Abstract
The identification and removal of host cell proteins (HCPs) from biologic products is a critical step in drug development. Despite recent improvements to purification processes, biologics such as monoclonal antibodies, enzyme replacement therapies, and vaccines that are manufactured in a range of cell lines and purified using diverse processes may contain HCP impurities, making it necessary for developers to identify and quantify impurities during process development for each drug product. HCPs that contain sequences that are less conserved with human homologs may be more immunogenic than those that are more conserved. We have developed a computational tool, ISPRI-HCP, that estimates the immunogenic potential of HCP sequences by evaluating and quantifying T cell epitope density and relative conservation with similar T cell epitopes in the human proteome. Here we describe several case studies that support the use of this method for classifying candidate HCP impurities according to their immunogenicity risk.
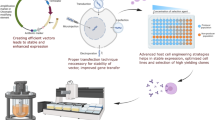
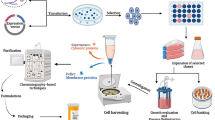
Graphical Abstract

Similar content being viewed by others
Introduction
More than 600 biologics of many different types have been licensed by the FDA for human use, with a market value of approximately US$ 365 billion. This market value is expected to expand to more than US$ 700 billion by 2030 (1, 2). In fact, FDA approvals of biologics such as monoclonal antibodies, enzyme replacement therapies, and vaccines exceeded those of small molecules for the first time in 2022, following decades of investment by biopharma companies (3). This growth in the biologics sector highlights the significant contributions this sector is making towards improving human and animal health. The number of manufacturers of biologics continues to expand as the earliest biologic therapeutics come off patent and biosimilar products enter the approval process.
Occasionally, protein impurities derived from cell culture systems can present a challenge for therapeutic development. The types of host cells used to produce biologics include bacterial cells like Escherichia coli, yeast cells such as Saccharomyces cerevisiae, and mammalian cells, including Chinese Hamster Ovary (CHO) cells and human embryonic kidney (HEK) cells. These cells are often chosen for their capacity to be genetically manipulated to express the desired therapeutic. In addition to expressing the desired product, host cells make a wide range of other proteins that may be difficult to separate from the product during processing and formulation. While many of these proteins may be innocuous, some host cell protein impurities, or HCPs (also known as PRPIs, or Process Related Protein Impurities), can have an adverse effect on the safety and efficacy of biologic products (4,5,6,7). As a result, efforts have been made to improve methods for identification, removal, and immunogenicity assessment of HCPs (8,9,10,11).
In response to the above concerns about HCPs in biologic products, a range of analytical methods are used to identify and quantify HCPs during biologic development (See references (12,13,14) for reviews). For example, ELISA kits for identifying HCPs are sold for commercial use or developed internally for HCP detection (15, 16). These tests are high throughput, and relatively sensitive, although some HCP are missed due to similarities with the host that is the source of the antibody panel. Anti-HCP antibodies can also be used to identify HCP in two-dimensional (2D) gels in which HCP are separated by isoelectric point and molecular weight. Once separated, proteins can be identified by mass spectrometry. These methods provide some indication of the volume and diversity of HCPs that may be present in a biologic product, but they may fail to fully identify all the potential HCPs in that given product, and they also do not directly facilitate the identification and selective removal of potentially immunogenic HCPs.
More recently, drug developers have turned to identification and evaluation of individual impurities by liquid chromatography-tandem mass spectrometry (LC–MS/MS), which has the additional benefit of making it possible to determine the identity of impurities that are present in biologic products even at low concentrations (12, 17). Moving from 3D western blot to LCMS has improved the potential for drug developers to identify and de-risk HCP that have high immunogenic potential.
Sequence-based immunogenicity risk assessment is then possible, and those HCPs that represent the highest risk can be the focus of process improvement efforts for impurity reduction. These methods are now well established in the biologics industry. In contrast, methods for assessing the immunogenicity risk of HCPs have not been very well defined, primarily due to the similarity of HCPs to human proteins, which may make it difficult to comprehend their potential for immunogenicity risk. Additional risk assessment and validation studies are needed to better quantify the immunogenicity risk of individual HCPs and to define thresholds that correlate with immunogenicity in the clinic.
Here we describe a computational approach for performing immunogenicity risk assessment of HCPs and PRPI. The method uses commercial tools that have been prospectively and retrospectively validated in biologics (see (18,19,20,21)) and the description of the method here would enable researchers to use similar tools to obtain proximal results (Supplemental Table S1). We expect that computational analyses of the T cell epitopes, as described, will provide a starting point for later validation studies performed in vitro, or post approval, in immune-monitoring studies.
Immunoinformatics Assessment of HCPs Using ISPRI-HCP
In 2017, EpiVax programmers developed a toolkit comprised of several integrated algorithms for immunogenicity screening of host cell proteins, known as ISPRI-HCP (Interactive Screening and Protein Reengineering Interface for Host Cell Proteins). ISPRI-HCP is accessible through a secure web-based interface (22, 23). ISPRI-HCP uses an established set of tools initially developed for the ISPRI website (24) for immunogenicity risk assessment of biologics that is currently used by many small and large biopharma companies to screen and re-engineer candidates in their biologics pipelines (25,26,27,28,29,30,31,32).
As will be illustrated in greater detail, ISPRI-HCP performs immunogenicity risk assessments of a host cell protein, using internal databases of HCP genomes obtained from GenBank or other sources (33,34,35,36,37). ISPRI-HCP estimates the immunogenic potential of protein sequences by evaluating their class II-restricted T cell epitope density, and the relative conservation of these epitopes with other, similar epitopes in the human genome, using the tools EpiMatrix, ClustiMer and JanusMatrix. We used ISPRI-HCP to classify a set of common HCP impurities according to their immunogenicity risk. These risk assessment thresholds generally correspond with clinical reports when those data are available. A brief description of the types of algorithms that are combined to generate the ISPRI-HCP immunogenicity risk assessment, including EpiMatrix, ClustiMer and JanusMatrix, is provided below.
The ISPRI-HCP web platform employs EpiMatrix, an epitope mapping tool that parses protein sequences into overlapping nine-mers (nine overlapping by eight) and evaluates each of the peptides for potential to bind to HLA DRB1 molecules. HLA DRB1 is selected for HCP analysis because it is the most common HLA expressed on antigen presenting cells and is associated with the development of anti-drug and autoantibodies, which are the main concern for HCP and for biologics in general (38, 39).
While there are hundreds of HLA DR alleles in the human population, the EpiMatrix tool reduces the complexity of the HLA landscape by focusing on nine “supertypes” that share common HLA DR binding pockets, covering the binding propensities of HLA DR alleles representing greater than 95% of the world’s population (40, 41). EpiMatrix is used in combination with the ClustiMer algorithm to find epitope-dense regions in protein sequences. Of key importance, HLA DR-restricted T cell epitopes are not randomly distributed within a protein, but instead tend to cluster in defined segments across the supertype alleles. ClustiMer helps define these epitope dense regions, and often highlights a common feature in immunogenic proteins known as an EpiBar (or epitope-bar) in which a given frame within a peptide is predicted to bind to four or more HLA supertype alleles (42). These regions of epitope density appear to be promiscuous epitopes (have a broad HLA-binding and immunogenic propensity) when tested in vitro (43). EpiBars have been shown to induce an immune response in a greater portion of any study cohort than regions that do not contain such density (Fig. 1) (23).
Example of a promiscuous epitope in PLBL2. The 15-mer peptide shown is derived from CHO Phospholipase B-like 2 (PLBL2), a host cell protein known to be immunogenic in humans. Z scores indicate the potential of each 9-mer frame to bind to a given HLA allele supertype; the strength of the score is indicated by the blue shading. All scores in the Top 5% (Z-Score ≥ 1.64) are considered “Hits” (highlighted in dark and medium blue). Scores in the top 10% are considered elevated, but not significant (light blue). Frames containing four or more alleles scoring above 1.64 are referred to as EpiBars (highlighted in yellow). The EpiBar highlighted in yellow on the left and right is a feature that is characteristic of a promiscuous T cell epitope. The PLBL2 promiscuous epitope sequence (IIKLLPGAR) has seven hits with scores in the top 5% for 6/9 and the top 1% for 1/9 HLA supertypes
Identification of HLA DR binding epitopes alone is not sufficient to determine the immunogenicity of peptides derived from therapeutic and host cell proteins. One important reason is that for HCPs derived from CHO or other mammalian cell lines such as HEK cells, the HCP epitopes may be very similar (or identical) to the epitopes found in the human genome, to which human beings may be tolerant, due to exposure of these epitopes in the T cell receptor selection process in the thymus (44). For that reason, ISPRI-HCP uses an additional algorithm, the JanusMatrix tool, which evaluates the homology of HLA-binding T cell epitope clusters identified in HCPs (e.g., from CHO) to epitopes with similar HLA-DR-restricted epitopes that are present in the human proteome (23, 42). JanusMatrix separates the amino acid sequence of T cell epitopes into TCR-facing residues and HLA binding cleft-facing residues, and then compares the TCR face to other putative T cell epitopes. Additional information on the EpiMatrix, ClustiMer and JanusMatrix tools can be found in Supplemental Methods S1.
The idea driving this comparison between HCPs and human proteins is that HCP-derived epitopes that are restricted by a specific HLA DR for which there are T cell receptor (TCR) facing residues that are conserved with human protein-derived epitopes that bind to the same HLA are likely to be tolerated by the human immune system, and thus are less likely to cause deleterious immune responses. This may be true even if the sequence of the HLA-binding face of the HCP is non-identical, presuming it still binds to the same HLA. JanusMatrix also defines the extent of cross conservation between epitope clusters and the human proteome based on the number of different human proteins that contain TCR-matching epitopes. TCR Epitopes derived from human proteins and non-human proteins from some bacterial and viral proteins, that have extensive cross-conservation have been shown to be tolerogenic, in vitro and in vivo, in studies measuring bystander suppression of T effector responses and cell surface markers (31, 32, 45).
Here, we will demonstrate the utility of this type of analysis for use with HCP found in biosimilar products and vaccines. Previously, EpiMatrix, ClustiMer and JanusMatrix have been used to evaluate the immunogenicity of biologics such as vaccine epitopes(23, 46), epitopes from monoclonal antibodies (11) and enzyme replacement therapies (47), and epitopes found in synthetic peptide generics including incretin mimetic peptides (e.g., semaglutide, liraglutide (48)) to ensure that the potential immunogenicity of the generic peptide drugs does not differ from that of the reference listed drug (RLD) (49). Both in silico and in vitro methods are used to evaluate the potential immunogenicity of generics and their peptide impurities (20, 49, 50).
Safety Risks of Immunogenic Host Cell Protein (HCP) Impurities
HCP immunogenicity may be a significant safety risk factor for drug products produced using cell culture systems. HCPs may cause adverse reactions in patients the nature of which could be hypersensitivity responses, generation of anti-drug antibodies with consequent abrogation of product efficacy, or, cross reactive neutralization of an endogenous protein homologue, especially those with non-redundant function. Even at low levels, HCPs may induce a detrimental immune response, contributing to the overall immunogenicity of the product (51). Detection of anti-host cell protein antibodies following exposure to therapeutic products has resulted in the suspension of advanced clinical trials (52, 53). Antibodies against HCP have the potential to cross-react with endogenous protein homologs, which may result in serious adverse events. While there are only few instances of safety risk to patients being attributed to HCP impurities, there is concern about high immunogenicity risk impurities as indicated by many recent FDA guidelines and other publications (11, 54,55,56). Relevant to the development of biosimilars, new HCP impurities that were not present in the RLD drug may result in a different immunogenicity profile and modify the overall safety profile of the biosimilar drug.
The significance of HCPs is illustrated by reports of unwanted immunogenicity and abrogated development of therapeutic proteins due to immunogenicity of residual HCPs present in monoclonal antibody and recombinant protein products manufactured in the most frequently used expression system for production of biologics and biosimilars, that of CHO cells (9,10,11,12,13). However, as reported in numerous articles, there are no reported examples wherein immunogenicity of CHO HCPs caused significant cross reactive immune responses to endogenous human protein homologs and produced clinically adverse effects. A framework for biologics development, highlighting key considerations in the risk assessment of HCPs, was published in 2015 (de Zafra et al., Biotech. Bioeng. 112: 2284–2291) (57). The most glaring example of robust immunogenicity to CHO HCP is the response in patients to CHO Phospholipase B Like 2 (PLBL2) which was found to be recognized by IgG4 mAbs by several companies and for which anti-PLBL2 antibodies were detected in treated patients (58). In theory, the greatest safety concerns related to this robust response is the potential for cross reactivity and neutralization of the activity of the endogenous human homolog, and the potential for the response to CHO PLBL2 to act as an adjuvant to generate immune responses to the therapeutic protein. Fortunately, no correlation was found between anti-PLBL2 antibodies and clinical adverse events. Another historical example of HCPs inducing immune responses to the therapeutic was in the production of human growth hormone (Somatropin) in an E. coli culture system. In clinical trials, approximately 60% of the patients receiving hGH that contained E. coli HCP impurities developed antibodies to both hGH and E. coli proteins, implicating E. coli HCPs as adjuvants (4). This response was especially worrisome in that immune response to the human HGH product had the potential to cross reactively neutralize endogenous HGH. Fortunately, the anti-drug antibodies (ADAs) were not cross-neutralizing in this instance.
Recent examples of HCPs found in therapeutics that had immunogenic activities of concern include the following: 1) clinical trials of a CHO cell produced Factor IX were halted after 26% of clinical trial subjects were discovered to have developed anti-HCP antibodies although, no adverse events related to the development of anti-HCP antibodies were reported and no relationship between the development of anti-HCP antibodies and the development of ADAs was observed (52, 53); 2) MCP-1 copurified with CHO-produced CTLA4-Ig and produced clinical adverse events due to histamine release, with a consequent clinical hold placed on the study (4); 3) TGFb1 copurified in a study of CHO-produced MUC1-Fc but its effects, if any were unknown (4). One factor worthy of note for several of these products regards the nature of the HCPs typically found: the majority of HCP are intracellular proteins (Fig. 2); thus, cross-reactive anti-drug antibodies would not necessarily reach the human homolog, minimizing potential adverse effects related to anti-homolog-antibodies.
The composition of extracellular versus intracellular CHO HCP impurities. Using the Gene Ontology (GO) knowledgebase, we retrieved the cellular component annotation for a compiled list of commonly found CHO HCP impurities. Shown is the percentage of extracellular (n = 21) versus intracellular (n = 89) proteins
Indeed, this latter point is captured in a 2015 risk assessment framework of HCPs, which includes additional factors to consider such as the HCP homology to human proteins, frequency and duration of treatment, route of administration, known biologically activity, and population age (57). The concentration of individual HCPs is also a critical risk factor. Many drug products may contain immunogenic HCPs present at low levels resulting in no measured response. The general guidelines for the maximum amount of allowed HCP impurities is 100 ng per mg of product (10). Nevertheless, the acceptable limit for HCP impurities in the final product should be determined on a case by case basis. This is due to the great diversity of production and purification systems used by pharmaceutical companies (57).
Immune System Processes Leading to HCP Immunogenicity
The immunogenicity of HCPs is routinely assessed by measuring antibodies against them, as well as antibody titer and isotype, which are attributes that impact therapeutic safety. Generation of antibodies to HCPs depends on uptake of HCP by antigen presenting cells, processing via cellular enzymatic machinery, and presentation of peptides derived from the protein in the context of HLA class II on antigen presenting cells that activate follicular helper T (Tfh) cells. Tfh cells provide help to B cells that make antibodies specific to epitopes in the HCP. Tfh cells induce the B cells to class switch (from IgM to IgG) and drive recombination, affinity maturation, and B cell differentiation into long-lived antibody-secreting plasma cells and memory cells (59, 60). Without T cell epitopes activating Tfh cells, B cell response remains weak and transient. For example, antigens that have high concentrations of T cell epitopes drive high affinity, long lasting B cell responses (e.g., Hepatitis B surface antigen), and antigens that have low T cell epitope concentrations (per unit length) only drive weak antibody responses (e.g., circumsporozoite protein from falciparum Malaria, and proteins such as albumin) (61).
However, presentation of biologic drug-derived T cell epitopes to T cells specific for the peptide-HLA complex does not always lead to generation of a strong immune response. In fact, presentation of T cell epitopes derived from biologic molecules can have quite diverse outcomes depending on whether the peptide produced is entirely foreign to the immune system, for which an immune response can be expected, or whether it has similarities to human proteins, which can lead to anergy (lack of response) or may even trigger a “regulatory” response to diminish immunogenicity. T cell epitopes, that appear to serve a regulatory function, were found in human pathogens and may have evolved to supress T cell response thereby protecting the pathogen from the induction of high titer, high affinity antibody responses (31, 62). It follows that the immunogenicity of any given host cell protein may be mitigated by the fact that its T cell epitopes appear similar at the TCR face to epitopes found in human proteins. This concept is likely to be familiar to scientists who have tried to developed antibodies to HCP proteins and found that some HCP do not effectively generate immune responses when they are used to immunize other mammalian species such as rodents, even when strong adjuvants are used (63, 64).
It is important to note that antibodies to HCPs can be observed in clinical studies, consistent with the observation that imperfect homology between animal model endogenous HCP and production line HCP contributes to the development of anti-HCP antibodies (63, 65). Increased immune response against the non-homologous epitope may lead to activation of T cells that recognize epitopes in the therapeutic drug, even if they are host-like (64). And while, “any protein is potentially immunogenic” (66) proteins that contain a large number of foreign-appearing T cell epitopes are even more so, particularly when delivered by a route that is not natural (intravenous, intramuscular), at a dose that is not consistent with natural expression, and in conjunction with innate immune response modifiers (20, 67). Cases of severe adverse immune responses to autologous proteins have been published (68, 69). Below, we give several examples of HCPs evaluated by ISPRI-HCP and illustrate the utility of the computational approach to HCP immunogenicity risk assessment.
ISPRI-HCP—Application Case Studies
To perform a host cell protein immunogenicity analysis, individual proteins (whether host-cell derived or from any other source) are scored for potential HLA DR ligands using EpiMatrix (70) (or this can be performed using other available epitope prediction tools, see for example, the list in Supplemental Table S1). In ISPRI-HCP, the density of ligands in the complete HCP protein sequence is expressed on a scale as an EpiMatrix Protein Score, which represents the difference between the number of putative HLA DR-restricted epitopes predicted by EpiMatrix in a given HCP sequence, normalized for length (per 1,000 nine-mer to allele assessments), and the median score (set as zero) for a set of random proteins (42).
The range of scores can be illustrated on a vertical or horizontal scale, in which the median represents the normalized EpiMatrix Score (HLA DR-restricted epitopes normalized for length) of 10,000 randomly generated protein sequences. This normalized scale enables the sorting of proteins into those that score higher or lower than the identified ‘random’ standard. In general, highly immunogenic antigens score twenty points higher than random proteins on this normalized scale, and non-immunogenic proteins score below zero. It is also interesting to note that the median score of proteins contained in the whole human proteome is minus nine (-9) on this scale, and the human secretome (which contains proteins such as albumin and immunoglobulin) scores below minus twenty (-20) on this scale. The median score for intracellular proteins is minus twelve (-12) (42).
As described above, HLA-DR-restricted epitopes tend to cluster. These regions of epitope density are uncovered using a tool called ClustiMer in the ISPRI toolbox and other epitope clustering algorithms. Once regions of epitope density have been identified using ClustiMer, the TCR-facing residues can be examined with JanusMatrix, or another similar tool, to identify putative T cell epitopes that are cross-conserved at the TCR face with similar epitopes in the human proteome. JanusMatrix assembles this information into an overall Human Homology Score which indicates the average depth of conservation with epitopes contained within proteins found in the human proteome, for each of the HLA-binding peptides contained in the source sequence. A high JanusMatrix Human Homology Score suggests greater conservation with the human genome, which may bias the immune response towards immune tolerance (due to extensive similarity with human proteins). Based on numerous application case studies, thresholds have been established for JanusMatrix that differentiate between more foreign, or potentially “immunogenic” epitopes, and human-like epitopes that are likely tolerated or potentially “tolerogenic” (23, 42, 71).
The relative immunogenicity risks for HCP impurities can be defined by graphing the EpiMatrix and JanusMatrix scores of each HCP on a 2D dot plot. The 2D plot is then divided into four quadrants using the EpiMatrix Score threshold of twenty (y = 20) and JanusMatrix Score threshold of three (x = 3). The immunogenicity risk classification is then determined by the HCP location within the four quadrants. HCPs (and proteins from other sources) with EpiMatrix Scores greater than 20 and JanusMatrix Scores less than 3 are classified as having higher risk of generating an immune response in a human host, and HCPs with EpiMatrix Scores less than 20 and JanusMatrix Scores greater than 3 are classified as lower risk proteins. It is not uncommon for HCPs to have many identical T cell epitopes within the human genome; however, known immunogenic HCPs generally have more non-human epitopes, on average, than HCPs that have not been reported to be immunogenic. In general, data on the immunogenicity of HCPs is limited (most developers do not publish their HCP data). When HCP immunogenicity data becomes more available, it will be important to identify “benchmark” proteins such as the ones identified in the case studies below.
Case Study #1—Immunogenicity Risk Classification of Commonly Found CHO Host Cell Protein Impurities
In preliminary work, ISPRI-HCP has been used to evaluate the immunogenic potential of the 143 CHO HCPs that are frequently found to co-purify with mAbs (9, 72). Shown in Fig. 3 is a subset of these proteins plotted on the Y axis by their EpiMatrix Protein Score and on the X axis by their JanusMatrix Human Homology Score. Several of these commonly identified HCPs with their EpiMatrix and JanusMatrix Scores are illustrated on the accompanying Quadrant plot (Fig. 3). The bubble plot shows each HCP in a quadrant that is used for classifying their immunogenicity risk based on EpiMatrix (EMX) and JanusMatrix (JMX) thresholds. We find that the predicted immunogenic potential of CHO HCPs covers a wide range of scores, both in terms of epitope content and “human-ness” as defined by JanusMatrix. We are currently performing in-vitro validation studies of ISPRI-HCP using several of the CHO proteins that are identified by name in Fig. 3 (PLBL2, CTSA, RAN, LPLA2, PLD3, and NUCB2) and their scores are listed in Supplemental Table S2.
ISPRI-HCP analysis of a typical CHO HCP Landscape. Using a list of commonly found CHO protein impurities, we calculated their EpiMatrix (EMX) and JanusMatrix (JMX) scores for each protein. Shown here are 143 CHO HCP impurities. Proteins are shown on a gradient scale from high (orange), medium (yellow), and low (green) immunogenicity. Proteins with EMX greater 20 and JMX scores less than 3 are predicted to be high risk (Q II). Proteins with EMX less than 20 and JMX greater than 3 are predicted to be low risk (Q IV). Proteins sourced from: Jones, M. et al. “High-risk” host cell proteins (HCPs): a multi-company collaborative view. Biotechnology and Bioengineering vol. 118 2870–2885 (2021); Molden et al. (2021), Host cell protein profiling of commercial therapeutic protein drugs as a benchmark for monoclonal antibody-based therapeutic protein development
It is important to note this plot does not show a third dimension that could impact immunogenicity, which is the prevalence or concentration of the HCP in the drug product. When that information is available, the size of the dot on this bubble chart can reflect relative concentration compared to other HCPs in the product, enabling drug developers to visualize this risk factor for further investigation.
This retrospective analysis of publications on CHO protein impurities suggests that the initial immunogenicity predictions made by combining the EpiMatrix and JanusMatrix Scores as above, and as calculated by ISPRI-HCP, are relatively accurate for some proteins for which clinical data are available.
For example, in a study performed by Genentech researchers, a “hitchhiker” protein identified as PLBL2 was confirmed to be immunogenic in clinical studies: ~ 90% of individuals receiving the mAb lebrikizumab containing PLBL2 developed anti-PLBL2 antibodies against this CHO protein impurity. In addition, a dose dependent production of anti-PLBL2 antibodies was observed eight weeks after the final dose. Fortunately, no association between drug safety and anti-PLBL2 antibodies was found in clinical studies, and anti-PLBL2 antibodies did not appear to contribute to induction of anti-lebrikizumab antibodies, in this instance (58).
Case Study #2—Assessment of Vaccine HCPs and Process-Related Protein Impurities
Immunogenicity of HCPs is not confined to biologic products produced in foreign cell culture systems (e.g., E. coli, CHO). HCPs can also be derived from human cell lines, such as human T-REx-293 cells that are used to produce viral-vectored vaccines. While the presence of many proteins in vaccine products is fairly common, since many vaccine products are produced in bacterial culture or in cell lines and consequently have PRPIs, and there is no well-defined limit for these impurities. The induction of immune responses to PRPIs may lead to unexpected adverse effects. For example, in the context of COVID-19 vaccination, rare cases of Vaccine-induced Thrombotic Thrombocytopenia (VTT) were observed for patients who received cell-culture derived COVID-19 vaccines developed by Oxford/AstraZeneca and Janssen (Johnson & Johnson). The condition was more common among young women than older individuals and men (73, 74). Usage of some of these COVID-19 vaccines was suspended in the US and their use is limited in other countries due to concern about VTT (75, 76). While the cause of VTT remains unknown, selected PRPIs in the Oxford/AstraZeneca chimpanzee adenovirus (ChAdOx1) that encodes the SARS-CoV-2 (nCoV-19) vaccine were subsequently identified in these vaccines, and some researchers have suggested that these PRPIs may have contributed to the unwanted autoimmune reactivity (77, 78), although additional studies may be required to confirm this.
Several research groups have published studies describing and identifying PRPIs that are present in clinical samples of the adenoviral vectored vaccines used for COVID prevention. These vaccine-associated PRPIs may be a mixture of cell-culture derived proteins and viral proteins. In the case of the Oxford/AstraZeneca vaccine, 29–55% of PRPIs were adenoviral proteins, with hexon being among the most abundant PRPIs (78). Furthermore, adenoviral hexon protein has been observed to form complexes with platelet factor 4 (PF4) and to be associated with the production of anti-PF4 antibodies that lead to VTT. In addition to anti-PF4 antibody production, PRPIs may also have an impact on the proinflammatory responses seen in VTT (73, 77).
Using ISPRI-HCP, we performed an in-silico immunogenicity risk assessment of the Oxford/AstraZeneca ChAdOx1 nCoV-19 vaccine protein impurities that were identified by Krutzke et al. (2021) (Fig. 4) (78). The PRPIs are derived from T-REx-293 cells, a derivative of human HEK293 cells and ChadOx1 composed of adenoviral proteins (79). A list of these PRPIs with their EpiMatrix and JanusMatrix Scores are found in Supplemental Table S3. The human PRPIs are located in QIII and QIV with EpiMatrix Scores less than twenty and JanusMatrix Scores ranging from 1.86 to 10.84. This classifies the human PRPIs as having a low to moderate immunogenicity risk. Unlike the human PRPIs, the adenoviral proteins are primarily clustered in a single quadrant (QIII) with EpiMatrix Scores less than twenty and JanusMatrix Scores less than three. These proteins are classified as moderate risk because they contain foreign epitopes and low T cell epitope content.
ISPRI-HCP analysis of the AstraZeneca ChadOx1 nCoV-19 vaccine protein impurities identified and published by Krutzke et al. (2021). Starting from a published list of protein impurities found in the AstraZeneca ChadOx1 nCoV-19 vaccine, we calculated their EpiMatrix and JanusMatrix Scores. A dot plot is shown for the adenoviral protein impurities (n = 12) derived from the chimpanzee adenovirus (ChadOx1) (Left graphic) (a) and human HCP impurities (n = 18) derived from (Human cell-derived) T-Rex-293 host cells (Right graphic) (b). Proteins are shown on a gradient scale from high (orange), medium (yellow), and low (green) immunogenicity. Proteins with EMX greater 20 and JMX scores less than 3 are predicted to be high risk (Q II). Proteins with EMX less than 20 and JMX greater than 3 are predicted to be low risk (Q IV). Benchmark proteins are shown as open circles
Out of all the PRPI assessed, adenoviral proteins pIIIA and pVII had the highest immunogenicity (EpiMatrix) Scores. Furthermore, adenoviral proteins, including hexon and DNA-BP, had the lowest predicted JanusMatrix Scores, reflecting low ‘human-ness’ and low likelihood of tolerance (Fig. 4). Using the same data, we assessed the difference in T cell epitope density and tolerance for the human versus adenoviral derived protein impurities (Fig. 5). The JanusMatrix Scores for human PRPI were higher when compared to the adenoviral PRPI. These differences were statistically significant when compared to the human protein impurities (Fig. 5b).
Comparison of T cell epitope density and tolerance of PRPI found in the AstraZeneca ChadOx1 nCoV-19 vaccine published by Krutzke et al. (2021). Starting from a published list of protein impurities found in the AstraZeneca ChadOx1 nCoV-19 vaccine, we calculated their EpiMatrix and JanusMatrix Scores. EpiMatrix Scores for human HCPs derived from T-Rex-293 host cells and adenoviral proteins derived from chimpanzee adenovirus (ChadOx1) are shown (a). JanusMatrix Scores for human HCPs derived from T-Rex-293 host cells and adenoviral proteins derived from chimpanzee adenovirus (ChadOx1) are shown (b). Proteins with an EpiMatrix Scores below 20 (dotted line) have a low epitope density. Proteins with a JanusMatrix Score less than 3 (dotted line) are likely to contain epitopes that are not tolerated (red dots). Bars represent the median for all data points, and error bars indicate the interquartile range. ****, P ≤ .0001, Mann–Whitney U test
Altogether these results highlight the potential for adenoviral-vector-derived PRPIs to activate T cells. The relatively low immunogenic potential of the mammalian cell culture proteins in the vaccine, contrasts with the higher potential for adenoviral PRPIs to induce unwanted or unexpected immune responses. However, the response to the adenoviral PRPIs, in conjunction with additional immune stimulation from vaccine adjuvants could theoretically contribute to a break in tolerance to the mammalian cell-derived HCPs for which immune tolerance is less robust.
Relevance to Biosimilar and Biologic Product Development
HCP analysis is particularly relevant to the development of biosimilar drugs. Since differences in the HCP content of biosimilar and reference listed drug (RLD) may change the immunogenicity profile and hinder their designation as “interchangeable products,” the FDA may be interested in evaluation of HCP and comparisons between originator HCP and biosimilar HCP content, to improve and expedite the development of fully interchangeable products.
The FDA is currently exploring whether it is possible to predict differences in immunogenicity using in silico and in vitro methodologies. As is stated by the FDA, “improving the efficiency of biosimilar development also includes... comparative immunogenicity assessment using in silico and in vitro methodologies” (80). Improving predictions of immunogenicity could improve the efficiency of biosimilar development, “and streamline the clinical data needed to support no clinically meaningful differences in immunogenicity between a proposed biosimilar product and the reference product” (80). More accurate immunogenicity estimates for PRPIs such as HCPs in biosimilar products may also reduce the duration of any clinical studies that may be required for a drug product (79).
It is widely believed that the amount and type of HCPs found in a therapeutic product can be shaped by the downstream purification process (81,82,83). It may be difficult for biosimilar developers to assure HCP comparability to an RLD due to differences in cell culture conditions. This is the case even in the likely scenario where the same host cell is used for manufacture, as HCP mixtures can be complex. Some components of a biosimilar’s HCP profile may be found in the RLD, but significant differences in amount and type of HCPs can exist, as demonstrated in a direct comparison of an RLD with a biosimilar candidate (84). Thus, both qualitative and quantitative differences between RLD and biosimilar HCP profiles should be assessed for biosimilar development and interchangeability of products.
Currently, there have been few studies comparing host cell protein immunogenicity as predicted in silico with clinical or pre-clinical studies of immunogenicity. It will be important to perform validating studies of in silico immunogenicity risk assessment to improve existing methods. Furthermore, combinations of in silico and in vitro studies are now being included in abbreviated new drug applications (not requiring clinical studies) for the office of generic drugs at the FDA (85). Similar approaches might be feasible regarding biosimilar biologic products once alternative methods like ISPRI-HCP are developed and validated. In any case, the evaluation of HCPs and PRPIs for potential immunogenicity is an important means of reducing the potential for problems related to drug safety and efficacy and should be an important component of the drug development pathway for all cell-culture derived products in the future.
Data Availability
All data generated or analysed during this study are included in this published article [and its supplementary information files].
Change history
19 December 2023
A Correction to this paper has been published: https://doi.org/10.1208/s12248-023-00868-5
Abbreviations
- HCP:
-
Host cell protein
- PRPI:
-
Process-related protein impurity
- EpiBar:
-
Epitope bar; promiscuous T cell epitope
- RLD:
-
Reference listed drug
References
(2022) Fact Sheet: FDA at a Glance. In: FDA U.S. Food and drug administration. https://www.fda.gov/about-fda/fda-basics/fact-sheet-fda-glance. Accessed 13 Feb 2023
BioSpace (2022) Biologics Market Size to Hit USD 719.94 Billion by 2030. In: BioSpace. https://www.biospace.com/article/biologics-market-size-to-hit-usd-719-94-billion-by-2030-/. Accessed 13 Feb 2023
Senior M. Fresh from the biotech pipeline: fewer approvals, but biologics gain share. Nat Biotechnol. 2023;41:174–82. https://doi.org/10.1038/s41587-022-01630-6.
Vanderlaan M, Zhu-Shimoni J, Lin S, Gunawan F, Waerner T, Van Cott KE. Experience with host cell protein impurities in biopharmaceuticals. Biotechnol Prog. 2018;34:828–37.
Park JH, Jin JH, Lim MS, An HJ, Kim JW, Lee GM. Proteomic Analysis of Host Cell Protein Dynamics in the Culture Supernatants of Antibody-Producing CHO Cells. Sci Rep. 2017;7:44246.
Gao SX, Zhang Y, Stansberry-Perkins K, Buko A, Bai S, Nguyen V, Brader ML. Fragmentation of a highly purified monoclonal antibody attributed to residual CHO cell protease activity. Biotechnol Bioeng. 2011;108:977–82.
Robert F, Bierau H, Rossi M, Agugiaro D, Soranzo T, Broly H, Mitchell-Logean C. Degradation of an Fc-fusion recombinant protein by host cell proteases: Identification of a CHO cathepsin D protease. Biotechnol Bioeng. 2009;104:1132–41.
Wang X, Hunter AK, Mozier NM. Host cell proteins in biologics development: Identification, quantitation and risk assessment. Biotechnol Bioeng. 2009;103:446–58.
Jones M, Palackal N, Wang F, et al. “High-risk” host cell proteins (HCPs): A multi-company collaborative view. Biotechnol Bioeng. 2021;118:2870–85.
Bracewell DG, Francis R, Smales CM. The future of host cell protein (HCP) identification during process development and manufacturing linked to a risk-based management for their control. Biotechnol Bioeng. 2015;112:1727–37.
Jawa V, Joubert MK, Zhang Q, Deshpande M, Hapuarachchi S, Hall MP, Flynn GC. Evaluating Immunogenicity Risk Due to Host Cell Protein Impurities in Antibody-Based Biotherapeutics. AAPS Journal. 2016;18:1439–52.
Pilely K, Johansen MR, Lund RR, Kofoed T, Jørgensen TK, Skriver L, Mørtz E. Monitoring process-related impurities in biologics-host cell protein analysis. Anal Bioanal Chem. 2022;414:747–58.
Guo J, Kufer R, Li D, Wohlrab S, Greenwood-Goodwin M, Yang F. Technical advancement and practical considerations of LC-MS/MS-based methods for host cell protein identification and quantitation to support process development. MAbs. 2023;15:2213365.
Mörtstedt H, Makower Å, Edlund P-O, Sjöberg K, Tjernberg A. Improved identification of host cell proteins in a protein biopharmaceutical by LC-MS/MS using the ProteoMiner™ Enrichment Kit. J Pharm Biomed Anal. 2020;185:113256.
Zhu-Shimoni J, Yu C, Nishihara J, Wong RM, Gunawan F, Lin M, Krawitz D, Liu P, Sandoval W, Vanderlaan M. Host cell protein testing by ELISAs and the use of orthogonal methods. Biotechnol Bioeng. 2014;111:2367–79.
Tscheliessnig AL, Konrath J, Bates R, Jungbauer A. Host cell protein analysis in therapeutic protein bioprocessing - methods and applications. Biotechnol J. 2013;8:655–70.
Strasser L, Oliviero G, Jakes C, Zaborowska I, Floris P, Ribeiro da Silva M, Füssl F, Carillo S, Bones J. Detection and quantitation of host cell proteins in monoclonal antibody drug products using automated sample preparation and data-independent acquisition LC-MS/MS. J Pharm Anal. 2021;11:726–31.
De Groot AS, Desai AK, Lelias S, et al. Immune Tolerance-Adjusted Personalized Immunogenicity Prediction for Pompe Disease. Front Immunol. 2021;12:636731. https://doi.org/10.3389/fimmu.2021.636731.
Bailey-Kellogg C, Gutiérrez AH, Moise L, Terry F, Martin WD, De Groot AS. CHOPPI: A Web Tool for the Analysis of Immunogenicity Risk from Host Cell Proteins in CHO-Based Protein Production. Biotechnol Bioeng. 2014;111:2170–82.
Jawa V, Terry F, Gokemeijer J, Mitra-Kaushik S, Roberts BJ, Tourdot S, De Groot AS. T-Cell Dependent Immunogenicity of Protein Therapeutics Pre-clinical Assessment and Mitigation-Updated Consensus and Review 2020. Front Immunol. 2020;11:1301. https://doi.org/10.3389/fimmu.2020.01301.
Hrdinová J, Verbij FC, Kaijen PHP, et al. Mass spectrometry-assisted identification of ADAMTS13-derived peptides presented on HLA-DR and HLA-DQ. Haematologica. 2018;103:1083–92.
Moise L, Gutierrez AH, Bailey-Kellogg C, et al. The two-faced T cell epitope: Examining the host-microbe interface with JanusMatrix. Hum Vaccin Immunother. 2013;9(7):1577–86.
De Groot AS, Moise L, Terry F, et al. Better Epitope Discovery, Precision Immune Engineering, and Accelerated Vaccine Design Using Immunoinformatics Tools. Front Immunol. 2020;11:442.
Mufarrege EF, Giorgetti S, Etcheverrigaray M, Terry F, Martin W, De Groot AS. De-immunized and Functional Therapeutic (DeFT) versions of a long lasting recombinant alpha interferon for antiviral therapy. Clin Immunol. 2017;176:31–41.
Moise L, Buller RM, Schriewer J, Lee J, Frey SE, Weiner DB, Martin W, de Groot AS. VennVax, a DNA-prime, peptide-boost multi-T-cell epitope poxvirus vaccine, induces protective immunity against vaccinia infection by T cell response alone. Vaccine. 2011;29:501–11.
Schanen BC, de Groot AS, Moise L, Ardito M, McClaine E, Martin W, Wittman V, Warren WL, Drake DR. Coupling sensitive in vitro and in silico techniques to assess cross-reactive CD4+ T cells against the swine-origin H1N1 influenza virus. Vaccine. 2011;29:3299–309.
Moise L, Tassone R, Latimer H, Terry F, Levitz L, Haran JP, Ross TM, Boyle C, Martin WD, de Groot AS. Immunization with cross-conserved H1N1 influenza CD4 + T-cell epitopes lowers viral burden in HLA DR3 transgenic mice. Hum Vaccin Immunother. 2013;9:2060–8.
Scholzen A, Richard G, Moise L, et al. Promiscuous Coxiella burnetii CD4 Epitope Clusters Associated With Human Recall Responses Are Candidates for a Novel T-Cell Targeted Multi-Epitope Q Fever Vaccine. Front Immunol. 2019;10:207. https://doi.org/10.3389/fimmu.2019.00207.
Winterling K, Martin WD, de Groot AS, Daufenbach J, Kistner S, Schüttrumpf J. Development of a novel fully functional coagulation factor VIII with reduced immunogenicity utilizing an in silico prediction and deimmunization approach. J Thromb Haemost. 2021;19:2161–70.
de Groot AS, Rosenberg AS, Miah SMS, Skowron G, Roberts BJ, Lélias S, Terry FE, Martin WD. Identification of a potent regulatory T cell epitope in factor V that modulates CD4+ and CD8+ memory T cell responses. Clin Immunol. 2021;224:108661.
Liu R, Moise L, Tassone R, Gutierrez AH, Terry FE, Sangare K, Ardito MT, Martin WD, de Groot AS. H7N9 T-cell epitopes that mimic human sequences are less immunogenic and may induce Treg-mediated tolerance. Hum Vaccin Immunother. 2015;11:2241–52.
Losikoff PT, Mishra S, Terry F, et al. HCV epitope, homologous to multiple human protein sequences, induces a regulatory T cell response in infected patients. J Hepatol. 2015;62:48–55.
Brinkrolf K, Rupp O, Laux H, et al. Chinese hamster genome sequenced from sorted chromosomes. Nat Biotechnol. 2013;31:694–5.
Partridge MA, Davidson MM, Hei TK. The complete nucleotide sequence of Chinese hamster (Cricetulus griseus) mitochondrial DNA. DNA Seq. 2007;18:341–6.
Xu X, Nagarajan H, Lewis NE, et al. The genomic sequence of the Chinese hamster ovary (CHO)-K1 cell line. Nat Biotechnol. 2011;29:735–41.
Jeong H, Kim HJ, Lee SJ. Complete Genome Sequence of Escherichia coli Strain BL21. Genome Announc. 2015;3(2):e00134-15. https://doi.org/10.1128/genomeA.00134-15.
Blattner FR, Plunkett G, Bloch CA, et al. The complete genome sequence of Escherichia coli K-12. Science. 1997;277:1453–62.
Tsakok T, Saklatvala J, Rispens T, et al. Development of antidrug antibodies against adalimumab maps to variation within the HLA-DR peptide-binding groove. JCI Insight. 2023;8(4):e156643. https://doi.org/10.1172/jci.insight.156643.
Hammer C, Ruppel J, Kamen L, Hunkapiller J, Mellman I, Quarmby V. Allelic variation in HLA-DRB1 is associated with development of antidrug antibodies in cancer patients treated with atezolizumab that are neutralizing in vitro. Clin Transl Sci. 2022;15:1393–9.
Lund O, Nielsen M, Kesmir C, et al. Definition of supertypes for HLA molecules using clustering of specificity matrices. Immunogenetics. 2004;55:797–810.
Southwood S, Sidney J, Kondo A, del Guercio MF, Appella E, Hoffman S, Kubo RT, Chesnut RW, Grey HM, Sette A. Several common HLA-DR types share largely overlapping peptide binding repertoires. J Immunol. 1998;160:3363–73.
Moise L, Gutierrez A, Kibria F, Martin R, Tassone R, Liu R, Terry F, Martin B, De Groot AS. iVAX: An integrated toolkit for the selection and optimization of antigens and the design of epitope-driven vaccines. Hum Vaccin Immunother. 2015;11:2312–21.
Weber CA, Mehta PJ, Ardito M, Moise L, Martin B, De Groot AS. T cell epitope: friend or foe? Immunogenicity of biologics in context. Adv Drug Deliv Rev. 2009;61:965–76.
Klein L, Kyewski B, Allen PM, Hogquist KA. Positive and negative selection of the T cell repertoire: what thymocytes see (and don’t see). Nat Rev Immunol. 2014;14:377–91.
Wada Y, Nithichanon A, Nobusawa E, et al. A humanized mouse model identifies key amino acids for low immunogenicity of H7N9 vaccines. Sci Rep. 2017;7:1283.
Noe AR, Terry FE, Schanen BC, et al. Bridging Computational Vaccinology and Vaccine Development Through Systematic Identification, Characterization, and Downselection of Conserved and Variable Circumsporozoite Protein CD4 T Cell Epitopes From Diverse Plasmodium falciparum Strains. Front Immunol. 2021;12:689920.
Gámez A, Wang L, Sarkissian CN, Wendt D, Fitzpatrick P, Lemontt JF, Scriver CR, Stevens RC. Structure-based epitope and PEGylation sites mapping of phenylalanine ammonia-lyase for enzyme substitution treatment of phenylketonuria. Mol Genet Metab. 2007;91:325–34.
De Groot AS, Roberts BJ, Mattei A, Lelias S, Boyle C (2023) An Immunogenicity Risk Assessment Framework for Generic Peptide Drugs and their Impurities (Manuscript in preparation).
Mattei AE, Gutierrez AH, Martin WD, Terry FE, Roberts BJ, Rosenberg AS, de Groot AS. In silico immunogenicity assessment for sequences containing unnatural amino acids: A method using existing in silico algorithm infrastructure and a vision for future enhancements. Front Drug Discov. 2022;2:952326. https://doi.org/10.3389/fddsv.2022.952326.
Wullner D, Zhou L, Bramhall E, Kuck A, Goletz TJ, Swanson S, Chirmule N, Jawa V. Considerations for optimization and validation of an in vitro PBMC derived T cell assay for immunogenicity prediction of biotherapeutics. Clin Immunol. 2010;137:5–14.
Champion KM, Madden H, Dougherty J, Shacter E (2005) Defining Your Product Profile and Maintaining Control Over It , Part 2 Challenges of Monitoring Host Cell Protein Impurities. In: Bioprocess Int. https://bioprocessintl.com/2005/september-2005/defining-product-profile-maintaining-control-part-2/. Accessed 16 Jul 2023
Fierce Biotech (2012) Ipsen’s partner Inspiration Biopharmaceuticals announces hold of phase III clinical trials evaluating IB1001 for the treatment and prevention of hemophilia B. https://www.fiercebiotech.com/biotech/ipsen-s-partner-inspiration-biopharmaceuticals-announces-hold-of-phase-iii-clinical-trials. Accessed 16 Jul 2023
Jennifer Levin (2013) Fierce Biotech: Ipsen and Inspiration Biopharmaceuticals announce closing of the IB1001 sale to Cangene Corporation. https://www.fiercebiotech.com/biotech/ipsen-and-inspiration-biopharmaceuticals-announce-closing-of-ib1001-sale-to-cangene. Accessed 16 Jul 2023
Reijers JAA, Malone KE, Bajramovic JJ, Verbeek R, Burggraaf J, Moerland M. Adverse immunostimulation caused by impurities: The dark side of biopharmaceuticals. Br J Clin Pharmacol. 2019;85:1418–26.
U.S. FDA (2021) ANDAs for Certain Highly Purified Synthetic Peptide Drug Products That Refer to Listed Drugs of rDNA Origin Guidance for Industry. https://www.fda.gov/regulatory-information/search-fda-guidance-documents/andas-certain-highly-purified-synthetic-peptide-drug-products-refer-listed-drugs-rdna-origin. Accessed 16 Jul 2023
U.S. FDA, Center for Drug Evaluation and Research, Center for Biologics Evaluation and Research (2019) Immunogenicity Testing of Therapeutic Protein Products —Developing and Validating Assays for Anti-Drug Antibody Detection Guidance for Industry. https://www.fda.gov/regulatory-information/search-fda-guidance-documents/immunogenicity-testing-therapeutic-protein-products-developing-and-validating-assays-anti-drug. Accessed 16 Jul 2023
de Zafra CLZ, Quarmby V, Francissen K, Vanderlaan M, Zhu-Shimoni J. Host cell proteins in biotechnology-derived products: A risk assessment framework. Biotechnol Bioeng. 2015;112:2284–91.
Fischer SK, Cheu M, Peng K, Lowe J, Araujo J, Murray E, McClintock D, Matthews J, Siguenza P, Song A. Specific Immune Response to Phospholipase B-Like 2 Protein, a Host Cell Impurity in Lebrikizumab Clinical Material. AAPS J. 2017;19:254–63.
Suh W-K. Life of T Follicular Helper Cells. Mol Cells. 2015;38:195–201.
Crotty S. Follicular Helper CD4 T Cells (T FH ). Annu Rev Immunol. 2011;29:621–63.
Khan S, Parrillo M, Gutierrez AH, Terry FE, Moise L, Martin WD, de Groot AS. Immune escape and immune camouflage may reduce the efficacy of RTS, S vaccine in Malawi. Hum Vaccin Immunother. 2020;16:214–27.
de Groot AS, Moise L, Liu R, Gutierrez AH, Tassone R, Bailey-Kellogg C, Martin W. Immune camouflage: relevance to vaccines and human immunology. Hum Vaccin Immunother. 2014;10:3570–5.
Inaba H, Moise L, Martin W, de Groot AS, Desrosiers J, Tassone R, Buchman G, Akamizu T, de Groot LJ. Epitope Recognition in HLA-DR3 Transgenic Mice Immunized to TSH-R Protein or Peptides. Endocrinology. 2013;154:2234–43.
Grosenbaugh DA, Leard AT, Bergman PJ, et al. Safety and efficacy of a xenogeneic DNA vaccine encoding for human tyrosinase as adjunctive treatment for oral malignant melanoma in dogs following surgical excision of the primary tumor. Am J Vet Res. 2011;72:1631–8.
Prasad S, Kohm AP, McMahon JS, Luo X, Miller SD. Pathogenesis of NOD diabetes is initiated by reactivity to the insulin B chain 9–23 epitope and involves functional epitope spreading. J Autoimmun. 2012;39:347–53.
Rosenberg AS, Worobec A (2004) A Risk-Based Approach to Immunogenicity Concerns of Therapeutic Protein Products, Part 1: Considering Consequences of the Immune Response to a Protein. In: Biopharm Int. https://www.biopharminternational.com/view/risk-based-approach-immunogenicity-concerns-therapeutic-protein-products-part-1-considering-conseque. Accessed 17 Jul 2023
Hamuro L, Tirucherai GS, Crawford SM, Nayeem A, Pillutla RC, DeSilva BS, Leil TA, Thalhauser CJ. Evaluating a Multiscale Mechanistic Model of the Immune System to Predict Human Immunogenicity for a Biotherapeutic in Phase 1. AAPS J. 2019;21:94.
Koren E, Zuckerman LA, Mire-Sluis AR. Immune responses to therapeutic proteins in humans–clinical significance, assessment and prediction. Curr Pharm Biotechnol. 2002;3:349–60.
Casadevall N, Nataf J, Viron B, et al. Pure red-cell aplasia and antierythropoietin antibodies in patients treated with recombinant erythropoietin. N Engl J Med. 2002;346:469–75.
Koren E, de Groot AS, Jawa V, et al. Clinical validation of the “in silico” prediction of immunogenicity of a human recombinant therapeutic protein. Clin Immunol. 2007;124:26–32.
Kruiswijk C, Richard G, Salverda MLM, Hindocha P, Martin WD, De Groot AS, Van Riet E. In silico identification and modification of T cell epitopes in pertussis antigens associated with tolerance. Hum Vaccin Immunother. 2020;16:277–85.
Molden R, Hu M, Yen ES, Saggese D, Reilly J, Mattila J, Qiu H, Chen G, Bak H, Li N. Host cell protein profiling of commercial therapeutic protein drugs as a benchmark for monoclonal antibody-based therapeutic protein development. MAbs. 2021;13(1):1955811. https://doi.org/10.1080/19420862.2021.1955811.
Greinacher A, Thiele T, Warkentin TE, Weisser K, Kyrle PA, Eichinger S. Thrombotic Thrombocytopenia after ChAdOx1 nCov-19 Vaccination. N Engl J Med. 2021;384:2092–101.
Muir K-L, Kallam A, Koepsell SA, Gundabolu K. Thrombotic Thrombocytopenia after Ad26.COV2.S Vaccination. N Engl J Med. 2021;384:1964–5.
Elisabeth Mahase (2021) Covid-19: US suspends Johnson and Johnson vaccine rollout over blood clots. BMJ. https://doi.org/10.1136/bmj.n970. Accessed 16 Jul 2023
Jacqui Wise (2021) Covid-19: European countries suspend use of Oxford-AstraZeneca vaccine after reports of blood clots. In: BMJ. https://doi.org/10.1136/bmj.n699. Accessed 16 Jul 2023
Greinacher A, Selleng K, Palankar R, et al. Insights in ChAdOx1 nCoV-19 vaccine-induced immune thrombotic thrombocytopenia. Blood. 2021;138:2256–68.
Krutzke L, Rösler R, Allmendinger E, Engler T, Wiese S, Kochanek S. Process- and product-related impurities in the ChAdOx1 nCov-19 vaccine. Elife. 2022;11:e78513. https://doi.org/10.7554/eLife.78513.
Papp K, Bachelez H, Costanzo A, et al. Clinical similarity of biosimilar ABP 501 to adalimumab in the treatment of patients with moderate to severe plaque psoriasis: A randomized, double-blind, multicenter, phase III study. J Am Acad Dermatol. 2017;76:1093–102.
U.S. Food and Drug Administration (2022) FDA Biosimilar User Fee Act (BsUFA) Research Grant (U01). https://grants.nih.gov/grants/guide/rfa-files/RFA-FD-23-026.html. Accessed 17 Jul 2023
Hogwood CEM, Tait AS, Koloteva-Levine N, Bracewell DG, Smales CM. The dynamics of the CHO host cell protein profile during clarification and protein A capture in a platform antibody purification process. Biotechnol Bioeng. 2013;110:240–51.
Zhang Q, Goetze AM, Cui H, Wylie J, Trimble S, Hewig A, Flynn GC. Comprehensive tracking of host cell proteins during monoclonal antibody purifications using mass spectrometry. MAbs. 2014;6:659–70.
Schenauer MR, Flynn GC, Goetze AM. Profiling the effects of process changes on residual host cell proteins in biotherapeutics by mass spectrometry. Biotechnol Prog. 2013;29:951–7.
Mihara K, Ito Y, Hatano Y, et al. Host Cell Proteins: The Hidden Side of Biosimilarity Assessment. J Pharm Sci. 2015;104:3991–6.
U.S. Food & Drug Administration (2022) Abbreviated New Drug Application (ANDA). https://www.fda.gov/drugs/types-applications/abbreviated-new-drug-application-anda. Accessed 26 Feb 2023
Acknowledgements
The research for the case studies described in this report and for the review was performed by EpiVax team members with support from internal funds and with funding from the FDA Center for Drug Evaluation and Research (CDER) (# U01FD007760). CDER funding will support further validation of a method for immunogenicity risk assessment of host cell proteins (HCP) that may be useful for improving biosimilar product development and assessing product interchangeability.
Funding
This research was funded by EpiVax Inc.
Author information
Authors and Affiliations
Contributions
KH wrote the first draft of the manuscript and created figures. All authors contributed to the conceptualization and writing of the manuscript, made substantial edits, reviewed, and approved the manuscript for submission.
Corresponding author
Ethics declarations
Conflict of Interest
AD and WM are senior officer and shareholders, and KH, BJR, SK, FT, CMB, MM, and AR are employees of EpiVax, Inc., a privately owned biotechnology company located in Providence, RI. These authors acknowledge that there is a potential conflict of interest inherent in the publication of this manuscript, and attest that the work contained in this research report is free of any bias that might be associated with the commercial goals of the company.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
The original online version of this article was revised to replace "HCP" to "product" in the sentence "The general guidelines for the maximum amount of allowed HCP impurities is 100 ng per mg of product.".
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Haltaufderhyde, K., Roberts, B.J., Khan, S. et al. Immunoinformatic Risk Assessment of Host Cell Proteins During Process Development for Biologic Therapeutics. AAPS J 25, 87 (2023). https://doi.org/10.1208/s12248-023-00852-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1208/s12248-023-00852-z