Abstract
Glaucoma is a chronic neurodegenerative process of the optic nerve that is the leading cause of blindness worldwide, and early diagnosis of the disease could greatly affect patients’ prognoses. The pathophysiology of glaucoma is complicated by a combination of genetic and epigenetic factors. Deciphering the early diagnostic biomarkers in glaucoma could attenuate the disease's global burden and help us understand the exact mechanisms involved in glaucoma. The microRNAs are members of a larger family of non-coding RNAs that play an essential role in the epigenetic basis of glaucoma. A systematic study and meta-analysis of diagnostic microRNAs in glaucoma, jointly with network analysis of target genes, were carried out on published papers assessing differentially expressed microRNAs in human subjects. In total, 321 articles were found, and, after screening, six studies were eligible for further analysis. 52 differentially expressed microRNAs were found, of which 28 and 24 were up-regulated and down-regulated, respectively. Only 12 microRNAs were qualified for meta-analysis, with overall sensitivity and specificity of 80% and 74%, respectively. Then, using network analysis, it became apparent that the VEGF-A, AKT1, CXCL12, and HRAS genes were the most important targets for the microRNAs. Perturbations in WNT signaling, protein transport, and extracellular matrix organization pathways were discovered to be important in the etiology of glaucoma using the community detection approach. This study tries to uncover the promising microRNAs and their target genes that govern the epigenetics of glaucoma.
Similar content being viewed by others
Introduction
Glaucoma is a chronic neurodegenerative process within the optic nerve that is the leading cause of blindness, affecting more than 70 million people worldwide [1, 2]. The imperceptible and gradual nature of the disease leads to delayed diagnosis and irreversible neuropathy. Clinical diagnosis of glaucoma is relatively straightforward, but the delayed diagnosis puts a stop to the timely intervention; histopathologic changes are way before clinical changes in the optic disc and visual function [3]. The pathophysiology of glaucoma remains largely unknown. Our knowledge is limited except for several well-known risk factors, such as a strong family history of glaucoma, increased intraocular pressure, advanced age, corticosteroid use, and the black race [4]. Although increased intraocular pressure (IOP) is the most common and the only modifiable risk factor for glaucoma, in about thirty percent of patients with glaucoma, the IOP is within the normal range, which implies that there must be other contributing factors in the development of optic neuropathy [4, 5]. As we can infer from the risk factors indicated, there must be a combination of genetic and environmental interactions in the pathogenesis of glaucoma. There are various hypotheses for the pathophysiology of glaucoma that have yet to be completely explored, including neurotrophic factor deficiency, excitotoxicity, ischemia, oxidative stress, and axonal transport failure [6].
Epigenetics is the study of how environmental circumstances influence our genetics, and it should be acknowledged as an important component of glaucoma pathogenesis and development [7]. One of the key arms of the epigenetic regulatory effect is microRNAs (also called miRNAs). MicroRNAs are members of a larger family of non-coding RNAs, and the average length of microRNAs is 22 nucleotides [8].
MicroRNAs have garnered tremendous attention over the past few years due to the large number of processes they govern, such as cell growth and death. Basically, microRNAs interfere with messenger RNA translation, and in this way, they regulate about a third of human protein-coding genes [8]. Emerging studies suggests that the microRNAs could play a dual role in glaucoma. Protective microRNAs such as micro-RNA-483-3p decrease extracellular matrix fibrosis in response to stress. On the other hand, disease-promoting microRNAs such as microRNA-100 have been shown to reduce nerve growth [7]. A growing body of research has focused extensively on utilizing microRNAs in establishing the diagnosis and prognosis of chronic diseases such as glaucoma [9]. However, the accuracy and validity of these microRNAs in establishing the diagnosis of glaucoma are still under debate.
Identifying the most important evidence-based microRNAs can help us in the early diagnosis of the disease, in its prevention, and in the design of drugs that change the nature of the disease. Our study is nevertheless a valuable contribution to the field as it suggests a systematic review, meta-analysis, and genetic network study of the most critical microRNAs orchestrating the development of glaucoma.
Materials and methods
Databases and search strategy
PubMed, Web of Science, EMBASE, OVID, and Scopus databases were utilized, and papers from inception to December 30 2021 with the combination of the following search terms, were extracted: human, case or patient, control, microRNA, glaucoma, intraocular pressure, or hypertension. The full search strategy was shown in the method of literature search part.
Method of literature search
("human" OR "humans" OR "patient" OR "patients" OR "control" OR "controls" OR "group" OR "groups") AND ("microrna" OR "micro RNA" OR "micrornas" OR "mirs" OR "microRNA") AND ("glaucoma" OR "intraocular hypertension" OR "intraocular pressure" OR "ocular hypertension").
Inclusion and exclusion criteria
Articles are eligible for inclusion if they fulfill the following criteria: [1] original articles that are designed as a case–control study to assess the microRNA expression profiling in glaucoma patients; [2] the study must provide a source of samples obtained from the case and control groups [3]. MicroRNA profiling must be accomplished using one of the qPCR, microarray, or next-generation sequencing methods, and [4] the cut-off criteria for detection of differentially expressed microRNA must be reported. Review articles, studies not published in English, and animal and cell line studies were excluded.
Data extraction
Two independent researchers (HS and MSS) screened, evaluated, and extracted the data to minimize the risk of selection bias in the study. In cases where two researchers disagreed on the inclusion or exclusion of an article, the third researcher (MR) made the definitive decision.
Quality assessment
The quality of the included studies was assessed using the Quality Assessment of Diagnostic Accuracy Studies (QUADAS-2) criteria [10]. This method uses seven questions to evaluate the diagnostic accuracy studies in four key domains: patient selection, index test, reference standard, flow, and time. The seven questions are listed as follows: (1) Could the selection of patients have introduced bias? (2) Are there concerns that the included patients and setting do not match the review question? (3) Could the conduct or interpretation of the index test have introduced bias? (4) Are there concerns that the index test, its conduct, or its interpretation differ from the review question? (5) Could the reference standard, its conduct, or its interpretation have introduced bias? (6) Are there concerns that the target condition, as defined by the reference standard, does not match the question? (7) Could the patient flow have introduced bias? The first two address the assessment of patient selection; the next two address the index test; questions five and six address the reference standard; and the last one addresses the flow and timing.
Statistical analysis
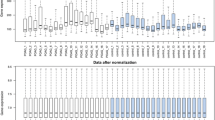
The analysis was conducted using receiver operator characteristic (ROC) curves to determine the accuracy of the discovered microRNAs in diagnosing glaucoma, as measured by the area under the curve (AUC) value and sensitivity and specificity, if available. The meta-analysis comprised studies that determined the sensitivity and specificity of particular microRNAs in diagnosing glaucoma. A random-effects model (DerSimonian–Laird technique) was utilized to account for the expected heterogeneity of investigations. All plots were constructed using STATA version 17 (StataCorp LP, College, Station, TX, USA). Also, all analyses were performed using STATA 17.0 software. The heterogeneity of included studies was assessed using I2 and χ2 statistics and was judged to be significant if I2 was > 50% or the p-value < 0.05. Where more than two studies of the same sample type were available, a subgroup analysis has been performed.
Network analysis
microRNA target identification
The MirTarBase database was used to find the microRNA targets. The MirTarBase is one of the most comprehensive databases that contains experimentally validated microRNA targets [11]. Targets discovered using at least one of the robust evidence methods, namely the Reporter Assay, Western Blot, or qPCR for the searched microRNAs, were included in the subsequent analyses.
Protein–protein interaction (PPI) network construction and hub genes identification
The STRING database was used to construct the PPI network. For PPI network construction, a minimum interaction score of at least 0.7 (high confidence) was needed. Only interactions between the given genes were allowed, and additional interactions were not considered in the final network. The non-interacting genes were removed. To find the most critical genes in the network (i.e., hub genes), we used a Cytoscape plugin, CytoHubba (version 0.1), which is able to find the essential genes. The top ten genes with the highest maximal clique centrality were considered the hub genes. Maximal clique centrality (MCC) is the largest subset of a network in which every two nodes are connected by a vertex. It has been demonstrated that the MCC score is an efficacious topologic metric for finding hub nodes.
Community detection and prioritization
Communities (also called clusters) are densely connected components in a network with a distinctive function. To find the algorithm with the highest level of modularity, we utilized a Friedman ranking statistic filter approach based on modularity. In other words, the algorithm with the highest level of modularity ranks first, along with the statistical difference between algorithms. Several different algorithms (walk-trap, Markov clustering, surprise, and spectral algorithms) have been used in this benchmark. The CDlib Python library was used to implement the community detection algorithms and ranking process [12].
After finding the most appropriate algorithm for community detection, we will apply a community prioritizing algorithm called CRANK to identify the most promising ones for future experimentation. The gene set enrichment analysis will be applied to each community using the Enrichr server [13]. This approach helps us guide future studies cost-effectively and prevent time-consuming research on drug discovery and repurposing.
Results
Data acquisition and characteristics of the included studies
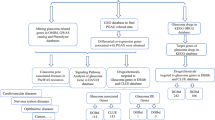
The primary search and study selection procedure is briefly demonstrated as a diagram, famously known as Preferred Reporting Items for Systematic Reviews and Meta-Analyses (PRISMA), in Fig. 1 [14]. As shown, our search strategy retrieved a total of 648 studies. After removing 327 duplicates, 321 studies remained for the title/abstract screening. Respectively, 284 irrelevant studies were omitted, and out of the remaining 37 studies, 6 met the inclusion criteria and entered our study [15,16,17,18,19,20]. These case–control studies measured the microRNA levels in glaucoma patients and healthy people; two of them analyzed the aqueous humor [16, 20], two assessed tears [18, 19], and the other two investigated serum and plasma samples [15, 17]. Table 1 displays the extracted data from the included studies. While one study included only female participants [15], the rest assessed both females and males. Moreover, real-time polymerase chain reaction (RT-PCR), next-generation sequencing, and microarray systems were administered. The risk of bias analysis of the articles based on the QUADAS-2 checklist is summarized in Figs. 2 and 3.
Differentially expressed microRNAs
After the screening of the 321 articles, finally, 6 of them were included [15,16,17,18,19,20]. Fifty-two differentially expressed mature microRNAs in these selected studies compared 115 glaucoma patients with 108 controls. The total number of individual microRNAs found in the included studies was 52, of which 24 were down-regulated, and 28 were up-regulated. The diagnostic accuracy of miR-210-3 was measured in two different scenarios: a screening set with a smaller sample size, followed by a validation in a larger group of patients; it was up-regulated in both steps [17]. Of the 52 individual microRNAs, 3, 3, 11, and 36 were taken from serum, plasma, tears, and aqueous humor, respectively.
All the patients' plasma and serum microRNA samples were up-regulated [15, 17]. However, most of the patients' tear samples contained microRNAs that were up-regulated [18, 19] compared with the control group (9 versus 2). In contrast, of the 36 samples of aqueous humor microRNAs, 22 were down-regulated and 14 were up-regulated [16, 20].
Meta-analysis of diagnostic test accuracy studies
The meta-analysis included studies that detected a single dysregulated microRNA in glaucoma cases compared to controls and the corresponding AUC, sensitivity, and specificity for the discovered microRNA. Using these criteria, four studies identified 12 individual microRNAs. The associated forest plot depicts the sensitivity (95% CIs) and specificity (95% CIs) for each microRNA (Fig. 4). Overall sensitivity and specificity of the 12 individual microRNAs in the diagnosis of glaucoma were 80% [95% CI 73–86%, test of heterogeneity: Q = 15.29, I2 = 28.08%, p-value = 0.17] and 74% [95% CI 63–83%, test of heterogeneity: Q = 28.15, I2 = 60.9%, p-value = 0.00], respectively. These findings suggest that utilizing microRNAs as biomarkers has a high discriminative ability in detecting glaucoma. For glaucoma detection, the slope coefficient was related to a p-value of 0.83 (Fig. 5), indicating no publication bias in our meta-analysis. In addition, the pooled diagnostic odds ratio (DOR) of the 12 individual microRNAs in the diagnosis of glaucoma was 11.2 [95% CI 5.3–24]. Afterward, random-effects models were used to re-analyze the data, and the diagnostic threshold was investigated. The Spearman's correlation coefficient was − 0.130 (p-value = 0.687), indicating that the diagnostic threshold was not responsible for the heterogeneity. Disparities in study techniques, specimen type, endogenous reference, or total sample size may also contribute to the existing heterogeneity. The publication bias of meta-analysis in diagnostic accuracy was investigated using Deeks' funnel plot asymmetry test, depicted in Fig. 6. The most well-known reporting bias is publication bias. It is the outcome of relevant trials being published or not, depending on the type and direction of the results. A paper, for instance, is more likely to be published if the findings are significant. Following subgroup analysis, it has been apparent that the most sensitive microRNA was mir-486-5p from aqueous humor (100%, 95% CI 66–100%). The most specific microRNAs were sampled from the plasma (mir-637, mir-1306-5p, and mir-3159). The data for subgroup analysis are provided in Additional file 1.
Network analysis
microRNA target identification and PPI network construction
The microRNA targets were found using the MirTarBase and were used in the STRING database to construct the final PPI network. The final network had 74 different genes with 120 interactions between them.
Identification of hub genes
Genes were sorted according to their maximal clique centrality (MCC) values, and the top ten genes with the highest MCC value are depicted in Fig. 7. Vascular endothelial growth factor A (VEGF-A) plays a vital role in angiogenesis regulation in normal and abnormal states, such as tumor angiogenesis [21]; AKT1 is a member of the serine/threonine AGC protein kinase family and is associated with cellular growth, metabolism, survival, and proliferation [22]; CXCL12 participates in a series of processes, such as inflammation and leukocyte trafficking, regulation of cell viability, and extracellular matrix remodeling [23]. The HRAS gene is involved primarily in regulating cell division [24].
Network communities
In the benchmark of the four different algorithms for community detection, walk-trap, a widely used algorithm in graph partitioning, achieved the first rank and was applied to the network to find the most important communities. Walk-trap is a hierarchical clustering algorithm based on a random walk iteration, which means members of a community have an overall shorter distance random walk than the rest of the network [25]. Figure 8 shows the community correlation matrix of the mentioned algorithms. The walk-trap results are consistent with the surprise and Markov clustering algorithms, but the spectral algorithm has found a different set of communities. The convergence of walk-trap, surprise, and Markov clustering algorithm communities may potentiate the probability of finding the actual pathobiological processes in glaucoma. The community prioritization ranking for the walk-trap result is provided in Table 2. In total, eight non-overlapping communities are found in the network (Fig. 9). The biggest community has 16 genes, and the smallest has only two. The details of each community, besides each community's gene ontology and biological processes, are provided in Table 2.
Interestingly, after using the CRANK algorithm on these communities, we could show that two small communities could get a higher rank, indicating the importance of community ranking in network analyses. Intriguingly, the eighth community, which contains SFRP1 and DKK2 genes, is involved in the WNT signaling pathway, and recent research findings support the crucial role of this pathway in regulating intraocular pressure [26, 27]. The other simultaneously first-ranked community is the sixth community, which involves the MYO6 and TOM1 genes. The TOM1 gene is involved in vesicular trafficking at the endosome, and mutations in this gene may lead to a combination of brain–eye–muscle anomalies [28]. On the other hand, Opineurin's weakened interaction with these two proteins results in dysregulated autophagosome maturation and clearance of inclusion bodies, eventually leading to glaucoma [29].
Discussion
MicroRNAs are one of the major epigenetic drivers in glaucoma. To the best of our knowledge, this paper is the first attempt to conduct a systematic review, meta-analysis, and genetic network analysis of microRNA profiling in glaucoma patients. The regulatory effects of microRNAs on their target genes may demonstrate some clues to the pathogenesis of glaucoma. Discovering diagnostic microRNAs that meticulously correlate with the presence of the disease could be beneficial in the early diagnosis and screening of glaucoma patients.
The connection between microRNAs and aqueous humor production and absorption, trabecular meshwork, the apoptosis of retinal ganglion cells, and glaucoma have been investigated extensively in human and other animals' eyes, like rats [15, 30,31,32]. It is worth mentioning that the expression patterns of the microRNAs in glaucoma are determined by the disease type and the clinical course of the disease.
The two most frequently dysregulated microRNAs in all studies include miR-486-5p and miR-143-3p. In this study, microRNAs altered in at least two studies were validated to find the corresponding target genes responsible for glaucoma development. AKT1, VEGF-A, CXCL12, and HRAS were the most notable targets.
MiR-486-5p is a potential biomarker for prognosis, diagnosis, and therapeutic target in several cancers [33]. Also, it has been reported to be correlated with disease severity and inflammation in sepsis [34]. Significant downregulation of miR-486-5P compared with the control group was indicated in two studies based on aqueous humor and plasma [16, 35].
On the other hand, it has been demonstrated that MiR-143-3p plays a tumor suppressor role in several tumors [36]; researchers identified the upregulation of this biomarker in patients with glaucoma [16].
Regarding hub genes in the network, except for VEGF-A, little is known about the exact function of the remaining genes. In [37], aqueous humor concentrations of VEGF-A had significantly higher levels in the eyes of patients with neovascular glaucoma. Correspondingly, there is evidence that patients with glaucoma might benefit from anti-VEGF-A therapy [38, 39]. Intriguingly, the results of this work have shed new light on the possible remarkable roles of AKT1, CXCL12, and HRAS genes in the pathogenesis of glaucoma.
Gene community findings are in line with those found in the literature, where the increased IOP causes perforation, deformation, and remodeling of the lamina cribrosa (a porous layer that allows nerve fibers to pass through the eye) and eventually interruption of retrograde transport of essential trophic factors to the retinal ganglion cells [40]; Previous studies indicate that perturbations in axonal transport are one of the earliest findings in the pathogenesis of glaucoma. A recent review of studies undertaken looking at the role of axonal transport in glaucoma has shown that both antegrade and retrograde protein transport are severely disrupted in glaucoma [41]. A causal loop exists as we do not exactly know whether the derangement in axonal transport causes increased IOP or vice versa. As indicated in our study, MYO6 is an unconventional reverse-direction myosin that is required for the degradation of harmful cellular components and the structural integrity of the Golgi apparatus via the p53-dependent pro-survival pathway [42].
Trabecular meshwork (TM) is a fenestrated structure that permits the aqueous humor to be drained from the eye [43]. The production rate of aqueous humor is monotonous; thus, the regulation of IOP is managed by controlling the outflow rate [44]. The increased IOP leads to the extracellular remodeling matrix (ECM) by activating a group of matrix metalloproteinases (MMPs) that instigate outflow rates by opening the pores. The types and expression levels of MMPs vary by the disease; for example, in age-related macular degeneration, the expression of MMP-2 and MMP-9 is significantly reduced [45]. Here we showed that MMP-7 (a matrilysin) and MMP-13 (also known as collagenase-3) are the most important MMPs in glaucoma. It should be mentioned that MMP-7, which is one of the smallest human MMPs, hypothetically can play a dual role in glaucoma; it facilitates ECM remodeling to increase outflow from the anterior chamber; on the other hand, it can cleave cell-surface molecules such as Fas-ligand and pro-TNF-α that induce apoptosis [46].
There is strong evidence for the presence of the canonical WNT signaling pathway in TM. sFRP1, a WNT signaling inhibitor, is significantly expressed in TM and regulated by microRNAs, not DNA methylation. Increased expression of sFRP1 is associated with TM stiffness and subsequently reduced outflow, loss of cell–cell communication, and ultimately apoptosis [47].
Steroid use is one of the risk factors for the pathogenesis of glaucoma [4]. As mentioned in Table 2, PGR and KRT7 in the steroid hormone-mediated signaling pathway are the most affected genes. Previous studies do not provide direct evidence of these genes in steroid-induced glaucoma; hence, further experiments are expected to improve the understanding of steroid-induced glaucoma concerning these two genes. Moreover, according to the literature, corticosteroids can inhibit MMPs and subsequently reduce the adaptation of the ECM to increased pressure [48].
A potential issue we need to consider is the variability of the methods for assessing microRNA profiling, which might compromise their reliability and reproducibility. One of the limitations of the current study is that the microRNAs' sources are different, which could affect the subsequent analyses. MicroRNA releasing mechanisms might also be tissue-specific. Furthermore, it would also be interesting to consider TM regions where the biopsy has been taken because studies have shown that the rate of drainage can be different in TM (high-flow vs. low-flow draining parts) [49]. Moreover, our study findings are solely based on a meta-analysis, and the results might also be influenced by the small sample size, control heterogeneity in case–control studies, or the ethnicity of the glaucoma patients. Future large cohort studies are needed to thoroughly assess the clinical relevance of the obtained results. Despite these limitations, the current study still tries to provide a representative overview of the microRNAs and their target genes involved in the pathogenesis of glaucoma for directing future experiments, providing novel drug targets, and eventually cutting down the immense burden of this sight-threatening disease.
Conclusion
Glaucoma is a multifactorial sight-threatening disease with the complex interaction of genetic and environmental components. Emerging research reveals epigenetics as a key element in developing and preventing the disease. Epigenetic effects are conveyed through pathways regulating apoptosis, ECM protein synthesis, and inflammation. These pathways do not bear equal weight. Using the network analysis, we can estimate the extent of their influences.
Availability of data and materials
The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request.
References
Flaxman SR, Bourne RRA, Resnikoff S, Ackland P, Braithwaite T, Cicinelli MV, et al. Global causes of blindness and distance vision impairment 1990–2020: a systematic review and meta-analysis. Lancet Glob Health. 2017;5(12):e1221–34.
Quigley HA, Broman AT. The number of people with glaucoma worldwide in 2010 and 2020. Br J Ophthalmol. 2006;90(3):262–7.
Tatham AJ, Medeiros FA, Zangwill LM, Weinreb RN. Strategies to improve early diagnosis in glaucoma. Prog Brain Res. 2015;221:103–33.
Weinreb RN, Aung T, Medeiros FA. The pathophysiology and treatment of glaucoma: a review. JAMA. 2014;311(18):1901–11.
Killer HE, Pircher A. Normal tension glaucoma: review of current understanding and mechanisms of the pathogenesis. Eye (Lond). 2018;32(5):924–30.
Kuehn MH, Fingert JH, Kwon YH. Retinal ganglion cell death in glaucoma: mechanisms and neuroprotective strategies. Development. 2005;1:32005.
Gauthier AC, Liu J. Epigenetics and signaling pathways in glaucoma. Biomed Res Int. 2017;2017:5712341.
Sato F, Tsuchiya S, Meltzer SJ, Shimizu K. MicroRNAs and epigenetics. FEBS J. 2011;278(10):1598–609.
Condrat CE, Thompson DC, Barbu MG, Bugnar OL, Boboc A, Cretoiu D, et al. miRNAs as biomarkers in disease: latest findings regarding their role in diagnosis and prognosis. Cells. 2020;9(2):276.
Whiting PF, Rutjes AW, Westwood ME, Mallett S, Deeks JJ, Reitsma JB, et al. QUADAS-2: a revised tool for the quality assessment of diagnostic accuracy studies. Ann Intern Med. 2011;155(8):529–36.
Huang H-Y, Lin Y-C-D, Li J, Huang K-Y, Shrestha S, Hong H-C, et al. miRTarBase 2020: updates to the experimentally validated microRNA–target interaction database. Nucleic Acids Res. 2019;48(D1):D148–54.
Rossetti G, Milli L, Cazabet R. CDLIB: a python library to extract, compare and evaluate communities from complex networks. Appl Netw Sci. 2019;4(1):1–26.
Chen EY, Tan CM, Kou Y, Duan Q, Wang Z, Meirelles GV, et al. Enrichr: interactive and collaborative HTML5 gene list enrichment analysis tool. BMC Bioinform. 2013;14(1):1–14.
Normando AGC, Pérez-de-Oliveira ME, Guerra ENS, Lopes MA, Rocha AC, Brandão TB, et al. To extract or not extract teeth prior to head and neck radiotherapy? A systematic review and meta-analysis. Support Care Cancer. 2022;30(11):8745–59.
Hindle AG, Thoonen R, Jasien JV, Grange RMH, Amin K, Wise J, et al. Identification of candidate miRNA biomarkers for glaucoma. Invest Ophthalmol Vis Sci. 2019;60(1):134–46.
Hubens WHG, Krauskopf J, Beckers HJM, Kleinjans JCS, Webers CAB, Gorgels TGMF. Small RNA sequencing of aqueous humor and plasma in patients with primary open-angle glaucoma. Invest Ophthalmol Vis Sci. 2021;62(7):24.
Liu Y, Wang Y, Chen Y, Fang X, Wen T, Xiao M, et al. Discovery and validation of circulating Hsa-miR-210-3p as a potential biomarker for primary open-angle glaucoma. Invest Ophthalmol Vis Sci. 2019;60(8):2925–34.
Raga-Cervera J, Bolarin JM, Millan JM, Garcia-Medina JJ, Pedrola L, Abellán-Abenza J, et al. MiRNAs and genes involved in the interplay between ocular hypertension and primary open-angle Glaucoma. Oxidative stress, inflammation, and apoptosis networks. J Clin Med. 2021;10(11):2227.
Tamkovich S, Grigor’eva A, Eremina A, Tupikin A, Kabilov M, Chernykh V, et al. What information can be obtained from the tears of a patient with primary open angle glaucoma? Clin Chim Acta. 2019;495:529–37.
Tanaka Y, Tsuda S, Kunikata H, Sato J, Kokubun T, Yasuda M, et al. Profiles of extracellular miRNAs in the aqueous humor of glaucoma patients assessed with a microarray system. Sci Rep. 2014. https://doi.org/10.1038/srep05089.
Ho QT, Kuo CJ. Vascular endothelial growth factor: Biology and therapeutic applications. Int J Biochem Cell Biol. 2007;39(7):1349–57.
Lascaratos G, Chau K-Y, Zhu H, Gkotsi D, Kamal D, Gout I, et al. Systemic PTEN–Akt1–mTOR pathway activity in patients with normal tension glaucoma and ocular hypertension: a case series. Mitochondrion. 2017;36:96–102.
Zhou Y, Cao H-B, Li W-J, Zhao L. The CXCL12 (SDF-1)/CXCR4 chemokine axis: oncogenic properties, molecular targeting, and synthetic and natural product CXCR4 inhibitors for cancer therapy. Chin J Nat Med. 2018;16(11):801–10.
Gripp KW, Lin AE. Costello syndrome: a Ras/mitogen activated protein kinase pathway syndrome (rasopathy) resulting from HRAS germline mutations. Genet Med. 2012;14(3):285–92.
Pons P, Latapy M. Computing communities in large networks using random walks. International symposium on computer and information sciences. Berlin: Springer; 2005.
Webber HC, Bermudez JY, Millar JC, Mao W, Clark AF. The role of Wnt/β-Catenin Signaling and K-Cadherin in the regulation of intraocular pressure. Invest Ophthalmol Vis Sci. 2018;59(3):1454–66.
Lo Faro V, ten Brink JB, Snieder H, Jansonius NM, Bergen AA. Genome-wide CNV investigation suggests a role for cadherin, Wnt, and p53 pathways in primary open-angle glaucoma. BMC Genomics. 2021;22(1):590.
Gwon Y, Kam T-I, Kim S-H, Song S, Park H, Lim B, et al. TOM1 regulates neuronal accumulation of amyloid-β oligomers by FcγRIIb2 variant in Alzheimer’s disease. J Neurosci. 2018;38(42):9001–18.
Shen W-C, Li H-Y, Chen G-C, Chern Y, Tu P-h. Mutations in the ubiquitin-binding domain of OPTN/optineurin interfere with autophagy-mediated degradation of misfolded proteins by a dominant-negative mechanism. Autophagy. 2015;11(4):685–700.
Lagos-Quintana M, Rauhut R, Meyer J, Borkhardt A, Tuschl T. New microRNAs from mouse and human. RNA. 2003;9(2):175–9.
Guo R, Shen W, Su C, Jiang S, Wang J. Relationship between the pathogenesis of glaucoma and miRNA. Ophthalmic Res. 2017;57(3):194–9.
Lu L, Yue J, Xu F, Jablonski M, Williams R. miRNA-mediated gene expression changes in a glaucoma mouse model. Investig Ophthalmol Vis Sci. 2018;59(9):5160.
Ninawe A, Guru SA, Yadav P, Masroor M, Samadhiya A, Bhutani N, et al. miR-486-5p: a prognostic biomarker for Chronic Myeloid Leukemia. ACS Omega. 2021;6(11):7711–8.
Sun B, Guo S. miR-486-5p Serves as a diagnostic biomarker for sepsis and its predictive value for clinical outcomes. J Inflamm Res. 2021;14:3687.
Liu Y, Chen Y, Wang Y, Zhang X, Gao K, Chen S, et al. microRNA profiling in glaucoma eyes with varying degrees of optic neuropathy by using next-generation sequencing. Invest Ophthalmol Vis Sci. 2018;59(7):2955–66.
He Z, Yi J, Liu X, Chen J, Han S, Jin L, et al. MiR-143-3p functions as a tumor suppressor by regulating cell proliferation, invasion and epithelial–mesenchymal transition by targeting QKI-5 in esophageal squamous cell carcinoma. Mol Cancer. 2016;15(1):1–17.
Zhou M, Wang J, Wang W, Huang W, Ding X, Zhang X. Placenta growth factor in eyes with neovascular glaucoma is decreased after intravitreal ranibizumab injection. PLoS ONE. 2016;11(1): e0146993.
Park SC, Su D, Tello C. Anti-VEGF therapy for the treatment of glaucoma: a focus on ranibizumab and bevacizumab. Expert Opin Biol Ther. 2012;12(12):1641–7.
Xiong Q, Li Z, Li Z, Zhu Y, Abdulhalim S, Wang P, et al. Anti-VEGF agents with or without antimetabolites in trabeculectomy for glaucoma: a meta-analysis. PLoS ONE. 2014;9(2): e88403.
Almasieh M, Wilson AM, Morquette B, Vargas JLC, Di Polo A. The molecular basis of retinal ganglion cell death in glaucoma. Prog Retin Eye Res. 2012;31(2):152–81.
Dias MS, Luo X, Ribas VT, Petrs-Silva H, Koch JC. The role of Axonal transport in Glaucoma. Int J Mol Sci. 2022;23(7):3935.
Jung EJ, Liu G, Zhou W, Chen X. Myosin VI is a mediator of the p53-dependent cell survival pathway. Mol Cell Biol. 2006;26(6):2175–86.
Tektas O-Y, Lütjen-Drecoll E. Structural changes of the trabecular meshwork in different kinds of glaucoma. Exp Eye Res. 2009;88(4):769–75.
Acott TS, Kelley MJ, Keller KE, Vranka JA, Abu-Hassan DW, Li X, et al. Intraocular pressure homeostasis: maintaining balance in a high-pressure environment. J Ocul Pharmacol Ther. 2014;30(2–3):94–101.
Weinreb RN, Robinson MR, Dibas M, Stamer WD. Matrix metalloproteinases and glaucoma treatment. J Ocul Pharmacol Ther. 2020;36(4):208–28.
Williams H, Johnson JL, Jackson CL, White SJ, George SJ. MMP-7 mediates cleavage of N-cadherin and promotes smooth muscle cell apoptosis. Cardiovasc Res. 2010;87(1):137–46.
Mao W, Millar JC, Wang W-H, Silverman SM, Liu Y, Wordinger RJ, et al. Existence of the canonical Wnt signaling pathway in the human trabecular meshwork. Invest Ophthalmol Vis Sci. 2012;53(11):7043–51.
Raghunathan VK, Morgan JT, Park SA, Weber D, Phinney BS, Murphy CJ, et al. Dexamethasone stiffens trabecular meshwork, trabecular meshwork cells, and matrix. Invest Ophthalmol Vis Sci. 2015;56(8):4447–59.
Battista SA, Lu Z, Hofmann S, Freddo T, Overby DR, Gong H. Reduction of the available area for aqueous humor outflow and increase in meshwork herniations into collector channels following acute IOP elevation in bovine eyes. Invest Ophthalmol Vis Sci. 2008;49(12):5346–52.
Acknowledgements
Not applicable.
Funding
This research did not receive any specific grant from funding agencies in the public, commercial, or not-for-profit sectors.
Author information
Authors and Affiliations
Contributions
MR and MF conceptualized the study. MSS and HS accomplished literature review, screening, and data extraction. PS performed meta-analysis. MR and MF performed network analysis and community detection. SD was the corresponding author of the article. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Additional file 1: Figure S1.
The Hierarchical Summary Receiver Operating Characteristic (HSROC) curve of circulating miRNAs for the diagnosis glaucoma patients. Figure 2. The scatter plot of the Likelihood ratio of circulating miRNAs for the diagnosis glaucoma patients. Figure S3. Fagan's plot for pre-test and post-probabilities with likelihood ratios. Figure S4. Probability modifying plot. Figure S5. Subgroup analysis of the microRNAs based on the sample type. Figure 6. Sensitivity vs specificity plot for various microRNAs based on sample site subgroups.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated in a credit line to the data.
About this article
Cite this article
Rezaei, M., Faramarzpour, M., Shobeiri, P. et al. A systematic review, meta-analysis, and network analysis of diagnostic microRNAs in glaucoma. Eur J Med Res 28, 137 (2023). https://doi.org/10.1186/s40001-023-01093-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s40001-023-01093-8