Abstract
Background
The burden of hereditary breast cancer in India is not well defined. Moreover, genetic testing criteria (National Comprehensive Cancer Network [NCCN] and Mainstreaming Cancer Genetics [MCG] Plus) have never been validated in the Indian population.
Methods
All new female breast cancer patients from 1st March 2019 to 28th February 2020 were screened. Those providing informed consent and without previous genetic testing were recruited. Multigene panel testing (107 genes) by next-generation sequencing was performed for all patients. The frequency of pathogenic/likely pathogenic (P/LP) mutations between patients qualifying and not qualifying the testing criteria was compared and their sensitivity was computed.
Results
Overall, 275 breast cancer patients were screened and 236 patients were included (median age 45 years); 30 patients did not consent and 9 patients previously underwent genetic testing. Thirty-four (14%) women had a positive family history and 35% had triple-negative breast cancer. P/LP mutations were found in 44/236 (18.64%) women; mutations in BRCA1 (22/47, 46.8%) and BRCA2 (9/47, 19.1%) were the most common, with 34% of mutations present in non-BRCA genes. Patients qualifying the testing criteria had a higher risk of having a P/LP mutation (NCCN: 23.6% vs. 7.04%, p = 0.03; MCG plus: 24.8% vs. 7.2%, p = 0.01). The sensitivity of the NCCN criteria was 88.6% (75.4–96.2) and 86.36% (72.65–94.83) for MCG plus. More than 95% sensitivity was achieved if all women up to 60 years of age were tested. Cascade testing was performed in 31 previous (16/44 families), with 23 testing positive.
Conclusions
The frequency of P/LP mutations in India is high, with significant contribution of non-BRCA genes. Testing criteria need modification to expand access to testing.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Breast cancer is the most common cancer among Indian women and worldwide, and constitutes the most common cause of cancer-related mortality among women.1,2 According to Western literature, around 10–15% of breast cancers can be attributed to germline pathogenic mutations, with BRCA1/2 causing around 40–50% of hereditary breast cancers (4–6% of the unselected breast cancer population).3,4,5,6,7 With increasing use of multigene panel testing, mutations in other breast cancer-causing genes, such as PALB2, ATM, CHEK, and TP53, are being increasingly detected.8,9 The Indian scenario is characterized by a younger median age of onset (< 50 years) and a higher prevalence of triple-negative breast cancer (TNBC; up to 30%), which when combined with delayed access to effective breast cancer screening programs and limited access to treatment leads to a low incidence-to-mortality ratio compared with the West.10,11,12,13 Since BRCA mutations are enriched in the young TNBC population, the population prevalence of these mutations is expected to be higher in India.14
There are two main issues in the Indian context that need to be addressed. First, the genetic burden of breast cancer is not well-defined in the Indian population. Previous studies have reported a high prevalence of pathogenic mutations, however they were performed on non-consecutive patients with a limited gene panel and suffered from strong selection and referral bias.15,16,17 Second, it remains unclear as to which patients should be offered genetic testing. National Comprehensive Cancer Network (NCCN) and Mainstreaming Cancer Genetics (MCG) Plus criteria are commonly used to select patients for testing, however other groups have used even more restrictive criteria for testing without any definite evidence for doing so.18,19 Recent studies have highlighted the limitations of the NCCN criteria when applied to a North American population20,21,22 and have suggested expanding the testing to include all patients up to the age of 65 years to improve sensitivity.20 However, no such data are available from India and the application of these criteria to the Indian population remains empiric.
In the present study, we subjected all consecutive patients with breast cancer attending the breast cancer clinic (BCC) at our institute over a one-year period to multigene panel testing by next-generation sequencing (NGS). Our objective was to define the prevalence and describe the mutational profile of hereditary breast cancer, as well as to test the performance of NCCN and MCG Plus criteria in our population.
Methods
All new patients above 18 years of age registered at the BCC over a 1-year period from 1st March 2019 to 28th February 2020, with a diagnosis of invasive breast cancer, were screened for participation in this study. Patients who had not undergone genetic testing previously and who provided written informed consent were included. A detailed clinical history was recorded in a predesigned proforma. Staging was performed according to the American Joint Committee on Cancer (AJCC) 8th edition. A three-generation pedigree chart was built using standard symbols. Patients were classified into those qualifying either the NCCN 2018 criteria or MCG Plus criteria23 for testing and those who did not using a predesigned checklist (electronic supplementary Tables e1 and e2). After pretest counseling by a medical oncologist, 5 mL of peripheral blood was taken from each patient and a targeted NGS was performed using the Trusight cancer sequencing panel (Illumina, San Diego, CA, USA) covering 107 high-risk genes associated with cancer predisposition at a centralized College of American Pathologists (CAP) and Clinical Laboratory Improvement Amendments (CLIA) certified laboratory. The reflex multiplex ligation by probe amplification (MLPA) method was performed in all cases that were negative by NGS to identify large genomic rearrangements (LGRs) in BRCA1 and 2 genes. Technical details, including the genes that were tested as part of the panel, are provided in the appendix (electronic supplementary e3 and Table e4). All variants obtained by NGS were classified as benign, likely benign, pathogenic, likely pathogenic, and variant of uncertain significance (VUS) according to the American College of Medical Genetics (ACMG) guidelines. Sanger sequencing was used to confirm single nucleotide variants (SNVs) and small insertions and deletions (indels). All patients received post-test counseling. Patients having pathogenic/likely pathogenic (P/LP) mutations in any of the breast cancer-related genes were managed according to NCCN guidelines. Cascade testing was initiated to identify unaffected previvors (Fig. 1). The study was approved by the Institute Ethics Committee (IEC) and was conducted according to good clinical practice (GCP) guidelines as outlined in the Declaration of Helsinki.
Statistical Analysis
Descriptive statistics were used to describe the baseline characteristics of the entire cohort. The Chi-square test was used to compare the rates of P/LP mutations in patients qualifying and not qualifying the testing criteria. A p value < 0.05 was considered significant. Sensitivity, specificity, positive predictive value, and negative predictive value of both the NCCN and MCG Plus criteria at different cut-off ages was calculated using 2 × 2 tables. All analysis was performed using STATA software version 13 (StataCorp LLC, College Station, TX, USA).
Results
Study Population
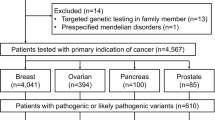
A total of 275 breast cancer patients were screened and 236 women with breast cancer were recruited in the study. Thirty patients (10.9%) did not provide consent and nine (3.3%) had previously undergone genetic testing. Median age of the entire cohort was 45 years (18–80 years); 73 (31.1%) women were under 40 years of age, 77 (32.8%) were between 40 and 49 years, 48 (20.4%) were between 50 and 59 years, and 38 (16.1%) were aged 60 years or above. Thirty-four women (14.4%) had a significant family history suggestive of HBOC. Stage one tumors were uncommon (18/236, 7.6%), and an almost equal proportion of women had stage II and III disease (35.1% and 34.3% respectively). Around 35.2% (83/236) of patients had TNBC among our cohort, with 45.8% (108/236) and 33.5% (79/236) having hormone-positive and human epidermal growth factor receptor 2 (HER2)-positive disease, respectively (Table 1 and electronic supplementary Table e5)
Mutational Profile in the Overall Cohort
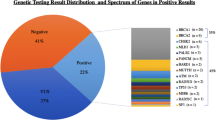
A total of 44/236 (18.64%) patients had a P/LP mutation; however, 47 P/LP mutations were seen, as three patients had two simultaneous P/LP mutations in different genes. Thirty-four (72.3%) mutations were present in one of the high penetrance genes [22 BRCA1 (46.8%), 9 BRCA2 (19.1%), 3 TP53 (6.3%)] and 11 mutations (23.4%) were seen in moderate penetrance genes (4 PALB2, 2 RAD50, 2 ATM, 2 FANCI, 1 RAD51D). One mutation each was identified in MSH2 and MUTYH. Both these mutations were subsequently confirmed on Sanger sequencing but these patients did not have any other features suggestive of hereditary colon cancer syndrome (Fig. 2 and electronic supplementary Table e5). Interestingly, only one patient with a TP53 mutation had the features of Li-Fraumeni syndrome. We identified only one patient with an Ashkenazi Jewish (AJ) founder mutation. A VUS was detected in 89 patients (37.71%), yielding a VUS/pathogenic ratio of 2.02. The VUS rates were not significantly different in women qualifying and not qualifying the NCCN criteria (34.5% vs. 45%, p = 0.14). No LGRs were detected by MLPA in patients who were negative by NGS.
Comparison of National Comprehensive Cancer Network (NCCN) Qualifiers and Non-Qualifiers
A total of 165 patients (69.9%) fulfilled the NCCN criteria for testing; 88 (37.3%), 59 (25%), 16 (6.8%), and 2 (0.8%) women fulfilled one, two, three, and four of the NCCN criteria, respectively (Table 2). Median age was younger in the NCCN qualifiers (39 years vs. 50 years). All patients with a significant family history and with metachronous breast cancer qualified the NCCN criteria for testing. Although baseline stage was equitably distributed in both groups, more patients in the NCCN qualifier group had TNBC (p = 0.003). Patients who qualified the NCCN criteria had a significantly higher incidence of P/LP mutation (23.6% vs. 7.04%, p = 0.03) or a BRCA1/2 mutation (18.18% vs. 1.4%, p < 0.001) (Table 3).
Comparison of Mainstreaming Cancer Genetics (MCG) Plus Qualifiers and Non-Qualifiers
Overall, 153 (64.8%) patients qualified the MCG Plus criteria for testing, whereas 83 (35.2%) patients did not (electronic supplementary Table e7). Baseline characteristics between the two groups are summarized in electronic supplementary Table e6. As among the NCCN groups, patients who qualified the MCG Plus criteria were younger and had a higher probability of having TNBC (p < 0.001) [electronic supplementary Table e6]. A significantly higher proportion of patients qualifying the MCG Plus criteria had a P/LP or BRCA1/2 mutation compared with those not qualifying the criteria (24.8% vs. 7.2%, p = 0.01; and 19.6% vs. 1.2%, p < 0.001) (Table 3).
Characteristics of Patients with Pathogenic/Likely Pathogenic Mutations but Not Qualifying the Testing Criteria
A total of five (11.36%) and six patients (13.6%) would have been missed if only the NCCN and MCG Plus criteria had been used for selecting patients for testing, respectively. Analyzing these six patients, all of whom were > 45 years of age (46–64 years), all five had hormone-positive tumors, while four had HER2-positive tumors. The mutations found were variable, with one patient each having mutation in the RAD50, TP53, MUTYH, MSH2, BRCA1 and PALB2 genes (supplementary e8).
Sensitivity of the NCCN and MCG Plus Criteria
The sensitivity of the NCCN and MCG Plus criteria for picking up patients with pathogenic mutations was 88.6% (75.4–96.2) and 86.36% (72.65–94.83) respectively (Table 4). When age criteria were adjusted to include all women < 40 years of age, the sensitivity decreased to 79.55% (64.7–90.2%); however, if all women up to the age of 60 years were to be tested, sensitivity of both the NCCN and MCG Plus criteria would have increased to 97.73% (87.98–99.94), with only one patient not being picked up. Sensitivity of both criteria at various cut-off ages is given in Table 5. Relaxing the cut-off age to 60 years (from 45 years for MCG Plus and NCCN 2019) would have meant testing an additional 22 women (9.3%) for picking up five additional mutations, giving a yield of 22.72%. It would also mean leaving 22 women out of the entire cohort untested (9.3%) and one mutation not being picked up.
Interventions Based on Genetic Testing Results
Of the 31 patients who tested positive for a mutation in any of the BRCA genes, 27 were offered a contralateral risk-reducing mastectomy (RRM) and 24 were offered a bilateral salpingo-oophorectomy (RRBSO). Ten patients underwent an RRM (37.37%) and seven underwent an RRBSO (29.16%). Occult malignancy was not identified in any of the operated patients. The delay in elective surgeries during the coronavirus disease 2019 (COVID-19) pandemic was the major reason for patients not opting for risk-reducing interventions (14/17, 82.35%). Only three patients refused surgery due to personal reasons (2) or fear of complications even after extended counseling (1). The family members of 16 patients with a P/LP mutation were able to undergo cascade testing, which was also affected by the COVID-19 pandemic. A total of 31 of 310 (10%) first-degree relatives of affected patients were tested, of whom a P/LP mutation was identified in 23 previvors (74.19%) [electronic supplementary Table e9]
Discussion
In this prospective study of consecutive breast cancer patients from a North Indian tertiary care cancer center, we report a high prevalence of P/LP mutations (18.64%) in hereditary cancer susceptibility genes among breast cancer patients. It is also evident that although patients qualifying the NCCN/MCG Plus criteria have a higher likelihood of having a P/LP mutation, a finding that is similar to some studies and discordant with other studies,20,21,22 a significant proportion of patients are missed by these restrictive criteria (11.3 and 13.6%, respectively), thus confirming their relative insensitivity when applied to an Indian population.
Few studies from India have reported on the prevalence and spectrum of pathogenic mutations. The largest study by Mannan et al., which was published in 2016 and later updated in 2018, evaluated 1010 patients with either breast or ovarian cancers, or both, and found a mutation rate of 30.3%.16,17 In their study, around 80% of patients qualified the NCCN criteria and those qualifying had a higher rate of mutation (34.4%) compared with those not qualifying the criteria (13.5%). Around 85% of patients in this cohort had mutation in the BRCA1/2 gene. A similar laboratory-based study by Chheda et al. analyzed the prevalence of BRCA1/2 mutations in 160 women fulfilling the NCCN criteria for testing and found a mutation frequency of 31.2%.15 However, these studies were laboratory based and were conducted on a non-consecutive population of mixed breast and ovarian cancer patients, and thus suffered from significant referral and selection bias and likely overestimated the prevalence of pathogenic mutations. Although we recognize the limitation of a relatively small sample size in our study, the prevalence of 18.6% reported in our study is likely a closer estimate of the actual prevalence of pathogenic mutations in breast cancer patients, as consecutive patients were studied. This prevalence is significantly higher than what is reported from large Western literature and is similar to the young TNBC rich African population (11%).3,4,20,21,24,25,26 We also found a higher proportion of non-BRCA mutations compared with Singh et al. (34% vs. 15%), as we used a larger and more comprehensive panel.16 We found a higher prevalence of P/LP mutations despite only 14% of patients having a significant family history, compared with almost 50% of patients with a significant history in large US studies.20,21
There was a significant difference in the frequency of P/LP mutations between the NCCN qualifiers and non-qualifiers in our study, which is in stark contrast to the findings in the American populations.21 As both these studies had consecutive patients and used large multigene panels, the spectrum of mutations appears different in India. However, Yadav et al. quoted a significant difference between NCCN qualifiers and non-qualifiers, but this was likely due to the use of a small restrictive panel of genes.20
We did not find any recurrent mutations in our cohort and only one patient had an AJ founder mutation (BRCA1 c68_69 delAG). This spectrum of mutations is significantly different from what has been previously reported from India.15,16,18,19,27,28,29 Studies from Western and Southern India have reported a high prevalence of founder mutation, particularly c68_69 delAG, with a frequency of 0.5–4.1%.17,28,30 Singh et al. reported a founder mutation prevalence of 3.5%. Other recurrent mutations, including c.5074 + 1G > A, which have been previously reported in Marwari families of Western India, were absent in our population.16 The absence of founder mutations in a predominantly North Indian population signifies that a strategy of screening patients by testing for founder mutations, as followed in some parts of India, is unlikely to be fruitful.
We also tried to address which patients should undergo genetic testing in India. Data are available to suggest that 50–80% of individuals at risk of having a hereditary breast cancer are never tested as they do not meet the current criteria31,32 and, overall, < 10% of patients carrying a BRCA1/2 mutation have been identified.33 Recent guidelines from the American Society of Breast Surgeons (ASBRS) recommend the testing of all women with breast cancer for pathogenic mutations, a recommendation that is based on studies by Yang et al. and Beitsch et al., which showed that there was no significant difference in the prevalence of pathogenic mutations in patients qualifying and not qualifying the NCCN criteria for testing.21,22 This suggests that almost 50% of mutations are missed if they are only tested by criteria-based testing. This was an important landmark in breast cancer genetics research as previously the NCCN guidelines, which are published and updated by expert groups, were universally accepted without having prospective validation. There are merits to testing all patients as systemic therapy can be tailored to mutation status,34,35,36 and risk-reducing interventions, including RRBSO and RRM, can be offered to patients and affected previvors.37,38 Although an argument can be made that testing more women leads to an increase in cost, more demand for genetic counsellors, and higher VUS detection rates, all these issues stem from not testing enough women in the first place.39,40,41 The cost has come down significantly over the past decade, with multiple studies showing the cost effectiveness of even a population-based testing strategy in high-income countries.42,43 VUS rates are also likely to decrease as more data become available from diverse populations. Recent studies have suggested that mainstreaming genetic testing and counseling by an oncologist is acceptable, and this strategy, often used in India and also in this study, may further decrease the need for certified genetic counsellors.23,44
Although patients qualifying the NCCN and MCG criteria had a higher likelihood of having a P/LP mutation in our study, the sensitivity of these criteria remain low and suggests that these criteria need modification in order to identify more patients for testing. In our study, a sensitivity analysis by age revealed that relaxing the age cut-off from the current 45 years in both the NCCN and MCG Plus criteria to 55 years could increase the sensitivity of these criteria to > 90% in our population, and testing all women up to the age of 60 years further improved the sensitivity to 97.7%. In the study by Yadav et al., a sensitivity cut-off of > 90% was reached at age 65 years, on which recommendations to consider testing in all women until 65 years of age, followed by risk stratification, were made in the recent NCCN guidelines.20 Although our sample size was small, a cut-off age of 60 years seems plausible in our setting in view of the younger age of onset of breast cancer in the Indian population. Testing women above the age of 60 years is unlikely to be of benefit as the prevalence of BRCA1/2 mutation and non-BRCA genes is low in this cohort (< 1%), as reported in a recent Women’s Health Initiative (WHI) study.45 None of the patients in our study aged > 65 years had a P/LP mutation, thus lending credence to these observations. Alternatively, incorporating the CANRISK tool for prediction of mutations at a cut-off of > 5%, as suggested by the recent NCCN guidelines, could improve the sensitivity of the testing criteria to 93.18%; however, this tool is cumbersome to use and utility in a busy routine practice is questionable.
An important aspect of genetic testing is the prevention of further cancers in patients and unaffected previvors. Although the NCCN guidelines offer recommendations for testing unaffected individuals, the sensitivity of these criteria for unaffected previvors has not been prospectively validated and are likely inadequate for the same. Despite the demonstrated benefits of RRBSO in terms of all-cause mortality and reduction in breast cancer incidence in multiple studies, the major focus of guidelines remains on identifying genetic susceptibility in patients after cancer diagnosis, rather than before the diagnosis.37,38,46 Although the uptake of risk-reducing interventions has increased over the past decade, with one study suggesting increased frequency of RRM in women diagnosed after 2009 than before,47 the willingness to undergo prophylactic surgeries among cancer survivors has varied widely, from 30 to 70% among studies, and depends on a number of ethnic and sociocultural factors besides cancer risk.47,48,49,50 Delay in elective surgeries due to the COVID-19 pandemic prevented a significant number of women in our cohort from undergoing preventive surgery (the current uptake of 37% for RRM and 29% for RRBSO would have been > 90% for both without the pandemic). None of the unaffected previvors were able to undergo preventive surgeries due to the pandemic. Uptake of cascade testing among first-degree relatives was low (10%), primarily due to the pandemic. A recent study by Offit et al. emphasized the importance of cascade testing among first-, second- and third-degree relatives of affected individuals in order to estimate the actual burden of hereditary cancer.51,52 However, a study from Singapore showed that even when free cascade testing was offered, only 29% of the first-degree relatives actually underwent testing, highlighting the need to extend counseling to unaffected previvors to meet this huge unmet need.53 Several factors, including hesitancy to reach out to health care providers and lack of communication on the part of the affected proband, contribute to this gap. Telephonic counseling is a potential solution to this problem, especially in a country such as India, and a recent study has shown the feasibility of such an approach.54
To summarize, this is the first prospective study from India to report a high prevalence of pathogenic mutations in a cohort of consecutive breast cancer patients. Testing by NGS using a broad multigene panel at a centralized CAP- and CLIA-certified laboratory for all patients with confirmation of a pathogenic variant by Sanger sequencing, followed by reflex MLPA for all patients negative by NGS, is a significant strength of this study. Pre- and post-counseling was performed by a medical oncologist in the study showing the feasibility of mainstreaming of genetic testing, which is an urgent need in a low middle-income country setting due to the shortage of trained cancer genetic counsellors.23,55 The relatively small sample size of the study, as well as its single-center nature, are important limitations of this study and larger prospective multi-institutional studies are needed to confirm these observations.
Conclusion
The prevalence of pathogenic mutations in breast cancer patients is higher in Indian patients compared with most other populations. AJ founder mutations are uncommon in the North Indian population and a strategy to screen for them before going on to full sequencing is unlikely to be beneficial. Although criteria-based testing still has a role in the current scenario, age criteria needs relaxation to 55–60 years to increase the sensitivity of the criteria to > 90% and expand the access to genetic testing. At the same time, there is an urgent need to improve post-test counseling and extend it to the unaffected previvors in order to improve the uptake of prophylactic interventions and cascade testing.
References
Bray F, Ferlay J, Soerjomataram I, Siegel RL, Torre LA, Jemal A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2018;68(6):394–424.
Mathur P, Sathishkumar K, Chaturvedi M, et al. Cancer statistics, 2020: report from national cancer registry programme, India. JCO Glob Oncol. 2020;6:1063–75. https://doi.org/10.1200/GO.20.00122.
Claus EB, Risch N, Thompson WD. Autosomal dominant inheritance of early-onset breast cancer. Implications for risk prediction. Cancer. 1994;73(3):643–51.
Peto J, Collins N, Barfoot R, et al. Prevalence of BRCA1 and BRCA2 gene mutations in patients with early-onset breast cancer. J Natl Cancer Inst. 1999;91(11):943–9.
King M-C, Marks JH, Mandell JB. New York breast cancer study group. Breast and ovarian cancer risks due to inherited mutations in BRCA1 and BRCA2. Science. 2003;302(564):643–6.
Sun J, Meng H, Yao L, Lv M, Bai J, Zhang J, et al. Germline mutations in cancer susceptibility genes in a large series of unselected breast cancer patients. Clin Cancer Res. 2017;23(20):6113–9.
Petrucelli N, Daly MB, Pal T. BRCA1- and BRCA2-associated hereditary breast and ovarian cancer. In: Adam MP, Ardinger HH, Pagon RA, Wallace SE, Bean LJ, Stephens K, et al. (eds). GeneReviews®. Seattle (WA): University of Washington, Seattle; 1993 (cited 13 Dec 2020). Available from http://www.ncbi.nlm.nih.gov/books/NBK1247/
Apostolou P, Fostira F. Hereditary breast cancer: the era of new susceptibility genes. Biomed Res Int. 2013;2013:747318.
Han M-R, Zheng W, Cai Q, Gao Y-T, Zheng Y, Bolla MK, et al. Evaluating genetic variants associated with breast cancer risk in high and moderate-penetrance genes in Asians. Carcinogenesis. 2017;38(5):511–8.
Sandhu DS, Sandhu S, Karwasra RK, Marwah S. Profile of breast cancer patients at a tertiary care hospital in north India. Indian J Cancer. 2010;47(1):16–22.
Sandhu GS, Erqou S, Patterson H, Mathew A. Prevalence of triple-negative breast cancer in India: systematic review and meta-analysis. J Glob Oncol. 2016;2(6):412–21.
Kulkarni A, Kelkar DA, Parikh N, Shashidhara LS, Koppiker CB, Kulkarni M. Meta-analysis of prevalence of triple-negative breast cancer and its clinical features at incidence in indian patients with breast cancer. JCO Global Oncol. 2020;6:1052–62.
Thaku KK, Bordoloi D, Kunnumakkara AB. Alarming burden of triple-negative breast cancer in India. Clin Breast Cancer. 2017;18(3):e393–9.
Engel C, Rhiem K, Hahnen E, Loibl S, Weber KE, Seiler S, et al. Prevalence of pathogenic BRCA1/2 germline mutations among 802 women with unilateral triple-negative breast cancer without family cancer history. BMC Cancer. 2018;18(1):265.
Chheda P, Pande S, Dama T, Vinarkar S, Chanekar M, Limaye S, et al. Spectrum of germline BRCA mutations in hereditary breast and ovarian cancer syndrome in Indian population: a central reference laboratory experience. Cancer Res Stat Treat. 2020;3(1):32.
Singh J, Thota N, Singh S, Padhi S, Mohan P, Deshwal S, et al. Screening of over 1000 Indian patients with breast and/or ovarian cancer with a multi-gene panel: prevalence of BRCA1/2 and non-BRCA mutations. Breast Cancer Res Treat. 2018;170(1):189–96.
Mannan AU, Singh J, Lakshmikeshava R, Thota N, Singh S, Sowmya TS, et al. Detection of high frequency of mutations in a breast and/or ovarian cancer cohort: implications of embracing a multi-gene panel in molecular diagnosis in India. J Hum Genet. 2016;61(6):515–22.
Soumittra N, Meenakumari B, Parija T, Sridevi V, Nancy KN, Swaminathan R, et al. Molecular genetics analysis of hereditary breast and ovarian cancer patients in India. Hered Cancer Clin Pract. 2009;7(1):13.
Darooei M, Poornima S, Salma BU, Iyer GR, Pujar AN, Annapurna S, et al. Pedigree and BRCA gene analysis in breast cancer patients to identify hereditary breast and ovarian cancer syndrome to prevent morbidity and mortality of disease in Indian population. Tumour Biol. 2017;39(2):1010428317694303.
Yadav S, Hu C, Hart SN, Boddicker N, Polley EC, Na J, et al. Evaluation of Germline genetic testing criteria in a hospital-based series of women with breast cancer. J Clin Oncol. 2020;38(13):1409–18.
Beitsch PD, Whitworth PW, Hughes K, Patel R, Rosen B, Compagnoni G, et al. Underdiagnosis of hereditary breast cancer: are genetic testing guidelines a tool or an obstacle? J Clin Oncol. 2019;37(6):453–60.
Yang S, Axilbund JE, O’Leary E, Michalski ST, Evans R, Lincoln SE, et al. Underdiagnosis of hereditary breast and ovarian cancer in medicare patients: genetic testing criteria miss the mark. Ann Surg Oncol. 2018;25(10):2925–31.
Kemp Z, Turnbull A, Yost S, Seal S, Mahamdallie S, Poyastro-Pearson E, et al. Evaluation of cancer-based criteria for use in mainstream BRCA1 and BRCA2 genetic testing in patients with breast cancer. JAMA Netw Open. 2019;2(5):e194428.
Armstrong N, Ryder S, Forbes C, Ross J, Quek RG. A systematic review of the international prevalence of BRCA mutation in breast cancer. Clin Epidemiol. 2019;11:543–61.
Tung N, Lin NU, Kidd J, Allen BA, Singh N, Wenstrup RJ, et al. Frequency of Germline mutations in 25 cancer susceptibility genes in a sequential series of patients with breast cancer. J Clin Oncol. 2016;34(13):1460–8.
Fackenthal JD, Zhang J, Zhang B, Zheng Y, Hagos F, Burrill DR, et al. High prevalence of BRCA1 and BRCA2 mutations in unselected Nigerian breast cancer patients. Int J Cancer. 2012;131(5):1114–23.
Vaidyanathan K, Lakhotia S, Ravishankar HM, Tabassum U, Mukherjee G, Somasundaram K. BRCA1 and BRCA2 germline mutation analysis among Indian women from south India: identification of four novel mutations and high-frequency occurrence of 185delAG mutation. J Biosci. 2009;34(3):415–22.
Hedau S, Jain N, Husain SA, Mandal AK, Ray G, Shahid M, et al. Novel germline mutations in breast cancer susceptibility genes BRCA1, BRCA2 and p53 gene in breast cancer patients from India. Breast Cancer Res Treat. 2004;88(2):177–86.
Hansa J, Kannan R, Ghosh SK. Screening of 185DelAG, 1014DelGT and 3889DelAG BRCA1 mutations in breast cancer patients from North-East India. Asian Pac J Cancer Prev. 2012;13(11):5871–4.
Saxena S, Chakraborty A, Kaushal M, Kotwal S, Bhatanager D, Mohil RS, et al. Contribution of germline BRCA1 and BRCA2 sequence alterations to breast cancer in Northern India. BMC Med Genet. 2006t;7:75.
Levy-Lahad E, Lahad A, King M-C. Precision medicine meets public health: population screening for BRCA1 and BRCA2. J Natl Cancer Inst. 2015;107(1):420.
Childers KK, Maggard-Gibbons M, Macinko J, Childers CP. National distribution of cancer genetic testing in the United States. JAMA Oncol. 2018;4(6):876–9.
Drohan B, Roche CA, Cusack JC, Hughes KS. Hereditary breast and ovarian cancer and other hereditary syndromes: using technology to identify carriers. Ann Surg Oncol. 2012;19(6):1732–7.
Robson ME, Tung N, Conte P, Im S-A, Senkus E, Xu B, et al. OlympiAD final overall survival and tolerability results: Olaparib versus chemotherapy treatment of physician’s choice in patients with a germline BRCA mutation and HER2-negative metastatic breast cancer. Ann Oncol. 2019;30(4):558–66.
Litton JK, Hurvitz SA, Mina LA, Rugo HS, Lee K-H, Gonçalves A, et al. Talazoparib versus chemotherapy in patients with germline BRCA1/2-mutated HER2-negative advanced breast cancer: final overall survival results from the EMBRACA trial. Ann Oncol. 2020;31(11):1526–35.
Tung NM, Robson ME, Ventz S, Santa-Maria CA, Nanda R, Marcom PK, et al. TBCRC 048: phase II study of olaparib for metastatic breast cancer and mutations in homologous recombination-related genes. J Clin Oncol. 2020;38(36):4274–82.
Marchetti C, De Felice F. Palaia I et al risk-reducing salpingo-oophorectomy: a meta-analysis on impact on ovarian cancer risk and all cause mortality in BRCA 1 and BRCA 2 mutation carriers. BMC Womens Health. 2014;14:150.
Rebbeck TR, Kauff ND, Domchek SM. Meta-analysis of risk reduction estimates associated with risk-reducing salpingo-oophorectomy in BRCA1 or BRCA2 mutation carriers. J Natl Cancer Inst. 2009;101(2):80–7.
Hoffman-Andrews L. The known unknown: the challenges of genetic variants of uncertain significance in clinical practice. J Law Biosci. 2017;4(3):648–57.
Stoll K, Kubendran S, Cohen SA. The past, present and future of service delivery in genetic counseling: keeping up in the era of precision medicine. Am J Med Genet C Semin Med Genet. 2018;178(1):24–37.
Milliron KJ, Griggs JJ. Advances in genetic testing in patients with breast cancer, high-quality decision making, and responsible resource allocation. J Clin Oncol. 2019;37(6):445–7.
Sun L, Brentnall A, Patel S, Buist DSM, Bowles EJA, Evans DGR, et al. A cost-effectiveness analysis of multigene testing for all patients with breast cancer. JAMA Oncol. 2019;5(12):1718–30.
Manchanda R, Patel S, Gordeev VS, Antoniou AC, Smith S, Lee A, et al. Cost-effectiveness of population-based BRCA1, BRCA2, RAD51C, RAD51D, BRIP1, PALB2 mutation testing in unselected general population women. J Natl Cancer Inst. 2018;110(7):714–25.
Richardson M, Min HJ, Hong Q, Compton K, Mung SW, Lohn Z, et al. Oncology clinic-based hereditary cancer genetic testing in a population-based health care system. Cancers (Basel). 2020;12(2):E338.
Kurian AW, Bernhisel R, Larson K, Caswell-Jin JL, Shadyab AH, Ochs-Balcom H, et al. prevalence of pathogenic variants in cancer susceptibility genes among women with postmenopausal breast cancer. JAMA. 2020;323(10):995–7.
Carbine NE, Lostumbo L, Wallace J, Ko H. Risk-reducing mastectomy for the prevention of primary breast cancer. Cochrane Database Syst Rev. 2018;4:CD002748.
Metcalfe K, Eisen A, Senter L, Armel S, Bordeleau L, Meschino WS, et al. International trends in the uptake of cancer risk reduction strategies in women with a BRCA1 or BRCA2 mutation. Br J Cancer. 2019;121(1):15–21.
Long J, Evans TG, Bailey D, Lewis MH, Gower-Thomas K, Murray A. Uptake of risk-reducing surgery in BRCA gene carriers in Wales, UK. Breast J. 2018;24(4):580–5.
Beattie MS, Crawford B, Lin F, Vittinghoff E, Ziegler J. Uptake, time course, and predictors of risk-reducing surgeries in BRCA carriers. Genet Test Mol Biomark. 2009;13(1):51–6.
Evans DGR, Lalloo F, Ashcroft L, Shenton A, Clancy T, Baildam AD, et al. Uptake of risk-reducing surgery in unaffected women at high risk of breast and ovarian cancer is risk, age, and time dependent. Cancer Epidemiol Biomark Prev. 2009;18(8):2318–24.
Kurian AW, Katz SJ. Emerging opportunity of cascade genetic testing for population-wide cancer prevention and control. J Clin Oncol. 2020;38(13):1371–4.
Offit K, Tkachuk KA, Stadler ZK, Walsh MF, Diaz-Zabala H, Levin JD, et al. Cascading after peridiagnostic cancer genetic testing: an alternative to population-based screening. J Clin Oncol. 2020;38(13):1398–408.
Courtney E, Chok AK-L, Ting Ang ZL, Shaw T, Li S-T, Yuen J, et al. Impact of free cancer predisposition cascade genetic testing on uptake in Singapore. NPJ Genom Med. 2019;4(1):1–7.
Caswell-Jin JL, Zimmer AD, Stedden W, Kingham KE, Zhou AY, Kurian AW. Cascade genetic testing of relatives for hereditary cancer risk: results of an online initiative. J Natl Cancer Inst. 2018;111(1):95–8.
Verma A, Nag S, Hasan Q, Selvakumar VPP. Mainstreaming genetic counseling for BRCA testing into oncology clinics – Indian perspective. Indian J Cancer. 2019;56(5):38.
Acknowledgment
The authors would like to thank Tata Center of Development, University of Chicago, for funding this study.
Funding
This study was supported by the TATA Trusts at the University of Chicago, New Delhi Center.
Author information
Authors and Affiliations
Contributions
RP, OO, AM, SVSD, AG, AB, AK, SB, KSD, ST, and ED contributed to the concept design and conduct of the study; RP and AM conducted the genetic counseling; AM and RP performed the statistical analysis; and RP and AM drafted the manuscript. All authors edited the manuscript.
Corresponding author
Ethics declarations
Disclosure
Abhenil Mittal, SVS Deo, Ajay Gogia, Atul Batra, Akash Kumar, Sandeep Bhoriwal, Koushik Sinha Deb, Ekta Dhamija, Sanjay Thulkar, VL Ramprasad, Olufunmilayo Olopade, and Raja Pramanik declare no potential conflicts of interest.
Consent to Participate
Informed consent was obtained from all patients.
Data Availability
Data regarding this study will be available from the corresponding author (RP) upon reasonable request.
Ethical Approval
The study protocol was approved by the IEC, vide letter number IECPG-35/23.01.2019
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Mittal, A., Deo, S.V.S., Gogia, A. et al. Profile of Pathogenic Mutations and Evaluation of Germline Genetic Testing Criteria in Consecutive Breast Cancer Patients Treated at a North Indian Tertiary Care Center. Ann Surg Oncol 29, 1423–1432 (2022). https://doi.org/10.1245/s10434-021-10870-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1245/s10434-021-10870-w