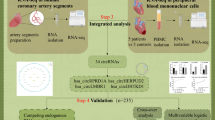
Abstract
Activated nuclear factor-κB (NF-κB) plays an important role in the development of cardiovascular disease (CVD) through its regulated genes and microRNAs (miRNAs). However, the gene regulation profile remains unclear. In this study, primary mouse vascular endothelial cells (pMVECs) were employed to detect CVD-related NF-κB-regulated genes and miRNAs. Genechip assay identified 77 NF-κB-regulated genes, including 45 upregulated and 32 downregulated genes, in tumor necrosis factor α (TNFα)-treated pMVECs. Ten of these genes were also found to be regulated by NF-κB in TNFα-treated HeLa cells. Quantitative real-time PCR (qRT-PCR) assay confirmed the upregulation of Egr1, Tnf, and Btg2 by NF-κB in the TNFα-treated pMVECs. The functional annotation revealed that many NF-κB-regulated genes identified in pMVECs were clustered into classical NF-κB-involved biological processes. Genechip assay also identified 26 NF-κB-regulated miRNAs, of which 21 were upregulated and 5 downregulated, in the TNFα-treated pMVECs. Further analysis showed that nine of the identified genes are regulated by seven of these miRNAs. Finally, among the identified NF-κB-regulated genes and miRNAs, 5 genes and 12 miRNAs were associated with CVD by miRWalk and genetic association database analysis. Taken together, these findings show an intricate gene regulation network raised by NF-κB in TNFα-treated pMVECs. The network provides new insights for understanding the molecular mechanism underlying the progression of CVD.
概要
目 的
鉴定与心血管疾病相关的核因子 κB(NF-κB)调控的基因和小 RNA (microRNA), 探讨疾病发生发展的机制。
创新点
构建心血管疾病相关的NF-κB 调控网络。
方 法
基于 NF-κB 转录活性差异的小鼠原代血管内皮细胞模型, 采用基因芯片 (Genechip) 检测 NF-κB 调控的基因和 microRNA。 再通过实时荧光定量聚合酶链反应 (qRT-PCR) 和生物信息学方法进行差异基因和 microRNA 的筛选、 验证、 功能注释, 从而发现与心血管疾病相关的 NF-κB 调控的基因和 microRNA。
结 论
在肿瘤坏死因子 α (TNFα) 处理的小鼠原代血管内皮细胞中: NF-κB 调控 77 个基因, 其中 45 个基因上调, 32 个基因下调。 NF-κB 还在 TNFα 处理的 HeLa 细胞中调控其中 10 个基因。 通过 qRT-PCR 验证了 NF-κB 上调 Egr1、 Tnf 和 Btg2 的表达。 基因功能注释表明, 许多 NF-κB 调节的基因聚类到经典的 NF-κB 参与的生物学过程。 在 TNFα 处理的小鼠原代血管内皮细胞中还发现: NF-κB 调控 26 个 microRNA, 其中 21 个上调, 5 个下调。 进一步研究发现, 7 个 NF-κB 调控的 microRNA 还可能调控 9 个 NF-κB 调控的基因。 最后通过检索数据库发现, 5 个 NF-κB 调控的基因和 12 个 NF-κB 调控的 microRNA 与心血管疾病相关。 因此, 本研究提升了对心血管疾病进展分子机制的理解。
Similar content being viewed by others
References
Becker KG, Barnes KC, Bright TJ, et al., 2004. The genetic association database. Nat Genet, 36(5):431–432. https://doi.org/10.1038/ng0504-431
Bond M, Fabunmi RP, Baker AH, et al., 1998. Synergistic upregulation of metalloproteinase-9 by growth factors and inflammatory cytokines: an absolute requirement for transcription factor NF-κB. FEBS Lett, 435(1):29–34. https://doi.org/10.1016/S0014-5793(98)01034-5
Borghaei RC, Rawlings PL Jr, Javadi M, et al., 2004. NF-κB binds to a polymorphic repressor element in the MMP-3 promoter. Biochem Biophys Res Commun, 316(1):182–188. https://doi.org/10.1016/j.bbrc.2004.02.030
Boulanger CM, 1999. Secondary endothelial dysfunction: hypertension and heart failure. J Mol Cell Cardiol, 31(1): 39–49. https://doi.org/10.1006/jmcc.1998.0842
Chen L, Wang FY, Zeng ZY, et al., 2017. MicroRNA-199a acts as a potential suppressor of cardiomyocyte autophagy through targeting Hspa5. Oncotarget, 8(38):63825–63834. https://doi.org/10.18632/oncotarget.19133
Chen R, Alvero AB, Silasi DA, et al., 2008. Regulation of IKKβ by miR-199a affects NF-κB activity in ovarian cancer cells. Oncogene, 27(34):4712–4723. https://doi.org/10.1038/onc.2008.112
Chistiakov DA, Sobenin IA, Orekhov AN, et al., 2015. Human miR-221/222 in physiological and atherosclerotic vascular remodeling. Biomed Res Int, 2015:354517. https://doi.org/10.1155/2015/354517
Collart MA, Baeuerle P, Vassalli P, 1990. Regulation of tumor necrosis factor alpha transcription in macrophages: involvement of four κB-like motifs and of constitutive and inducible forms of NF-κB. Mol Cell Biol, 10(4):1498–1506. https://doi.org/10.1128/MCB.10.4.1498
Dweep H, Sticht C, Pandey P, et al., 2011. miRWalk-database: prediction of possible miRNA binding sites by “walking” the genes of three genomes. J Biomed Inform, 44(5):839–847. https://doi.org/10.1016/j.jbi.2011.05.002
Estruch R, 2014. Cardiovascular mortality: how can it be prevented? Nefrologia, 34(5):561–569. https://doi.org/10.3265/Nefrologia.pre2014.Apr.12481
Frantz S, Ertl G, Bauersachs J, 2007. Mechanisms of disease: Toll-like receptors in cardiovascular disease. Nat Clin Pract Cardiovasc Med, 4(8):444–454. https://doi.org/10.1038/ncpcardio0938
Galardi S, Mercatelli N, Farace MG, et al., 2011. NF-κB and c-Jun induce the expression of the oncogenic miR-221 and miR-222 in prostate carcinoma and glioblastoma cells. Nucleic Acids Res, 39(9):3892–3902. https://doi.org/10.1093/nar/gkr006
Hein H, Schlüter C, Kulke R, et al., 1997. Genomic organization, sequence, and transcriptional regulation of the human eotaxin gene. Biochem Biophys Res Commun, 237(3): 537–542. https://doi.org/10.1006/bbrc.1997.7169
Hullinger TG, Montgomery RL, Seto AG, et al., 2012. Inhibition of miR-15 protects against cardiac ischemic injury. Circ Res, 110(1):71–81. https://doi.org/10.1161/CIRCRESAHA.111.244442
Iademarco MF, McQuillan JJ, Rosen GD, et al., 1992. Characterization of the promoter for vascular cell adhesion molecule-1 (VCAM-1). J Biol Chem, 267(23): 16323–16329.
Lerman A, Zeiher AM, 2005. Endothelial function: cardiac events. Circulation, 111(3):363–368. https://doi.org/10.1161/01.CIR.0000153339.27064.14
Li YX, Song YH, Li F, et al., 2009. MicroRNA-221 regulates high glucose-induced endothelial dysfunction. Biochem Biophys Res Commun, 381(1):81–83. https://doi.org/10.1016/j.bbrc.2009.02.013
Li Z, Song Y, Liu L, et al., 2017. miR-199a impairs autophagy and induces cardiac hypertrophy through mTOR activation. Cell Death Differ, 24(7):1205–1213. https://doi.org/10.1038/cdd.2015.95
Lim YC, Garcia-Cardena G, Allport JR, et al., 2003. Heterogeneity of endothelial cells from different organ sites in T-cell subset recruitment. Am J Pathol, 162(5): 1591–1601. https://doi.org/10.1016/S0002-9440(10)64293-9
Liu YX, Wang JK, 2013. Effects of DMSA-coated Fe3O4 nanoparticles on the transcription of genes related to iron and osmosis homeostasis. Toxicol Sci, 131(2):521–536. https://doi.org/10.1093/toxsci/kfs300
Ma XD, Becker Buscaglia LE, Barker JR, et al., 2011. MicroRNAs in NF-κB signaling. J Mol Cell Biol, 3(3): 159–166. https://doi.org/10.1093/jmcb/mjr007
Madamanchi NR, Vendrov A, Runge MS, 2005. Oxidative stress and vascular disease. Arterioscler Thromb Vasc Biol, 25(1):29–38. https://doi.org/10.1161/01.ATV.0000150649.39934.13
Mangge H, Becker K, Fuchs D, et al., 2014. Antioxidants, inflammation and cardiovascular disease. World J Cardiol, 6(6):462–477. https://doi.org/10.4330/wjc.v6.i6.462
Menghini R, Stöhr R, Federici M, 2014. MicroRNAs in vascular aging and atherosclerosis. Ageing Res Rev, 17:68–78. https://doi.org/10.1016/j.arr.2014.03.005
Murdaca G, Colombo BM, Cagnati P, et al., 2012. Endothelial dysfunction in rheumatic autoimmune diseases. Atherosclerosis, 224(2):309–317. https://doi.org/10.1016/j.atherosclerosis.2012.05.013
Newby AC, 2008. Metalloproteinase expression in monocytes and macrophages and its relationship to atherosclerotic plaque instability. Arterioscler Thromb Vasc Biol, 28(12): 2108–2114. https://doi.org/10.1161/ATVBAHA.108.173898
Prisco AR, Hoffmann BR, Kaczorowski CC, et al., 2016. Tumor necrosis factor α regulates endothelial progenitor cell migration via CADM1 and NF-κB. Stem Cells, 34(7): 1922–1933. https://doi.org/10.1002/stem.2339
Santhekadur PK, Das SK, Gredler R, et al., 2012. Multifunction protein staphylococcal nuclease domain containing 1 (SND1) promotes tumor angiogenesis in human hepatocellular carcinoma through novel pathway that involves nuclear factor κB and miR-221. J Biol Chem, 287(17):13952–13958. https://doi.org/10.1074/jbc.M111.321646
Shannon P, Markiel A, Ozier O, et al., 2003. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res, 13(11):2498–2504. https://doi.org/10.1101/gr.1239303
Shiao MS, Chang AYF, Liao BY, et al., 2012. Transcriptomes of mouse olfactory epithelium reveal sexual differences in odorant detection. Genome Biol Evol, 4(5):703–712. https://doi.org/10.1093/gbe/evs039
Song DM, Fang GQ, Mao SZ, et al., 2018. Selective inhibition of endothelial NF-κB signaling attenuates chronic intermittent hypoxia-induced atherosclerosis in mice. Atherosclerosis, 270:68–75. https://doi.org/10.1016/j.atherosclerosis.2018.01.027
Song Y, Wu ZC, Ding W, et al., 2018. NF-κB in mitochondria regulates PC12 cell apoptosis following lipopolysaccharideinduced injury. J Zhejiang Univ-Sci B (Biomed & Biotechnol), 19(6):425–435. https://doi.org/10.1631/jzus.B1700488
Steyers CM III, Miller FJ Jr, 2014. Endothelial dysfunction in chronic inflammatory diseases. Int J Mol Sci, 15(7):11324–11349. https://doi.org/10.3390/ijms150711324
Sun SC, Ganchi PA, Ballard DW, et al., 1993. NF-κB controls expression of inhibitor IκBα: evidence for an inducible autoregulatory pathway. Science, 259(5103):1912–1915. https://doi.org/10.1126/science.8096091
Sun XH, He SL, Wara AKW, et al., 2014. Systemic delivery of microRNA-181b inhibits nuclear factor-κB activation, vascular inflammation, and atherosclerosis in apolipoprotein E-deficient mice. Circ Res, 114(1):32–40. https://doi.org/10.1161/CIRCRESAHA.113.302089
Tousoulis D, Koutsogiannis M, Papageorgiou N, et al., 2010. Endothelial dysfunction: potential clinical implications. Minerva Med, 101(4):271–284.
Vanhoutte PM, 2009. Endothelial dysfunction: the first step toward coronary arteriosclerosis. Circ J, 73(4):595–601. https://doi.org/10.1253/circj.CJ-08-1169
Varley CL, Armitage S, Hassanshahiraviz G, et al., 2003. Regulation of the C-X-C chemokine, mob-1, gene expression in primary rat hepatocytes. Cytokine, 23(3): 64–75. https://doi.org/10.1016/S1043-4666(03)00198-4
Vincenti MP, Coon CI, Brinckerhoff CE, 1998. Nuclear factor κB/p50 activates an element in the distal matrix metalloproteinase 1 promoter in interleukin-1β-stimulated synovial fibroblasts. Arthritis Rheum, 41(11):1987–1994. https://doi.org/10.1002/1529-0131(199811)41:11<1987::AID-ART14>3.0.CO;2-8
Wei W, Zhang WY, Bai JB, et al., 2016. The NF-κBmodulated microRNAs miR-195 and miR-497 inhibit myoblast proliferation by targeting Igf1r, Insr and cyclin genes. J Cell Sci, 129(1):39–50. https://doi.org/10.1242/jcs.174235
Widlansky ME, Gokce N, Keaney JF Jr, et al., 2003. The clinical implications of endothelial dysfunction. J Am Coll Cardiol, 42(7):1149–1160. https://doi.org/10.1016/S0735-1097(03)00994-X
Xing YJ, Zhou F, Wang JK, 2013. Subset of genes targeted by transcription factor NF-κB in TNFα-stimulated human HeLa cells. Funct Integr Genomics, 13(1):143–154. https://doi.org/10.1007/s10142-012-0305-0
Yang JCS, Wu SC, Rau CS, et al., 2015. TLR4/NF-κBresponsive microRNAs and their potential target genes: a mouse model of skeletal muscle ischemia-reperfusion injury. Biomed Res Int, 2015:410721. https://doi.org/10.1155/2015/410721
Zampetaki A, Dudek K, Mayr M, 2013. Oxidative stress in atherosclerosis: the role of microRNAs in arterial remodeling. Free Radic Biol Med, 64:69–77. https://doi.org/10.1016/j.freeradbiomed.2013.06.025
Zhang HC, Taylor WR, Joseph G, et al., 2013. mRNAbinding protein ZFP36 is expressed in atherosclerotic lesions and reduces inflammation in aortic endothelial cells. Arterioscler Thromb Vasc Biol, 33(6):1212–1220. https://doi.org/10.1161/ATVBAHA.113.301496
Zhou F, Wang W, Xing YJ, et al., 2014. NF-κB target microRNAs and their target genes in TNFα-stimulated HeLa cells. Biochim Biophys Acta, 1839(4):344–354. https://doi.org/10.1016/j.bbagrm.2014.01.006
Zhou F, Xu XH, Wang DY, et al., 2017. Identification of novel NF-κB transcriptional targets in TNFα-treated HeLa and HepG2 cells. Cell Biol Int, 41(5):555–569. https://doi.org/10.1002/cbin.10762
Zhu BB, Ye J, Ashraf U, et al., 2016. Transcriptional regulation of miR-15b by c-Rel and CREB in Japanese encephalitis virus infection. Sci Rep, 6:22581. https://doi.org/10.1038/srep22581
Zhu J, Zhu LWS, Yang JH, et al., 2018. Proteomic analysis of human umbilical vein endothelial cells exposed to PM2.5. J Zhejiang Univ-Sci B (Biomed & Biotechnol), 19(6):458–470. https://doi.org/10.1631/jzus.B1700103
Acknowledgments
We thank Prof. Jin-ke WANG (State Key Laboratory of Bioelectronics, Southeast University, Nanjing, China) for his work on this study.
Author information
Authors and Affiliations
Contributions
Fei ZHOU designed the project, performed data analysis, and wrote the paper. Hui ZHU, Yun LI, and Mao-xian WANG conducted the experimental studies, evaluated the data, and partially wrote the manuscript. Wen-xin DU and Ju-hong WANG helped in the data analysis and revised the manuscript. All authors read and approved the final manuscript. Therefore, all authors have taken part in the study and take responsibility for the integrity and security of the data.
Corresponding author
Ethics declarations
Hui ZHU, Yun LI, Mao-xian WANG, Ju-hong WANG, Wen-xin DU, and Fei ZHOU declare that they have no conflict of interest.
All institutional and national guidelines for the care and use of laboratory animals were followed.
Additional information
Project supported by the Natural Science Foundation of Guangdong Province (Nos. 2017A030310606 and 2016A030307039), the Science and Technology Planning Project of Guangdong Province (Nos. 2014A070713039 and 2016A030303063), the Science and Technology Planning Project of Chaozhou City (No. 2016GY18), and the National Natural Science Foundation of China (No. 31770584)
Electronic supplementary material
11585_2019_368_MOESM1_ESM.pdf
Analysis of cardiovascular disease-related NF-κB-regulated genes and microRNAs in TNFα-treated primary mouse vascular endothelial cells
Rights and permissions
About this article
Cite this article
Zhu, H., Li, Y., Wang, Mx. et al. Analysis of cardiovascular disease-related NF-κB-regulated genes and microRNAs in TNFα-treated primary mouse vascular endothelial cells. J. Zhejiang Univ. Sci. B 20, 803–815 (2019). https://doi.org/10.1631/jzus.B1800631
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1631/jzus.B1800631