 molecules were extracted from an expression array. A correlation matrix was formed for the 15 molecules and statistically significant molecular pairs were used as pairs for network analysis of coexpression. The number of molecular or gene expression pairs significantly correlated among the SIRS or control and sepsis groups are as follows for datasets: EMR, 15 and 15; GEO66099-1, 13 and 15; GEO9960, 13 and 11; GSE95233, 13 and 8; GSE66099-2, 15 and 14; GSE57065, 14 and 13, respectively. Network analysis revealed that network diameter, number of nodes and shortest path were equal to or lower in the sepsis group.
Conclusions
The coexpression network in sepsis patients was relatively small sized and had lower shortest paths compared with the SIRS group or healthy control group. Cytokines with one degree (k = 1) are increased in sepsis group compared with SIRS or healthy control group. IL-9 and IL-2 were not included in network of sepsis group indicating that these cytokines showed no correlation with other cytokines. These data might imply that cytokines tend to be dysregulated in the sepsis group compared to that of SIRS or normal control groups
Similar content being viewed by others
References
Bone RC, Balk RA, Cerra FB, et al. Definitions for sepsis and organ failure and guidelines for the use of innovative therapies in sepsis. Chest 1992; 101: 1644.
Seymour CW, Liu VX, Iwashyna TJ, et al. Assessment of clinical criteria for sepsis for the third international consensus definitions for sepsis and septic shock (Sepsis-3). JAMA 2016; 315:762.
Singer M, Deutschman C, Seymour CW, et al. The third international consensus definitions for sepsis and septic shock (Sepsis-3). JAMA 2016; 315:801.
Angus DC, van der Poll T. Severe sepsis and septic shock. N Engl J Med 2013; 369:840.
Hotchkiss RS, Karl IE. The pathophysiology and treatment of sepsis. N Engl J Med 2003; 348:138.
Kaukonen KM, Bailey M, Pilcher D, Cooper DJ, Bellomo R. Systemic inflammatory response syndrome criteria in defining severe sepsis. N Engl J Med 2015; 372: 1629.
Cohen J. The immunopathogenesis of sepsis. Nature 2002; 420:885.
Boomer JS, To K, Chang KC, et al. Immunosuppression in patients who die of sepsis and multiple organ failure. JAMA 2011; 306: 2594.
Bermejo-Martin JF, Andaluz-Ojeda D, Almansa R, et al. Defining immunological dysfunction in sepsis: a requisite tool for precision medicine. J Infect 2016; 72:525.
Cavailon JM, Adib-Conquy M, Cloez-Tayarani I, Fitting C. Immunodepression in sepsis and SIRS assessed by ex vivo cytokine production is not a generalized phenomenon: a review. J Endotoxin Res 2001; 7:85.
Zhao H, Li W, Lu Z, Sheng Z, Yao Y. The growing specrtrum of anti-inflammatory interleukins and their potential roles in the development of sepsis. J Interferon Cytokine Res 2015; 35:242.
Trevelin S, Alves-Filho J, Sonego F, et al. Toll-like receptor 9 activation in neutrophils impairs chemotaxis and reduces sepsis outcome. Crit Care Med 2012; 40: 2631.
Jekarl DW, Lee S, Lee J, et al. Procalcitonin as a diagnostic marker and IL-6 as a prognostic marker for sepsis. Diagn Microbiol Infect Dis 2013; 75:342.
Jekarl DW, Kim JY, Lee S, et al. Diagnosis and evaluation of severity of sepsis via the use of biomarkers and profiles of 13 cytokines: a multiplex analysis. Clin Chem Lab Med 2015; 53:575.
Tang BM, Huang S, McLean A. Genome-wide transcription profiling of human sepsis: a systematic review. Crit Care 2010; 14: R237.
Sweeney TE, Shidham A, Wong HR, Khatri P. A comprehensive time-course-based multicohort analysis of sepsis and sterile inflammation reveals a robust diagnostic gene set. Sci Transl Med 2015; 7: 287ra71.
Sutherland A, Thomas M, Brandon RA, et al. Development and validation of a novel molecular biomarker diagnostic test for the early detection of sepsis. Crit Care 2011; 15: R149.
Barabasi AL, Oltvai Z. Network biology: understanding the cell’s functional organization. Nat Rev Genet 2004; 5:101.
Barabasi AL, Gulbahce N, Loscalzo J. Network medicine: a networkbased approach to human disease. Nat Rev Genet 2011; 12:56.
Tang B, McLean A, Dawes I, Huang S, Lin R. Gene-expression profiling of peripheral blood mononuclear cells in sepsis. Crit Care Med 2009; 37:882.
Venet F, Schilling J, Cazalis MA, et al. Modulation of LILRB2 and mRNA expression in septic shock patients and after ex vivo lipopolysaccharide stimulation. Hum Immunol 2017; 78:441.
Cazalis MA, Lepape A, Venet F, et al. Early and dynamic changes in gene expression in septic shock patients: a genome-wide approach. Intensive Care Med Exp 2014; 2:20.
Tamayo E, Fernandez A, Almansa R, et al. Pro-and antiinflammatory responses are regulated simultaneously from the first moments of septic shock. Eur Cytokine Netw 2011; 22:82.
Bozza FA, Salluh JI, Japiassu AM, et al. Cytokine profiles as markers of disease severity in sepsis: a multiplex analysis. Crit Care 2007; 11: R49.
Lvovschi V, Arnaud L, Parizot C, et al. Cytokine profiles in sepsis have limited relevance for stratifying patients in the emergency department: a prospective observational study. PLoS One 2011; 6: e28870.
R-project. http://cran.r-project.org (Accessed 2 Feb. 2017).
Davis S, Meltzer P. GEOquery: a bridge between the gene expression omnibus (GEO) and BioConductor. Bioinformatics 2007; 23: 1846.
Falcon S, Morgan M, Gentleman R. An introduction to Bioconductor’s expression set class. 2007. http://www.lcg.unam.mx/∼lcollado/R /resources/Expression-SetIntroduction.pdf (Accessed 2 Feb. 2017).
Cho SB, Kim J, Kim JH. Identifying set-wise differential coexpression in gene expression microarray data. BMC Bioinfomatics 2009; 16:109.
Kim J, Kim K, Kim H, Yoon G, Lee K. Characterization of age signatures of DNA methylation in normal and cancer tissues from multiple studies. BMC Bioinfomatics 2014; 15:997.
Draghici S. Statistics and data analysis for microarray using R and bioconductor, 2nd ed. Boca Raton: CRC Press, 2011.
Pages H, Carlson M, Falcon S, Li N. AnnotationDbi: annotation database interface. R-package version 1.38.0. 2017. https://bioconductor.org/packages/release/bioc/html/AnnotationDbi.html (Accessed 2 Feb. 2017).
Andrade J, Riyue B. Analysis of microarray data with R and Bioconductor. 2013. http//cir.uchicago.edu (Accessed 2 Feb. 2017).
Chang W. R graphics cook book, 1st ed. Sebastopol: O’Reilly, 2013.
Zhu Z, Gerstein M, Snyder M. Getting connected: analysis and principles of biological networks. Genes Dev 2007; 21: 1010.
Doncheva NT, Assenov Y, Domingues FS, Albrecht M. Topological analysis and interactive visualization of biological networks and protein structures. Nat Protoc 2012; 7:670.
Saito R, Smoot ME, Ono K, et al. Atravel guide to cytoscape plugins. Nat Method 2012; 9: 1069.
Netanalyzer. http://med.bioinf.mpi-inf.mpg.de/netanalyzer/(Accessed 15 Apr. 2017).
Xu Ke, Bezakova I, Bunimovich L, Yi SV. Path lengths in proteinprotein interaction networks and biological complexity. Proteomics 2011; 11: 1857.
Zhang Z, Zhan J. A big world inside small-world networks. PLoS One 2009; 4: e5686.
Pavlopoulos GA, Secrier M, Moschopoulos CN, et al. Using graph theory to analyze biological networks. BioData Mining 2011; 4: 10.
Author information
Authors and Affiliations
Corresponding author
About this article
Cite this article
Jekarl, D.W., Kim, K.S., Lee, S. et al. Cytokine and molecular networks in sepsis cases: a network biology approach. Eur Cytokine Netw 29, 103–111 (2018). https://doi.org/10.1684/ecn.2018.0414
Published:
Issue Date:
DOI: https://doi.org/10.1684/ecn.2018.0414