Abstract
Autogene cevumeran is an individualized neoantigen-specific therapy (iNeST) under development for the treatment of various solid tumors. It consists of an RNA-Lipoplex (RNA-LPX) in which the encapsulated mRNA molecule encodes up to ten neoepitopes identified from each individual patient. In association with major histocompatibility complex (MHC) class I and MHC class II, these neoantigens can potentially stimulate and expand neoantigen-specific CD4+ and CD8+ T cells, leading to antitumor responses. As part of the pharmacokinetic (PK) property assessment of Autogene cevumeran in patients, both the lipid and mRNA content in circulation are measured. This work focused on our efforts to establish a sensitive and robust method for the measurement of mRNA levels of RNA-LPX in plasma. Due to the chemical characteristics of mRNA, extra precautions are required in order to effectively preserve mRNA integrity in human plasma during sample collection, handling and storage. To this end, a number of sample collection tubes and storage conditions were evaluated in order to inform the most optimal and operationally feasible conditions by which to preserve mRNA integrity during sample collection and upon freeze–thaw. PAXgene Blood ccfDNA tubes successfully prevented mRNA degradation and were subsequently selected for patient sample collection in the clinical trial. A branched DNA (bDNA)-based mRNA PK assay was developed to achieve the desired assay performance. Here, we discuss the evaluation of various sample collection and processing conditions as well as the optimization of the work flow during bDNA PK method development.
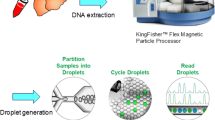
Graphical Abstract






Similar content being viewed by others
References
Blass E, Ott PA. Advances in the development of personalized neoantigen-based therapeutic cancer vaccines. Nat Rev Clin Oncol. 2021;18(4):215–29. https://doi.org/10.1038/s41571-020-00460-2.
Haen SP, Rammensee HG. The repertoire of human tumor-associated epitopes--identification and selection of antigens and their application in clinical trials. Curr Opin Immunol. 2013;25(2):277–83. https://doi.org/10.1016/j.coi.2013.03.007.
Vogelstein B, Papadopoulos N, Velculescu VE, Zhou S, Diaz LA Jr, Kinzler KW. Cancer genome landscapes. Science. 2013;339(6127):1546–58. https://doi.org/10.1126/science.1235122.
Chalmers ZR, Connelly CF, Fabrizio D, Gay L, Ali SM, Ennis R, et al. Analysis of 100,000 human cancer genomes reveals the landscape of tumor mutational burden. Genome Med. 2017;9(1):34. https://doi.org/10.1186/s13073-017-0424-2.
Shemesh CS, Hsu JC, Hosseini I, Shen BQ, Rotte A, Twomey P, et al. Personalized cancer vaccines: clinical landscape, challenges, and opportunities. Mol Ther. 2021;29(2):555–70. https://doi.org/10.1016/j.ymthe.2020.09.038.
Kranz LM, Diken M, Haas H, Kreiter S, Loquai C, Reuter KC, et al. Systemic RNA delivery to dendritic cells exploits antiviral defence for cancer immunotherapy. Nature. 2016;534(7607):396–401. https://doi.org/10.1038/nature18300.
Sahin U, Oehm P, Derhovanessian E, Jabulowsky RA, Vormehr M, Gold M, et al. An RNA vaccine drives immunity in checkpoint-inhibitor-treated melanoma. Nature. 2020;585(7823):107–12. https://doi.org/10.1038/s41586-020-2537-9.
Kreiter S, Selmi A, Diken M, Sebastian M, Osterloh P, Schild H, et al. Increased antigen presentation efficiency by coupling antigens to MHC class I trafficking signals. J Immunol. 2008;180(1):309–18. https://doi.org/10.4049/jimmunol.180.1.309.
Henderson N, Wilson A. Measurement of mRNA therapeutics: method development and validation challenges. Bioanalysis. 2019;11(21):2003–10. https://doi.org/10.4155/bio-2019-0120.
Hawthorne G, Henderson N, Holtta M, Khan S, Lindqvist J, Wilson A. Overcoming analytical challenges to generate data critical to understanding lipid nanoparticle-delivered modified mRNA biodistribution. Bioanalysis. 2019;11(21):1993–2001. https://doi.org/10.4155/bio-2019-0138.
Lubelchek RJ, Max B, Sandusky CJ, Hota B, Barker DE. Reliability at the lower limits of HIV-1 RNA quantification in clinical samples: a comparison of RT-PCR versus bDNA assays. PLoS One. 2009;4(6):e6008. https://doi.org/10.1371/journal.pone.0006008.
Tsongalis GJ. Branched DNA technology in molecular diagnostics. Am J Clin Pathol. 2006;126(3):448–53. https://doi.org/10.1309/90BU6KDXANFLN4RJ.
Collins ML, Irvine B, Tyner D, Fine E, Zayati C, Chang C, et al. A branched DNA signal amplification assay for quantification of nucleic acid targets below 100 molecules/ml. Nucleic Acids Res. 1997;25(15):2979–84. https://doi.org/10.1093/nar/25.15.2979.
Redrup MJ, Igarashi H, Schaefgen J, Lin J, Geisler L, Ben M'Barek M, et al. Sample management: recommendation for best practices and harmonization from the Global Bioanalysis Consortium Harmonization Team. AAPS J. 2016;18(2):290–3. https://doi.org/10.1208/s12248-016-9869-2.
Inglut CT, Sorrin AJ, Kuruppu T, Vig S, Cicalo J, Ahmad H, et al. Immunological and Toxicological Considerations for the Design of Liposomes. Nanomaterials (Basel). 2020;10(2). https://doi.org/10.3390/nano10020190.
van de Merbel N, Savoie N, Yadav M, Ohtsu Y, White J, Riccio MF, et al. Stability: recommendation for best practices and harmonization from the Global Bioanalysis Consortium Harmonization Team. AAPS J. 2014;16(3):392–9. https://doi.org/10.1208/s12248-014-9573-z.
Reddi KK, Holland JF. Elevated serum ribonuclease in patients with pancreatic cancer. Proc Natl Acad Sci U S A. 1976;73(7):2308–10. https://doi.org/10.1073/pnas.73.7.2308.
Ilinskaya ON, Mahmud RS. Ribonucleases as antiviral agents. Mol Biol. 2014;48(5):615–23. https://doi.org/10.1134/S0026893314040050.
Tsui NB, Ng EK, Lo YM. Stability of endogenous and added RNA in blood specimens, serum, and plasma. Clin Chem. 2002;48(10):1647–53.
Larson MH, Pan W, Kim HJ, Mauntz RE, Stuart SM, Pimentel M, et al. A comprehensive characterization of the cell-free transcriptome reveals tissue- and subtype-specific biomarkers for cancer detection. Nat Commun. 2021;12(1):2357. https://doi.org/10.1038/s41467-021-22444-1.
Gleaves CA, Welle J, Campbell M, Elbeik T, Ng V, Taylor PE, et al. Multicenter evaluation of the Bayer VERSANT HIV-1 RNA 3.0 assay: analytical and clinical performance. J Clin Virol. 2002;25(2):205–16. https://doi.org/10.1016/s1386-6532(02)00011-2.
Trimoulet P, Halfon P, Pohier E, Khiri H, Chene G, Fleury H. Evaluation of the VERSANT HCV RNA 3.0 assay for quantification of hepatitis C virus RNA in serum. J Clin Microbiol. 2002;40(6):2031–6. https://doi.org/10.1128/JCM.40.6.2031-2036.2002.
Bahl K, Senn JJ, Yuzhakov O, Bulychev A, Brito LA, Hassett KJ, et al. Preclinical and clinical demonstration of immunogenicity by mRNA vaccines against H10N8 and H7N9 influenza viruses. Mol Ther. 2017;25(6):1316–27. https://doi.org/10.1016/j.ymthe.2017.03.035.
Sedic M, Senn JJ, Lynn A, Laska M, Smith M, Platz SJ, et al. Safety evaluation of lipid nanoparticle-formulated modified mRNA in the sprague-dawley rat and cynomolgus monkey. Vet Pathol. 2018;55(2):341–54. https://doi.org/10.1177/0300985817738095.
Acknowledgements
We want to thank Chris Petry and Sara Wichner for providing critical reagents. The graphical abstract was designed using BioRender.
Funding
All work was funded by Genentech Inc., a member of the Roche group.
Author information
Authors and Affiliations
Contributions
All authors contributed to the experimental design and data analysis. YZ and AB did the experimental work. SG and KP wrote the manuscript.
Corresponding author
Ethics declarations
Conflict of Interest
All authors are employees of Genentech Inc., a member of the Roche group and Roche shareholders.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Guelman, S., Zhou, Y., Brady, A. et al. A Fit-for-Purpose Method to Measure Circulating Levels of the mRNA Component of a Liposomal-Formulated Individualized Neoantigen-Specific Therapy for Cancer. AAPS J 24, 64 (2022). https://doi.org/10.1208/s12248-022-00709-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1208/s12248-022-00709-x