Abstract
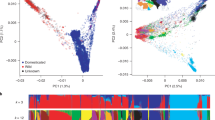
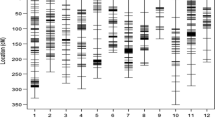
As one of the world’s earliest domesticated crops, barley is a model species for the study of evolution and domestication. Domestication is an evolutionary process whereby a population adapts, through selection; to new environments created by human cultivation. We describe the genome-scanning of molecular diversity to assess the evolution of barley in the Tibetan Plateau. We used 667 Diversity Arrays Technology (DArT) markers to genotype 185 barley landraces and wild barley accessions from the Tibetan Plateau. Genetic diversity in wild barley was greater than in landraces at both genome and chromosome levels, except for chromosome 3H. Landraces and wild barley accessions were clearly differentiated genetically, but a limited degree of introgression was still evident. Significant differences in diversity between barley subspecies at the chromosome level were observed for genes known to be related to physiological and phenotypical traits, disease resistance, abiotic stress tolerance, malting quality and agronomic traits. Selection on the genome of six-rowed naked barley has shown clear multiple targets related to both its specific end-use and the extreme environment in Tibet. Our data provide a platform to identify the genes and genetic mechanisms that underlie phenotypic changes, and provide lists of candidate domestication genes for modified breeding strategies.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Åberg E. 1938. Hordeum agriocrithon nova sp.: a wild six-rowed barley. Ann. Roy. Agric. Coll. Sweden 6:159–216.
Badr, A., Sch. R., Rabey, H.El., Effgen, S., Ibrahim, H., Pozzi, C., Rohde, W., Salamini, F. 2000. On the origin and domestication history of barley (Hordeum vulgare). Mol. Biol. Evol. 17:499–510.
Bauer, E., Weyen, J., Schiemann, A., Graner, A., Ordon, F. 1997. Molecular mapping of novel resistance genes against Barley Mild Mosaic Virus (BaMMV). Theor. Appl. Genet. 95:1263–1269.
Beattie, A.D., Edney, M.J., Scoles, G.J., Rossnagel, B.G. 2010. Association mapping of malting quality data from Western Canadian two-row barley cooperative trials. Crop Sci. 50:1649–1663.
Boyd, W.J.R., Li, C.D., Grime, C.R., Cakir, M., Potipibool, S., Kaveeta, L., Men, S., Kamali, M.R., Barr, A.R., Moody, D.B., Lance, R.C.M., Logue, S.J., Raman, H., Read, B.J. 2003. Conventional and molecular genetic analysis of factors contributing to variation in the timing of heading among spring barley (Hordeum vulgare L.) genotypes grown over a mild winter growing season. Aust. J. Agric. Res. 54:1277–1301.
Bradbury, P.J., Zhang, Z., Kroon, D.E., Casstevens, T.M., Ramdoss, Y., Buckler, E.S. 2007. TASSEL: software for association mapping of complex traits in diverse samples. Bioinformatics 23:2633–2635.
Brueggeman, R., Rostoks, N., Kudrna, D., Kilian, A., Han, F., Chen, J., Druka, A., Steffenson, B., Kleinhofs, A. 2002. The barley stem rust-resistance gene Rpg1 is a novel disease-resistance gene with homology to receptor kinases. Proc. Natl Acad. Sci. USA 99:9328–9333.
Cai, S., Wu, D., Jabeen, Z., Huang, Y., Huang, Y., Zhang, G. 2013. Genome-wide association analysis of aluminum tolerance in cultivated and Tibetan wild barley. PLoS ONE 8:e69776.
Cakir, M., Poulsen, D., Galwey, N.W., Ablett, G.A., Chalmers, K.J., Platz, G.J., Park, R.F., Lance, R.C.M., Panozzo, J.F., Read, B.J., Moody, D.B., Barr, A.R., Johnston, P., Li, C.D., Boyd, W.J.R., Grime, C.R., Appels, R., Jones, M.G.K, Langridge, P. 2003. Mapping and QTL analysis of the barley population Tallon × Kaputar. Aust. J. Agric. Res. 54:1155–1162.
Collins, H.M., Panozzo, J.F., Logue, S.J., Jefferies, S.P., Barr, A.R. 2003. Mapping and validation of chromosome regions associated with high malt extract in barley (Hordeum vulgare L.). Aust. J. Agric. Res. 54:1223–1240.
Coventry, S.J., Barr, A.R., Eglinton, J.K., Mcdonald, G.K. 2003. The determinants and genome locations influencing grain weight and size in barley (Hordeum vulgare L.). Aust. J. Agric. Res. 54:1103–1115.
Dai, F., Nevo, E., Wu, D., Comadran, J., Zhou, M., Qiu L., Chen, Z., Beiles, A., Chen, G., Zhang, G. 2012. Tibet is one of the centers of domestication of cultivated barley. Proc. Natl Acad. Sci. USA. 109:16969–16973.
Doebley, J.F., Gaut, B.S., Smith, B.D. 2006. The molecular genetics of crop domestication. Cell 127:1309–1321.
Doyle, J.J. 1990. Isolation of plant DNA from fresh tissue. Focus 12:13–15.
Edwards, M.C., Steffenson, B.J. 1996. Genetics and mapping of barley stripe mosaic virus resistance in barley. Phytopathol. 86:184–187.
Excoffier, L., Lischer, H.E. 2010. Arlequin suite ver 3.5: a new series of programs to perform population genetics analyses under Linux and Windows. Mol. Ecol. Resour. 10:564–567.
Faure, S., Turner, A.S., Gruszka, D., Christodoulou, V., Davis, S.J., von Korff, M., Laurie, D.A. 2012. Mutation at the circadian clock gene EARLY MATURITY 8 adapts domesticated barley (Hordeum vulgare) to short growing seasons. Proc. Natl. Acad. Sci. USA. 109:8328–8333.
Garvin, D.F., Millergarvin, J.E., Viccars, E.A., Jacobsen, J.V., Brown, A.H.D. 1998. Identification of molecular markers linked to ant28–484, a mutation that eliminates proanthocyanidin production in barley seeds. Crop Sci. 38:1250–1255.
Genger, R.K., Williams, K.J., Raman, H., Read, B.J., Wallwork, H., Burdon, J.J., Brown, A.H.D. 2003. Leaf scald resistance genes in Hordeum vulgare and Hordeum vulgare ssp. spontaneum: parallels between cultivated and wild barley. Aust. J. Agric. Res. 54:1335–1342.
Graner, A., Tekauz, A. 1996. RFLP mapping in barely of a dominant gene conferring resistance to scald (Rhynchosporium secalis). Theor. Appl. Genet. 93: 421–425.
Hill, W.G., Robertson, A. 1968. Linkage disequilibrium in finite populations. Theor. Appl. Genet. 38:226–231.
Holsinger, K.E., Lewis, P.O., Dey, D.K. 2002. A Bayesian approach to inferring population structure from dominant markers. Mol. Ecol. 11:1157–1164.
Hsu, T.W. 1975. On the origin and phylogeny of cultivated barley with reference to the discovery of Ganze wild two-rowed barley Hordeum spontaneum C. Koch. Acta Genet. Sin. 2:129–137. (in Chinese with English abstract)
Igartua, E., Moralejo, M., Casas, A.M., Torres, L., Molina-Cano, J.L. 2012. Whole-genome analysis with SNPs from BOPA1 shows clearly defined groupings of Western Mediterranean, Ethiopian, and Fertile Crescent barleys. Genet. Resour. Crop. Evol. 60:251–264.
Jefferies, S.P., Barr, A.R., Karakousis, A., Kretschmer, J.M., Manning, S., Chalmers, K.J., Nelson, J.C., Islam, A.K.M.R., Langridge, P. 1999. Mapping of chromosome regions conferring boron toxicity tolerance in barley (Hordeum vulgare L.). Theor. Appl. Genet. 98:1293–1303.
Kikuchi, R., Kawahigashi, H., Oshima, M., Ando, T., Handa, H. 2012. The differential expression of HvCO9, a member of the CONSTANS-like gene family, contributes to the control of flowering under short-day conditions in barley. J. Exp. Bot. 63:773–784.
Larson, S.R., Kadyrzhanova, D., Mcdonald, C., Sorrells, M., Blake, T.K. 1996. Evaluation of barley chromosome- 3 yield QTLs in a backcross F2 population using STS-PCR. Theor. Appl. Genet. 93:618–625.
Li, C., Ni, P., Francki, M., Hunter, A., Zhang, Y., Schibeci, D., Li, H., Tarr, A., Wang, J., Cakir, M., Yu, J., Bellgard, M. Lance, R., Appels, R. 2004. Genes controlling seed dormancy and pre-harvest sprouting in a rice-wheat-barley comparison. Funct. Integr. Genomics. 4:84–93.
Li, C.D., Lance, R.C.M., Collins, H.M., Tarr, A., Roumeliotis, S., Harasymow, S., Cakir, M., Fox, G.P., Grime, C.R., Broughton, S., Young, K.J., Raman, H., Barr, A.R., Moody, D.B., Read, B.J. 2003. Quantitative trait loci controlling kernel discoloration in barley (Hordeum vulgare L.). Aust. J. Agric. Res. 54:1251–1259.
Lonergan, P. F. 2001. Genetic characterisation and QTL mapping of zinc nutrition in barley (Hordeum vulgare). Ph.D. thesis, University of Adelaide, Australia. http://hdl.handle.net2440/21718
Mano, Y., Takeda, K. 1997. Mapping quantitative trait loci for salt tolerance at germination and the seedling stage in barley (Hordeum vulgare L.). Euphytica 94:263–272.
Meyer, R.C., Swanston, J.S., Young, G.R., Lawrence, P.E., Bertie, A., Ritchie, J., Wilson, A., Brosnan, J., Pearson, S., Bringhurst, T., Steele, G., Aldis, P.R., Filed, M., Jolliffe, T., Powell, W., Thomas, W.T.B. 2001. A genome based approach to improving barley for the malting and distilling industries. HGCA Project Report 264. http://cereals.ahdb.org.ukmedia/379983/264_complete_final_report.pdf Scottish Crop Research Institute pp. 70–74.
Nei, M. 1978. Estimation of average heterozygosity and genetic distance from a small number of individuals. Genetics 89:583–590.
Nevo, E. 2013. Evolution of wild barley and barley improvement. In: Zhang, G.P., Li, C.D., Liu, X. (eds), Advance in barley sciences. Springer. Dordrecht, The Netherlands. pp. 1–23.
Park, R.F., Karakousis, A. 2002. Characterization and mapping of gene Rph19 conferring resistance to Puccinia hordei in the cultivar ‘Reka 1’ and several Australian barleys. Plant Breed. 121:232–236.
Pourkheirandish, M., Hensel, G., Kilian, B., Senthil, N., Chen, G., Sameri, M., Azhaguvel, P., Sakuma, S., Dhanagond, S., Sharma, R., Mascher, M., Himmelbach, A., Gottwald, S., Nair, S.K., Tagiri, A., Yukuhiro, F., Nagamura, Y., Kanamori, H., Matsumoto, T., Willcox, G., Middleton, C.P., Wicker, T., Walther, A., Waugh, R., Fincher, G.B., Stein, N., Kumlehn, J., Sato, K., Komatsuda, T. 2015. Evolution of the grain dispersal system in barley. Cell 162:527–539.
Pourkheirandish, M., Komatsuda, T. 2007. The importance of barley genetics and domestication in a global perspective. Ann. Bot. 100:999–1008.
Pritchard, J.K., Wen, X., Falush, D. 2010. Documentation for structure software: Version 2.3. University of Chicago, Chicago, IL. Raman, H., Platz, G.J., Chalmers, K.J., Raman, R., Read, B.J., Barr, A.R., Moody, D.B. 2003. Mapping of genomic regions associated with net form of netblotch resistance in barley. Aust. J. Agric. Res. 54:1359–1367.
Read, B.J., Raman, H., Mcmichael, G., Chalmers, K.J., Ablett, G.A., Platz, G.J., Raman, R., Genger, R.K., Boyd, W.J.R., Li, C.D., Grime, C.R., Park, R.F., Wallwork, H., Prangnell, R., Lance, R.C.M. 2003. Mapping and QTL analysis of the barley population Sloop × Halcyon. Aust. J. Agric. Res. 54:1145–1153.
Ren, X., Nevo, E., Sun, D., Sun, G. 2013. Tibet as a potential domestication center of cultivated barley of China. PLoS ONE 8:e62700.
Rohlf, F.J. 2000. NTSYS-pc Numerical taxonomy and multivariate analysis system, version 2.1. Exeter Software, New York.
Rostoks, N., Ramsay, L., MacKenzie, K., Cardle, L., Bhat, P.R., Roose, M.L., Svensson, J.T., Stein, N., Varshney, R.K., Marshall, D.F., Graner, A., Close, T.J., Waugh, R. 2006. Recent history of artificial out-crossing facilitates whole-genome association mapping in elite inbred crop varieties. Proc. Natl Acad. Sci. USA. 103:18656–18661.
Roy, J.K., Smith, K.P., Muehlbauer, G.J., Chao, S., Close, T.J., Steffenson, B.J. 2010. Association mapping of spot blotch resistance in wild barley. Mol. Breed. 26:243–256.
Russell, J., Dawson, I.K., Flavell, A.J., Steffenson, B., Weltzien, E., Booth, A., Ceccarelli, S., Grando, S., Waugh, R. 2011. Analysis of >1000 single nucleotide polymorphisms in geographically matched samples of landrace and wild barley indicates secondary contact and chromosome-level differences in diversity around domestication genes. New Phytol. 191:564–578.
Shoeva, O.Y., Mock, H.-P., Kukoeva, T.V., Börner, A., Khlestkina, E.K. 2016. Regulation of the flavonoid Biosynthesis Pathway Genes in Purple and Black Grains of Hordeum vulgare. PloS one 11:e0163782
Spaner, D., Shugar, L.P., Choo, T.M., Falak, I., Briggs, K.G., Legge, W.G., Falk, D.E., Ullrich, S.E., Tinker, N.A., Steffenson, B.J., Mather, D.E. 1998. Mapping of disease resistance loci in barley on the basis of visual assessment of naturally occurring symptoms. Crop. Sci. 38:843–850.
Steffenson, B.J., Hayes, P. M., Kleinhofs, A. 1996. Genetics of seedling and adult plant resistance to net blotch (Pyrenophora teres f. teres) and spot blotch (Cochliobolus sativus) in barley. Theor. Appl. Genet. 92:552–558.
Tanno, K., Taketa, S., Takeda, K., Komatsuda, T. 2002. A DNA marker closely linked to the vrs1 locus (row-type gene) indicates multiple origins of six-rowed cultivated barley (Hordeum vulgare L.). Theor. Appl. Genet. 104:54–60.
Tashi, N., Tang, Y., Zeng, X. 2013. Food preparation from hulless barley in Tibet. In: Zhang, G.P., Li, C.D., Liu, X. (eds), Advance in barley sciences. Springer. Dordrect, The Netherlands. pp. 151–158.
Varshney, R.K., Paulo, M.J., Grando, S., van Eeuwijk, F.A., Keizer, L.C.P., Guo, P., Ceccarelli, S., Kilian, A., Baum, M., Graner, A. 2012. Genome wide association analyses for drought tolerance related traits in barley (Hordeum vulgare L.). Field Crop Res. 126:171–180.
von Bothmer, R., Sato, K., Komatsuda, T., Yasuda, S., Fischbeck, G. 2003. The domestication of cultivated barley. In: von Bothmer, R., van Hintum, T., Knüpffer, H., Sato, K. (eds), Diversity in barley (Hordeum vulgare). Elsevier, Amsterdam. pp. 9–27.
Wang, L., Xu, J.Q., Xia, T.F., Zhang, H.G., Liu, D.C., Shen, Y.H. 2014. Population structure and linkage disequilibrium in six-rowed barley landraces from the Qinghai-Tibetan Plateau. Crop. Sci. 54:2011–2022.
Wenzl, P., Li, H., Carling, J., Zhou, M., Raman, H., Paul, E., Hearnden, P., Maier, C., Xia, L., Caig, V., Ovesná, J., Cakir, M., Poulsen, D., Wang, J., Raman, R., Smith, K.P., Muehlbauer, G.J., Chalmer, K.J., Kleinhofs, A., Huttner, E., Kilian, A. 2006. A high-density consensus map of barley linking DArT markers to SSR, RFLP and STS loci and agricultural traits. BMC Genomics 7:206.
Williams, K.J., Lichon, A., Gianquitto, P., Kretschmer, J.M., Karakousis, A., Manning, S., Langridge, P., Wallwork, H. 1999 Identification and mapping of a gene conferring resistance to the spot form of net blotch (Pyrenophora teres f. maculata) in barley. Theor. Appl. Genet. 99:323–327.
Wu, D., Qiu, L., Xu, L., Ye, L., Chen, M., Sun, D., Chen, Z., Zhang, H., Jin, X., Dai, F., Zhang, G. 2011. Genetic variation of HvCBF genes and their association with salinity tolerance in Tibetan annual wild barley. PLoS ONE 6:e22938.
Zhang, F., Chen, G., Huang, Q., Orion, O., Krugman, T., Fahima, T., Korol, A.B., Nevo, E., Gutterman, Y. 2005. Genetic basis of barley caryopsis dormancy and seedling desiccation tolerance at the germination stage. Theor. Appl. Genet. 110:445–453.
Zhou, T.-S., Takashi, I., Ryouichi, K., Naohiko, H., Makoto, K., Takehiro, H., Kazuhiro, S. 2012. Malting quality quantitative trait loci on a high-density map of Mikamo golden × Harrington cross in barley (Hordeum vulgare L.). Mol. Breed. 30:103–112.
Author information
Authors and Affiliations
Corresponding authors
Additional information
Communicated by H. Grausgruber
Electronic supplementary material
Rights and permissions
This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Xu, J.Q., Wang, L., Liu, B.L. et al. Genome-wide Scan Using DArT Markers for Selection Footprints in Six-rowed Naked Barley from the Tibetan Plateau. CEREAL RESEARCH COMMUNICATIONS 46, 591–603 (2018). https://doi.org/10.1556/0806.46.2018.041
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1556/0806.46.2018.041