Abstract
Background
The epigenetic machinery is frequently altered in acute myeloid leukemia. Focusing on cytogenetically normal (CN) AML, we previously described an abnormal H3K27me3 enrichment covering 70 kb on the HIST1 cluster (6.p22) in CN-AML patient blasts. Here, we further investigate the molecular, functional, and prognosis significance of this epigenetic alteration named H3K27me3 HIST1 in NPM1-mutated (NPM1mut) CN-AML.
Results
We found that three quarter of the NPM1mut CN-AML patients were H3K27me3 HIST1high. H3K27me3 HIST1high group of patients was associated with a favorable outcome independently of known molecular risk factors. In gene expression profiling, the H3K27me3 HIST1high mark was associated with lower expression of the histone genes HIST1H1D, HIST1H2BG, HIST1H2AE, and HIST1H3F and an upregulation of genes involved in myelomonocytic differentiation. Mass spectrometry analyses confirmed that the linker histone protein H1d, but not the other histone H1 subtypes, was downregulated in the H3K27me3 HIST1high group of patients. H1d knockdown primed ATRA-mediated differentiation of OCI-AML3 and U937 AML cell lines, as assessed on CD11b/CD11c markers, morphological and gene expression analyses.
Conclusions
Our data suggest that NPM1mut AML prognosis depends on the epigenetic silencing of the HIST1 cluster and that, among the H3K27me3 silenced histone genes, HIST1H1D plays a role in AML blast differentiation.
Graphical abstract

Similar content being viewed by others
Background
Acute myeloid leukemias (AMLs) are a heterogeneous group of severe hematological malignancies that arise through the acquisition of oncogenic mutations by hematopoietic progenitor cells. Patient prognosis mainly relies on the given treatment, the outcome of which depends on cytogenetics and molecular alterations. Cytogenetically normal (CN) AML patients are usually assigned to an intermediate prognosis group that can be further subdivided through the detection of mutations in a growing number of genes [1]. Mutations in the nucleophosmin 1 (NPM1) gene are the commonest molecular lesions occurring in ≥ 50% of cases with cytogenetically normal acute myeloid leukemia (CN-AML). NPM1 mutations result in the generation of a nuclear export signal causing the delocalization of the protein from the nucleoli to the cytoplasm [2]. Analyses of large numbers of patients have shown that NPM1 mutations are associated with a relatively favorable prognosis, which can be mitigated by internal tandem duplications (ITD) of the tyrosine kinase receptor Fms-like tyrosine kinase 3 (FLT3) and mutations in DNA-methyl transferase 3A (DNMT3A). Indeed, in NPM1mut CN-AML, FLT3ITD mutation, and/or mutation in DNMT3A predict an increased risk of relapse and poorer outcome [3], [4]. Recent reports shed light on the importance of epigenetic deregulations that affect the epigenome and gene transcription in AML pathogenesis [5]. These deregulations are the consequence of numerous alterations found in genes encoding multiple classes of epigenetic proteins as DNA methylation and histone modification enzymes [6].
In addition to these mutations, recent reports underlined the importance of histone genes themselves in cancer onset. For example, the major histone cluster 1 (HIST1), encoding the vast majority of the redundant core and linker histones [7], has been found partially deleted in ALL [8] and deregulation of its expression is associated with breast cancer invasiveness [9]. Moreover, we previously described a new alteration that affects the HIST1 cluster in AML. This alteration consists in a marked histone H3 lysine 27 tri-methylation (H3K27me3) enrichment encompassing 70 kb of the HIST1 cluster, affecting histone genes and associates with the NPM1 mutation and a better leukemia-free survival [10]. In this study, we reported the clinical importance of this newly identified epigenetic alteration, called the H3K27me3 HIST1 mark, in relation to other known mutations and its functional consequences on the biology of CN-AML leukemic cells. We revealed that H3K27me3 HIST1 status and histone mRNA and protein levels define clinically and biologically different subgroups of NPM1mut CN-AML suggesting their importance in AML pathogenesis.
Results
H3K27me3 level on HIST1 locus is associated with better survival
To characterize the H3K27me3 HIST1 mark, we performed H3K27me3 ChIP-qPCR on samples obtained from 46 de novo CN-AML patients included in GOELAMS multicenter clinical trials LAM2006IR (NCT00860639) or LAM2007SA (NCT00590837). H3K27me3 level was determined at five HIST1 genomic locations that are representative of the H3K27me3 HIST1 signature, as described previously [10]. Heatmap showing H3K27me3 HIST1 gene enrichment confirmed the variation of H3K27me3 HIST1 level among CN-AML patients (Fig. 1a). The average of the five normalized H3K27me3 HIST1 values was calculated and this index showed a clear segregation of the H3K27me3 HIST1low and H3K27me3 HIST1high patients. With a cut-off value at 15, approximately 55% of CN-AML samples displayed an H3K27me3 HIST1 enrichment mark (Fig. 1b).
Analysis of H3K27me3 HIST1 level in CN-AML patients. a Heatmap of normalized H3K27me3 enrichment value obtained by H3K27me3 ChIP-qPCR on the indicated HIST1 genes. Enrichment was calculated as the percentage of bound/input and double normalized with HOXD4 and GAPDH. Each column represents a patient sample (n = 46) sorted by unsupervised hierarchical clustering. The separation between H3K27me3 HIST1low and H3K27me3 HIST1high groups is indicated by a dotted lane. Yellow square: mutation for the indicated gene. b Bimodal distribution of the patients according to H3K27me3 HIST1 mean level, obtained from the H3K27me3 enrichment values of the 5 HIST1 regions analyzed. In blue, patients with an H3K27me3 HIST1 mean value below 15 (H3K27me3 HIST1low, n = 21), in red, patients with a mean value above 15 (H3K27me3 HIST1high, n = 25). c OS and EFS in CN-AML patients according to H3K27me3 HIST1low and H3K27me3 HIST1high status
There was no association of H3K27me3 HIST1 status with age, gender, FLT3ITD, DNMT3A, IDH1, or CCAAT/enhancer-binding protein alpha (CEBPA) mutations (Fig. 1a and Additional file 1: Table S1). However, we noted that 25/33 (75.7%) of the NPM1mut samples were also H3K27me3 HIST1high (Fig. 1a and Additional file 1: Table S1) confirming our first observation [10]. In addition, we observed a significant association between H3K27me3 HIST1high and the presence of IDH2R140 mutation (36% vs. 4.7%, P = .01) (Fig. 1a and Additional file 1: Table S1).
H3K27me3 HIST1high patients had a better overall survival (OS) and event-free survival (EFS) than H3K27me3 HIST1low patients, with a median OS of 50.9 months versus 14.6 months (HR, 2.5 [1.5–5.5]; P = .02) and a median EFS of 37 months versus 9 months (HR, 2.7 [1.3–5.8]; P = .008) (Fig. 1c). The survival gain was independent in multivariate analyses taking age and NPM1wt/FLT3ITD status into account (Additional file 1: Table S2). These observations confirmed in an independent cohort the previously reported association of H3K27me3 HIST1high with a better prognosis [10].
H3K27me3 level on HIST1 locus is an independent biomarker predicting survival of NPM1mut CN-AML
NPM1-mutated AMLs represent a distinct clinical and biological entity in the World Health Organization (WHO) classification, commonly associated with a better prognosis [3]. We used the biological material obtained from 25 NPM1mut samples previously analyzed [10], and 78 samples either provided by the GOELAMSTHEQUE (n = 33), or the IPC/CRCM tumor bank (n = 45) (Fig. 2a). H3K27me3 status of the 78 new samples was analyzed (Additional file 2: Figure S1) and revealed that 75% of NPM1mut CN-AMLs were H3K27me3 HIST1high. The NPM1mut H3K27me3 HIST1high subgroup of patients was not enriched with DNMT3A or FLT3ITD mutations, the most frequently NPM1mut co-occurring alterations [11] (Table 1 and Additional file 2: Figure S1), but IDH2R140 was significantly overrepresented in this subgroup in comparison with the NPM1mut H3K27me3 HIST1low group (27.6% vs. 7.6%, P = .05). Interestingly, H3K27me3 HIST1high leukemic cells had a significantly lower CD34 expression than their H3K27me3 HIST1low counterparts (CD34 mean expression, 10.3% vs. 35%, P = 0.005) (Table 1). To further explore a potential association between CD34 expression and the presence of H3K27me3 HIST1 mark, we selected four patient samples (two in each H3K27me3 HIST1 group), containing both CD34low and CD34high blast populations. We next analyzed the H3K27me3 HIST1 level in CD34low and CD34high fluorescence-activated cell sorted (FACS) blasts. As shown in Fig. 2b and Additional file 2: Figure S2, H3K27me3 HIST1 status (high or low), in both CD34low and CD34high blast fractions, was similar to the bulk population, indicating that H3K27me3 HIST1high occurs independently of CD34 level of expression.
Analysis of H3K27me3 HIST1 level in CN-AML patients with NPM1 mutation. a Consort diagram showing number and overlap of NPM1mut AML samples analyzed in the different experiments. b Analysis of H3K27me3 HIST1 status in CD34low and CD34high blast populations. The left panel presents an H3K27me3 HIST1high patient and the right panel an H3K27me3 HIST1low patient. For each panel are shown: the gating strategy for separating blast cells according CD34 expression level (upper left), the H3K27me3 HIST1 status on the bulk population (upper right), on CD34low blast population (lower left) and on CD34high blast population (lower right). c OS and EFS in NPM1mut CN-AML patients according to H3K27me3 HIST1 status (n = 103). d OS and EFS in NPM1mut/FLT3wt CN-AML patients according to H3K27me3 HIST1 status (n = 53). Statistical significance was estimated using the log-rank test
Next, we analyzed the prognosis impact of the H3K27me3 HIST1 alteration. NPM1mut H3K27me3 HIST1high patients had a better OS and EFS than NPM1mut H3K27me3 HIST1low patients (median OS, 38.3 vs. 15.7 months; HR, 2 [range, 1.0–3.0]; P = .03; median EFS, 20.9 vs. 10.6 months; HR, 2.7 [range, 1.3–5.7]; P = .01) (Fig. 2c). In multivariate analysis, taking age and FLT3ITD/DNMT3Amut into account, H3K27me3 HIST1high remained significantly associated with a favorable EFS suggesting that the prognostic significance of the H3K27me3 HIST1 signature is independent of other known molecular alterations (Table 2).
Finally, we studied the impact of H3K27me3 HIST1 mark in the NPM1mut/FLT3wt subgroup, which is classically associated with a favorable prognosis [12]. H3K27me3 HIST1high patients (n = 36) displayed better OS and EFS than H3K27me HIST1low patients (n = 17), (median OS, 111.6 months vs. 23.2 months; P = .03; median EFS, 44.1 months vs. 13.9 months; P = .01, for H3K27me3 HIST1high and H3K27me3 HIST1low, respectively) (Fig. 2d).
Our results suggest that H3K27me3 HIST1 status is an independent epigenetic marker that identifies patients with a poor outcome within the NPM1/FLT3wt group of patients.
Histone mRNA expression is anti-correlated to H3K27me3 HIST1 level and predicts NPM1mut CN-AML patient outcome
To analyze the anti-correlation of histone mRNA level and the presence of the repressive H3K27me3 mark, we selected three histone genes; HIST1H1D, HIST1H2BG and HIST1H2BH, spread over the H3K27me3 HIST1 islet and associated with clinical outcome in public data (see below), and measured their mRNA levels. Expression of these three genes was lower in H3K27me3 HIST1high patients (n = 34) than in H3K27me3 HIST1low patients (n = 13) (Fig. 3a).
Gene expression from the HIST1 cluster impacts on survival in NPM1mut CN-AML. a mRNA analyses of HIST1 genes in the IPC cohort of NPM1mut CN-AML patients. HIST1H1D (n = 46), HIST1H2BH (n = 45), and HIST1H2BG (n = 47) mRNA levels were determined by Taqman, according to H3K27me3 HIST1 status. Gene expression was normalized on the average of two housekeeping genes (PGK1 and PPIA). Statistical significance was estimated using Mann Whitney test * p < 0.05; **p < 0.005 ***p < 0.0001. b Influence of histone gene expression on OS by univariate analysis. Only histone genes covered by the H3K27me3 HIST1 mark described in [10] were analyzed. c Survival analyzed according to the expression of HIST1H1D, HIST1H2BG, and HIST1H2BH mRNA on independent cohorts of patients (TGCA and Metzeler). Patients were split into two groups according to the expression of HIST1H1D, HIST1H2BH and HIST1H2BG genes using a Cox regression prognosis model. Statistical significance was estimated using the log-rank test
We next asked whether expression of these genes, as a consequence of H3K27me3 mark, was associated with patient survival. Given the small size of our cohort, we analyzed HIST1 gene expression in two published cohorts with publicly accessible clinical and mRNA expression data: TCGA [13] and Metzeler [14]. NPM1mut CN-AML patients, within these two cohorts, were identified by using a published gene expression signature that predicts the NPM1 mutational status [15] (see Additional file 3: Supplemental methods). Association of histone expression with survival was first tested for each of the 11 histone genes covered by the H3K27me3 HIST1 mark. This highlighted three histone genes, HIST1H1D, HIST1H2BG, and HIST1H2BH, for which a high level of expression was associated with a poor outcome (P = .004, .015 and .044 respectively, Fig. 3b). Then, we tested this 3-HIST1-mRNA signature in univariate analysis; 3-HIST1-mRNAlow patients had a favorable OS with a median OS of 17.7 months versus 9.6 months (HR = 1.66, range, 1.13–2.42, P = .009) (Fig. 3c). Multivariate analyses showed that the 3-HIST1-mRNAlow status was associated with a better prognosis (HR = 1.60, range 1.60–2.31, P = .01), independently of other prognosis markers including age, FAB classification and FLT3 status (Additional file 1: Table S3).
These results show that H3K27me3 HIST1high is associated with a lower expression of histone genes, and that 3-HIST1-mRNAlow signature defines a NPM1mut AML patient group with a better outcome.
Gene expression profiling associated with H3K27me3 HIST1 high identifies a “mature like” phenotype
We next characterized the gene expression profile (GEP) of H3K27me3 HIST1high samples (n = 16) in comparison to H3K27me3 HIST1low samples (n = 11) from the IPC cohort (see Additional File 1: Table S4 for patient clinical characteristics). Eighty-one genes were differentially expressed (p < .05, fold-change > 1.5) between the two groups, 58 being up- and 23 being downregulated in the H3K27me HIST1high group (Fig. 4a and Additional file 4: Table S5). Analysis of enhancer of zeste homolog 2 (EZH2) and suppressor of zeste 12 (SUZ12) expression revealed that the two groups, H3K27me3 HIST1high and H3K27me3 HIST1low, equally expressed PRC2 components (Fig. 4b). GSEA identified, in H3K27me3 HIST1high patients, genes associated with myelomonocytic differentiation such as immune or inflammatory responses (Fig. 4c and Additional file 4: Table S5). Downregulated genes in these patients belong to cell cycle and chromatin regulation categories, including histone genes from the HIST1 cluster (Fig. 4c and Additional file 4: Table S5). Using qPCR, we confirmed higher expression of three genes involved, in mature granulocyte functions, CYBB, FCN1, and CLEC4A [16], [17], [18] in H3K27me3 HIST1high patients (Fig. 4d). H3K27me3 level at the promoter of these genes was identical between H3K27me3 HIST1low and H3K27me3 HIST1high patients (Additional file 2: Figure S3) suggesting that the variation of expression was indirectly affected by H3K27me3 HIST1high.
H3K27me3 HIST1high is associated with a myelomonocytic Gene Expression Profile. a Volcano plot displaying differentially expressed genes between H3K27me3 HIST1high (n = 16) and H3K27me3 HIST1low patients (n = 11). The blue dots represent the histone genes covered by the H3K27me3 mark. b EZH2 and SUZ12 levels of expression obtained from our micro array data were analyzed in H3K27me3 HIST1low (n = 11) and H3K27me3 HIST1high (n = 16) patients. c GSEA (gene set enrichment analysis) of H3K27me3 HIST1high samples. d Expression of three genes associated with granulocytic functions according to H3K27me3 HIST1 status. CYBB (cytochrome B-245 beta chain); FCN1 (Ficolin 1); CLEC4A (C-type lectin domain family 4 member A). Data are represented in relative expression to HPRT. e–f Patients from TGCA and GSE 61804 cohorts were separated according to the 3-HIST1 mRNA signature in mRNAlow (n = 114) and mRNAhigh (n = 79) patients. e Myelomonocytic CYBB, FCN1, CLEC4A, ITGAM (integrin subunit alpha M) and f leukemic stem cell SOCS2 (suppressor of cytokine signaling 2), CDK6 (cyclin dependent kinase 6), LAPTM4B (lysosome-associated protein transmembrane-4β), and NGFRAP1 (nerve growth factor receptor-associated protein 1) gene expression was analyzed. Statistical significance was estimated using Mann Whitney test * p < 0.05; **p < 0.005 ***p < 0.0001. NS non-significant
To further validate the relation between low mRNA level of HIST1 genes and the expression of granulocytic markers, we tested mRNA expression of myelomonocytic maturation genes (CYBB, FCN1, CLEC4, and ITGAM) in the TCGA and Metzeler cohorts of patients stratified with the previously defined 3-HIST1-mRNA signature. The 3-HIST1-mRNAlow patient group overexpressed the differentiation genes in comparison to the 3-HIST1-mRNAhigh one (Fig. 4e), thus corroborating our previous observation (Fig. 4d). Reciprocally, genes such as SOCS2, CDK6, LAPTM4B, and NGFRAP1, which were described as associated with a leukemic stem cell signature [19], were less expressed in the 3-HIST1-mRNAlow patient group (Fig. 4f).
Taken together, these results suggest that HIST1 mRNA downregulation by the H3K27me3 HIST1high mark is associated with a more differentiated phenotype related to a committed state of leukemic cells.
The histone linker H1-3 is poorly expressed in H3K27me3 HIST1 high CN-AML patients
To study the role of histones on AML clinical and biological features, we looked at the effect of H3K27me3 HIST1 epigenetic silencing on histone protein level. First, we looked at the proportions of total histones and of each histone subtype (Additional file 2: Figure S4) in chromatin-bound fractions extracted from a series of 12 patient samples (six in each group) using intensity-based absolute quantification (iBAQ) approach. Normalized quantities of total linker histone H1 and core histones H2A, H2B, H3, and H4 were similar in both H3K27me3 HIST1high and H3K27me3 HIST1low patients (Additional file 2: Figure S5), suggesting that H3K27me3 HIST1high status did not globally modify histone protein abundance. Then, we decided to analyze specifically the histone linker H1-3, encoded by HIST1H1D, because its mRNA level is affected by H3K27me3 HIST1 status (Fig. 3a) and it is the leading gene for the mRNA signature (Fig. 3b). In addition, contrary to other histone subtypes (i.e., H2A, H2B, H3, and H4), H1 histone subtypes are heterogeneous in amino acid composition [20], which probably reflects a subtype-specific function. Indeed, when looking at the H1 subtype abundance, we observed that the H1-3 subtype was decreased in the H3K27me3 HIST1high group (normalized iBAQ value (Log2) = 6.09 vs 4.74; P = .04), whereas the other H1 subtypes, H1-5, H1-2, H1-4, H1-F0, and H1-FX, were unaffected (Fig. 5a). These results are consistent with the specific HIST1H1D mRNA expression decrease observed in AML samples harboring the H3K27me3 HIST1 mark. We confirmed the lower expression of H1-3 observed in H3K27me3 HIST1high group in comparison to H3K27me3 HIST1low group of patients by Western blot using pan H1 and specific H1-3 antibodies (Fig. 5b).
H1-3 protein expression is lower in H3K27me3 HIST1high patients. a Abundance of histone H1 subtypes. Results of each H1 subtype are expressed in iBAQ (Log2) adjusted on the median of the total protein extracted and normalized to total histone amount by patients. Statistical significance was estimated using unpaired t test, (*) p < 0.05. NS non significant. b Immunoblots of H3, H1, and H1-3 on patient chromatin bound extracts. The arrow points to the specific band for H1-3. Ratio of H1-3/total-H1 is calculated using Mean Signal Intensity on imageJ software
In conclusion, as a consequence of the presence of an H3K27me3 islet, NPM1mut CN-AML H3K27me3 HIST1high patients express low level of H1-3.
H1-3 knockdown confers a more mature phenotype in AML cell lines
We hypothesized that the better prognosis of H1-3 low AML may be related to a gain of differentiation and a loss of stem cell features of the AML blasts. Thus, we tested the consequences of H1-3 knockdown (KD) on differentiation potential of the NPM1-mutated OCI-AML3 cell line, which also harbor the DNMT3AR882C mutation [21]. Efficiency and specificity of our KD were assessed by testing mRNA levels of the different H1 subtypes (Fig. 6a and Additional file 2: Figure S6a and S6b) and by measuring H1-3 protein level, after H1-3 KD induction (Fig. 6b and Additional file 2: Figure S6c and S6d). Consequences of H1-3 KD on differentiation were evaluated upon all-trans retinoic acid (ATRA)-treatment. Albeit no increase in CD11b was observed upon H1-3 KD alone, addition of ATRA (0.5 μM and 1 μM) induced a significant increase in CD11b expression, with a marked increase at 0.5 μM (22.6 ± 2.5% vs. 41 ± 4.3%; P = .008) (Fig. 6c and Additional file 2: Figure S7a and S7b) and a significant increase in the proportion of the double positive CD11b/CD11c population (29.8 ± 1.3% vs. 42.5 ± 2.1%; P = .003) (Fig. 6d). In addition, morphological and quantification analyses showed that cytoplasmic granules, which reflect the beginning of a maturation process appeared upon H1-3 KD after 96 h of ATRA-treatment (0.5 μM), (Fig. 6e, f and Additional file 2: Figure S8). Finally, mRNA expression levels of two ATRA-induced genes, CYBB and ITGAM, were tested in H1-3 KD condition under ATRA-treatment; H1-3 downregulation increased the amplitude of ATRA-induced upregulation of these two genes (Fig. 6g). To test whether this ATRA-sensitization was dependent of the presence of NPM1mut or DNMT3AR882C, we performed H1-3 KD in the NPM1wt and DNMT3Awt myeloid U937 cell line. Interestingly, H1-3 KD increased the proportion of the double positive CD11b/CD11c in 0.1 μM ATRA-treated U937 cells and induced morphological changes (Fig. 6h). Altogether, these results suggest that downregulation of histone H1-3 induces ATRA-sensitization independently of NPM1 and DNMT3A mutations.
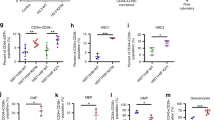
H1-3 KD promotes granulocytic differentiation in ATRA-treated AML cell lines. a qPCR expression analysis of the indicated histone H1 genes in shCtrl and shH1-3 (KD#1) without (Dox-) or with 6-days-induction of doxycycline (Dox+). Data represent three independent Dox inductions. Results are normalized on HPRT and expressed in fold change (FC) between Dox+ and Dox− conditions. b Immunoblot of H1-3, H1, and H3 in chromatin-bound extracts from shCtrl and KD#1 upon (+) or not (−) doxycycline (Dox). c, d Cell surface analysis of granulocytic markers consecutive to H1-3 KD in OCI-AML3 clones. c Percentage of CD11b positive cells in shCtrl and KD#1 upon doxycycline induction (Dox+) or not (Dox−) without ATRA (wo ATRA) or with 72 h of ATRA-treatment (0.5 μM or 1 μM). d Percentage of CD11b-CD11c cells upon 72 h of ATRA-treatment (0.5 μM). Data represent three independent experiments. Statistical significance was estimated using Mann Whitney test *p < 0.05; **p < 0.005. NS non-significant. e, f Cytological analysis of shCtrl and KD#1 clones upon doxycycline and after 96 h of ATRA treatment (0.5 μM). e May–Grünwald Giemsa coloration. Black arrows are pointing cytoplasmic azurophilic granules. f Quantification of cells with cytoplasmic granules. Results are presented as a percentage of positive cells (more than two granules). Statistical significance was estimated using Mann–Whitney test *p < 0.05; **p < 0.005. g Expression analysis of CYBB and ITGAM in untreated OCI-AML3 and in ATRA-treated (1 μM) KD#1 without or with doxycycline induction. Gene expression was normalized to two housekeeping genes (PGK1 and PPIA). Data represent three independent experiments. Statistical significance was estimated using T-test (one-tailed p value). h U937 cells were stably infected by a doxycycline inducible shCtrl (U-shCtrl) or shH1-3 (U-KD#1 and U-KD#2). Left panel: percentage of CD11b-CD11c cells upon 72 h of ATRA-treatment (0.1 μM). Level of H1d in the different conditions was measured by immunoblot. Right panel: May–Grünwald Giemsa staining of KD#1 clone with or without doxycycline upon 72 h hours of ATRA-treatment. Data represent two independent experiments. Statistical significance was estimated using unpaired Two-tailed t test * p < 0.05; ***p < 0.001
Discussion
Aberrant epigenetic patterns in leukemia have been described but determination of their importance in leukemia onset and progression is still lacking [22]. We have studied here the clinical and biological consequences of an epigenetic alteration of the HIST1 locus, the H3K27me3 HIST1 signature, which we previously described in CN-AML [10]. We first confirmed the presence of H3K27me3 HIST1high on 55% of CN-AML and 75% of NPM1mut CN-AML in an independent CN-AML cohort. H3K27me3 HIST1high impacts on the survival of CN-AML and NPM1mut CN-AML patients, and confers a better survival independently of other molecular markers. While H3K27me3 HIST1high is not associated with FLT3ITD or DNMT3A mutations, IDH2R140Q is overrepresented in H3K27me3 HIST1high patients (28% vs 7.6%); this is consistent with data highlighting the functional relationship between IDH2R140Q mutant expression and histone hypermethylation [23]. Although we do not know the mechanisms underlying the H3K27me3 HIST1high signature, this focal high level of H3K27me3 may reflect a maintained activity of EZH2 previously shown to be associated with favorable outcome in AML [24]. Our data provide the first evidence that detecting an aberrant epigenetic pattern in addition to gene mutations has clinical interest and could guide therapeutic choices.
Detection of our epigenetic biomarker in AML demonstrates that H3K27me3 HIST1high status is invariably associated with a favorable outcome and may reveal a less aggressive disease. The less aggressive form of AML associated with H3K27me3 HIST1high is supported by its anti-correlated stem cell signature, shown to be associated with an adverse prognosis [19]. It is also supported by its inflammatory and immune function signature, which has been associated with a better clinical response to dexamethasone in NPM1 CN-AML [25]. However, although CD34low AMLs, characterized by a leukemic stem cell arrested at a precursor-like stage [26], are enriched in the H3K27me3 HIST1high group, we could not find H3K27me3 HIST1high exclusively on CD34low blasts. This suggests that H3K27me3 HIST1high marks a more mature leukemia independently of the level of CD34 expression.
The main consequence of the H3K27me3 accumulation at the HIST1 locus may be the downregulation of the histone genes affected by the repressive epigenetic mark. Levels of replication-dependent histone gene expression may reflect aggressiveness of the disease and may have a survival impact. Indeed, the major histone gene cluster has been described as one of the most upregulated across breast cancer progression [9]. Our study highlighted a peculiar pattern of a lower histone gene expression and defined a three-HIST1 mRNA signature, containing three histone genes (HIST1H1D, HIST1H2BH, and HIST1H2BG) that are directly affected by the H3K27me3 enrichment and can predict survival on CN-AML patients. Interestingly, one of the most affected histone genes in our AML data encodes the histone linker H1-3. High level of H1.3 has been previously linked to cancer aggressiveness, as its overexpression is associated with malignant ovarian adenocarcinoma [27] and with a poor pancreatic ductal adenocarcinoma survival [28]. Given its tight link with differentiation blockage in cancer cells [29] and given that H1 subtype individual KDs induce changes, notably in genes involved in cell cycle and chromatin regulation [30], diminution of H1.3 level appears to be a valid effector of the less aggressive H3K27me3 HIST1high AML phenotype. If histone linker H1 subtypes have a well-described redundant roles in maintaining nucleosome architecture and regulating transcription [31], it becomes evident that H1 subtypes have also subtype-specific functions reflected by selective genomic binding and regulation of chromatin organization [20, 32]. H1 subtypes would influence chromatin compaction at definite loci affecting specific gene expression in a tissue specific manner. Thus, it is tempting to speculate that in our H3K27me3 HIST1high model, low H1-3 level would influence the differentiation state of the cells due to a specific change in chromatin targeting. In our cellular models, KD of H1-3 does not induce differentiation but sensitizes the cells to ATRA-treatment. Interestingly, this sensitivity is independent of DNMT3A activity, as the two cell line models (OCI-AML3 DNMT3Amut and U937 DNMT3Awt) are both sensitive to ATRA-treatment after H1-3 KD. In the light of previous reports, which documented an interaction between H1 proteins and DNMT3B [33, 34], one possibility is that DNMT3B, although not found mutated in AML, may be an influential epigenetic partner in AML and in ATRA sensitivity together with H1-3.
ATRA is successfully used for the treatment of acute promyelocytic leukemia (APL) by inducing terminal granulocytic differentiation of APL blasts [35]. To a lesser extent, ATRA can also induce differentiation in non-APL cells, in particular in AML with IDH1/IDH2 mutations [36] or induce cell death in NPM1mut AML when used in combination with Arsenic trioxide [37], [38]. Interestingly, ATRA sensitization of non-APL cells has been induced by targeting epigenetic enzymes: inhibition of histone deacetylase (HDAC) with valproic acid [39] or inhibition of the histone demethylase LSD1/KDM1A [40] or by targeting SUMOylation [41]. In the same line, our study suggests that epigenetic modifications induced by H1-3 targeting could prime AML cells toward differentiation, revealed by ATRA-sensitivity, which may explains the more mature phenotype found in H3K27me3 HIST1high leukemia and suggests that ATRA could be an efficient differentiating agent in AML with low H1-3 expression.
Conclusions
We showed that epigenetic silencing of a part of the HIST1 locus by the H3K27me3 mark is associated with a better outcome and a mature gene expression profile in NPM1mut CN-AML and we observed an important role of histone linker H1-3 expression in AML blast cell differentiation. Our study pinpoints the H3K27me3 HIST1 mark and HIST1H1D gene as two biomarkers potentially useful to stratify patient prognosis and defines targets that can be considered when developing epidrugs.
Methods
Patient samples
Blast cells were separated from blood or marrow samples through density-gradient (ficoll) separation, and stored in liquid nitrogen. Cryopreserved samples, with at least 70% of blasts, were collected from AML samples stored at Institut Paoli-Calmettes (IPC) Tumor Bank or at the Groupe Ouest Est d’Etude des Leucémies Aiguës et autres Maladies du Sang repository (GOELAMSTHEQUE). GOELAMS samples were extracted from multicenter clinical trials LAM2006IR (NCT00860639) or LAM2007SA (NCT00590837). All patients received conventional induction chemotherapy consisting in daunorubicine (DNR) and aracytine (ARAC) with or without Mylotarg in the LAM2006IR trial for patients < 60 years [42] and idarubicine (IDA) and ARAC with or without lomustine in the LAM2007SA trial for patients ≥ 60 years [43]. Informed consent was provided by all patients according to the Declaration of Helsinki and subjected to ethical institutional review board approval.
ChIP-qPCR
Chromatin immunoprecipitation (ChIP) was performed as previously described [10]. Briefly, frozen samples were thawed, washed twice in phosphate-buffered saline (PBS), and chromatin was extracted with TRIS buffer pH 8, 0.25% Triton. Samples were sonicated to obtain DNA fragments of 300–600 base pair (Bioruptor PICO) and chromatin was immunoprecipated with an anti-H3K27me3 antibody (Abcam #6002). After immunoprecipitation, DNA was purified with the I-Pure kit (Diagenode). Quantification of ChIPed DNA was performed by real-time PCR using the SsoADV Univer SYBR Green Supermix (Bio-Rad) and detected with a CFX96 Real-Time PCR Detection System (Bio-Rad). IgG control “cycle over the threshold” Ct values were subtracted to input or IP Ct values and converted into bound value by 2(−(IP Ct or input Ct- IgG IP Ct)). Data are expressed as percent of bound/input. For each HIST1 region, H3K27me3 ChIP signal was double normalized with ChIP signal obtained at a genomic location invariably enriched with H3K27me3 (HOXD4) and a genomic region depleted of H3K27me3 (GAPDH). Heatmaps were performed with gene-e software (Broad institute). Hierarchical clustering was done using Euclidian distance.
Histone gene nomenclature
The large HIST1 gene cluster on human chromosome region 6p22 is 2.1 Mb which contains 55 histone genes. Five genes (HIST1H1A-E) encode the canonical somatic histone linkers H1 (H1-1, H1-5, H1-2, H1-3, H1-4 respectively) while sets of 10–20 genes encode each of the core histone proteins (H2A, H2B, H3, and H4). Each of these genes is translated into a unique mRNA with distinct 5′ and 3′ extremities, as well as slight nucleotide changes in the coding region.
RT-qPCR analysis
Total RNA was isolated from patient samples using the RNeasy mini Kit (Qiagen). RNA was treated with RNase-free DNase set (Qiagen) to remove contaminating genomic DNA. The cDNA was synthesized using the Transcriptor High Fidelity cDNA Synthesis Kit (Roche Applied Science), quantified by Power SYBR Green (Roche Applied Science) or TaqMan qPCR. For SYBR green analyses, signals were detected with a CFX96 Real-Time PCR Detection System (Bio-Rad). Primers used for gene amplification were designed using Primer3 software and are listed in the table below. Relative expression levels were determined by the delta Ct method and expression level of HPRT was used for normalization. For TaqMan method, PCR Master Mix (Thermo Fisher) was used and signal detected with 7500 Fast Real-Time PCR System (Applied Biosystem). Probes used for TaqMan analyses were HIST1H1D: Hs00271187_s1; HIST1H2BG: Hs00374317_s1; HIST1H2BH: Hs00374322_s1; PGK1: Hs00943178_g1; PPIA: Hs04194521_s1 (Thermo Fisher). Relative expression levels were determined by the delta Ct method, taking the mean expression level of PKG1 and PPIA for normalization [44].
Gene expression profiling
RNA expression profiling of NPM1mut CN-AML was done with Affymetrix Human gene ST 2.0 and Human Genome U133 Plus 2.0 DNA microarrays (see Supplemental data). Microarray data are accessible under the accession number E-MTAB-6997.
Protein analysis
Cellular fractionation was done with the subcellular protein fractionation kit (Thermofisher). Mass spectrometry procedures are explained in supplemental data. Immunoblots were done as previously described [45]. Antibodies used were anti-Histone H1.3 (Abcam, ab24174, 1/750), anti-H1 (Active Motif, #39707, 1/2000), and anti-H3 (Active Motif, #39163, 1/10000).
Flow cytometry
Flow cytometry analyses were done using a BD-LSRII cytometer and analyzed using BD-DIVA Version 6.1.2 software (BD Biosciences). Antibodies used were CD11B-PE (Mac-1), 3:100, Beckman Coulter; CD11B-APC (M1/70), 1:500, eBioscience; CD11C-PeCy7 (BU-15), 3:100, Beckman Coulter; DRAQ7™, 1:400; Biostatus, CD34-PeCy7 (#343516) 1:33, Biolegend.
Cell culture, shRNA lentiviral infection, stable H1-3 knockdown, and treatments
The OCI-AML3 and U937 cells were grown in minimum essential medium alpha (MEMα) supplemented with 20% fetal bovine serum or Roswell Park Memorial Institute (RPMI) medium supplemented with 10% fetal bovine serum respectively. H1-3 knockdown (KD) was achieved using doxycycline-induced Dharmacon™SMARTvector™ short hairpin RNA (V3SH7669-229784413, shRNA-1 and V3SH7669-228676834, shRNA-2). A non-silencing shRNA (piSMART VSC10730) was used as a control (shCtrl). Cells containing the SMARTvector™ were sorted using an ARIAIII cytometer based on red fluorescent protein (RFP) expression and selected on puromycin (2 μg/mL). KD of H1-3 was obtained by the addition of doxycycline (2 μg/mL) during 5–7 days. All-trans-retinoic acid (ATRA; Sigma) was dissolved in dimethylsulfoxyde (DMSO) at 10 mM.
Statistical analyses
Statistical analyses were done using R software (version 2.15.2) (The Comprehensive R Archive Network. http://www.cran.r-project.org/) and Graph Pad Prism (Graph Pad Software, San Diego, CA, USA) and the significance of the differences between groups was determined via unpaired T test, Mann–Whitney test, or exact Fisher test. Data were presented as the median ± SEM. Overall survival (OS) and event-free survival (EFS) were calculated from the date of diagnosis to the date of death or to the date of relapse, death or the time to no response to intensive induction, respectively. Follow-up was measured from the date of diagnosis to the date of last news for living patients. Survivals were calculated using the Kaplan–Meier method and were compared with the log-rank test. Uni- and multivariate survival analyses were done using Cox regression analysis (Wald test). Variables with a p value < 0.05 were tested in multivariate analysis. All statistical tests were two-sided at the 5% level of significance, except when indicated otherwise.
Abbreviations
- APL:
-
Acute promyelocytic leukemia
- ARAC:
-
Aracytine
- ATRA:
-
All-trans retinoic acid
- CEBPA:
-
CCAAT/enhancer-binding protein alpha
- ChIP:
-
Chromatin immunoprecipitation
- CN-AML:
-
Cytogenetically normal acute myeloid leukemia
- DMSO:
-
Dimethylsulfoxyde
- DNMT3A:
-
DNA-methyl transferase 3A
- DNR:
-
Daunorubicine
- EFS:
-
Event-free survival
- EZH2:
-
Enhancer of zeste homolog 2
- FACS:
-
Fluorescence Activated Cell Sorting
- FLT3:
-
Fms-like tyrosine kinase 3
- H3K27me3:
-
Histone H3 Lysine 27 tri-methylation
- HIST1:
-
Histone cluster 1
- HR:
-
Hazard ratio
- IDA:
-
Idarubicine
- IDH:
-
Isocitrate dehydrogenase
- ITD:
-
Internal tandem duplications
- KD:
-
Knock down
- KDM1/LSD1:
-
Lysine specific demethylase 1
- MEMα:
-
Minimum essential medium alpha
- NPM1:
-
Nucleophosmin 1
- OS:
-
Overall survival
- PBS:
-
Phosphate-buffered saline
- RFP:
-
Red fluorescent protein
- RPMI:
-
Roswell Park Memorial Institute medium
- SUZ12:
-
Suppressor of zeste 12
References
Grimwade D, Ivey A, Huntly BJ. Molecular landscape of acute myeloid leukemia in younger adults and its clinical relevance. Blood. 2016;127:29–41.
Falini B, Mecucci C, Tiacci E, et al. Cytoplasmic nucleophosmin in acute myelogenous leukemia with a normal karyotype. N Engl J Med. 2005;352:254–66.
Heath EM, Chan SM, Minden MD, et al. Biological and clinical consequences of NPM1 mutations in AML. Leukemia. 2017;31:798–807.
Papaemmanuil E, Gerstung M, Bullinger L, et al. Genomic classification and prognosis in acute myeloid leukemia. N Engl J Med. 2016;374:2209–21.
Wouters BJ, Delwel R. Epigenetics and approaches to targeted epigenetic therapy in acute myeloid leukemia. Blood. 2016;127:42–52.
Pfister SX, Ashworth A. Marked for death: targeting epigenetic changes in cancer. Nat Rev Drug Discov. 2017;16:241–63.
Marzluff WF, Gongidi P, Woods KR, et al. The human and mouse replication-dependent histone genes. Genomics. 2002;80:487–98.
Holmfeldt L, Wei L, Diaz-Flores E, et al. The genomic landscape of hypodiploid acute lymphoblastic leukemia. Nat Genet. 2013;45:242–52.
Fritz AJ, Ghule PN, Boyd JR, et al. Intranuclear and higher-order chromatin organization of the major histone gene cluster in breast cancer. J Cell Physiol. 2018;233:1278–90.
Tiberi G, Pekowska A, Oudin C, et al. PcG methylation of the HIST1 cluster defines an epigenetic marker of acute myeloid leukemia. Leukemia. 2015;29:1202–6.
Schlenk RF, Dohner K, Krauter J, et al. Mutations and treatment outcome in cytogenetically normal acute myeloid leukemia. N Engl J Med. 2008;358:1909–18.
Dohner H, Estey E, Grimwade D, et al. Diagnosis and management of AML in adults: 2017 ELN recommendations from an international expert panel. Blood. 2017;129:424–47.
Cancer Genome Atlas Research Network. Genomic and epigenomic landscapes of adult de novo acute myeloid leukemia. N Engl J Med. 2013;368:2059–74. Ley TJ, Miller C et al
Metzeler KH, Hummel M, Bloomfield CD, et al. An 86-probe-set gene-expression signature predicts survival in cytogenetically normal acute myeloid leukemia. Blood. 2008;112:4193–201.
Kohlmann A, Bullinger L, Thiede C, et al. Gene expression profiling in AML with normal karyotype can predict mutations for molecular markers and allows novel insights into perturbed biological pathways. Leukemia. 2010;24:1216–20.
Bei L, Lu Y, Eklund EA. HOXA9 activates transcription of the gene encoding gp91Phox during myeloid differentiation. J Biol Chem. 2005;280:12359–70.
Richard M, Veilleux P, Rouleau M, et al. The expression pattern of the ITIM-bearing lectin CLECSF6 in neutrophils suggests a key role in the control of inflammation. J Leukoc Biol. 2002;71:871–80.
Bagger FO, Sasivarevic D, Sohi SH, et al. BloodSpot: a database of gene expression profiles and transcriptional programs for healthy and malignant haematopoiesis. Nucleic Acids Res. 2016;44:D917–24.
Ng SW, Mitchell A, Kennedy JA, et al. A 17-gene stemness score for rapid determination of risk in acute leukaemia. Nature. 2016;540:433–7.
Hergeth SP, Schneider R. The H1 linker histones: multifunctional proteins beyond the nucleosomal core particle. EMBO Rep. 2015;16:1439–53.
Quentmeier H, Martelli MP, Dirks WG, et al. Cell line OCI/AML3 bears exon-12 NPM gene mutation-A and cytoplasmic expression of nucleophosmin. Leukemia. 2005;19:1760–7.
Chatterjee A, Rodger EJ, Eccles MR. Epigenetic drivers of tumourigenesis and cancer metastasis. Semin Cancer Biol. 2018;51:149–59.
Kernytsky A, Wang F, Hansen E, et al. IDH2 mutation-induced histone and DNA hypermethylation is progressively reversed by small-molecule inhibition. Blood. 2015;125:296–303.
Gollner S, Oellerich T, Agrawal-Singh S, et al. Loss of the histone methyltransferase EZH2 induces resistance to multiple drugs in acute myeloid leukemia. Nat Med. 2017;23:69–78.
Bertoli S, Picard M, Berard E, et al. Dexamethasone in hyperleukocytic acute myeloid leukemia. Haematologica. 2018;103:988–98.
Quek L, Otto GW, Garnett C, et al. Genetically distinct leukemic stem cells in human CD34- acute myeloid leukemia are arrested at a hemopoietic precursor-like stage. J Exp Med. 2016;213:1513–35.
Medrzycki M, Zhang Y, McDonald JF, et al. Profiling of linker histone variants in ovarian cancer. Front Biosci (Landmark Ed). 2012;17:396–406.
Bauden M, Kristl T, Sasor A, et al. Histone profiling reveals the H1.3 histone variant as a prognostic biomarker for pancreatic ductal adenocarcinoma. BMC Cancer. 2017;17:810.
Medrzycki M, Zhang Y, Zhang W, et al. Histone h1.3 suppresses h19 noncoding RNA expression and cell growth of ovarian cancer cells. Cancer Res. 2014;74:6463–73.
Sancho M, Diani E, Beato M, et al. Depletion of human histone H1 variants uncovers specific roles in gene expression and cell growth. PLoS Genet. 2008;4:e1000227.
Izzo A, Schneider R. The role of linker histone H1 modifications in the regulation of gene expression and chromatin dynamics. Biochim Biophys Acta. 2016;1859:486–95.
Millan-Arino L, Islam AB, Izquierdo-Bouldstridge A, et al. Mapping of six somatic linker histone H1 variants in human breast cancer cells uncovers specific features of H1.2. Nucleic Acids Res. 2014;42:4474–93.
Yang SM, Kim BJ, Norwood Toro L, et al. H1 linker histone promotes epigenetic silencing by regulating both DNA methylation and histone H3 methylation. Proc Natl Acad Sci U S A. 2013;110:1708–13.
Li H, Kaminski MS, Li Y, et al. Mutations in linker histone genes HIST1H1 B, C, D, and E; OCT2 (POU2F2); IRF8; and ARID1A underlying the pathogenesis of follicular lymphoma. Blood. 2014;123:1487–98.
Warrell RP Jr, Frankel SR, Miller WH Jr, et al. Differentiation therapy of acute promyelocytic leukemia with tretinoin (all-trans-retinoic acid). N Engl J Med. 1991;324:1385–93.
Boutzen H, Saland E, Larrue C, et al. Isocitrate dehydrogenase 1 mutations prime the all-trans retinoic acid myeloid differentiation pathway in acute myeloid leukemia. J Exp Med. 2016;213:483–97.
El Hajj H, Dassouki Z, Berthier C, et al. Retinoic acid and arsenic trioxide trigger degradation of mutated NPM1, resulting in apoptosis of AML cells. Blood. 2015;125:3447–54.
Martelli MP, Gionfriddo I, Mezzasoma F, et al. Arsenic trioxide and all-trans retinoic acid target NPM1 mutant oncoprotein levels and induce apoptosis in NPM1-mutated AML cells. Blood. 2015;125:3455–65.
Tassara M, Dohner K, Brossart P, et al. Valproic acid in combination with all-trans retinoic acid and intensive therapy for acute myeloid leukemia in older patients. Blood. 2014;123:4027–36.
Schenk T, Chen WC, Gollner S, et al. Inhibition of the LSD1 (KDM1A) demethylase reactivates the all-trans-retinoic acid differentiation pathway in acute myeloid leukemia. Nat Med. 2012;18:605–11.
Baik H, Boulanger M, Hosseini M, et al. Targeting the SUMO pathway primes all-trans retinoic acid-induced differentiation of nonpromyelocytic acute myeloid leukemias. Cancer Res. 2018;78:2601–13.
Delaunay J, Recher C, Pigneux A, et al. Addition of Gemtuzumab Ozogamycin to chemotherapy improves event-free survival but not overall survival of AML patients with intermediate cytogenetics not eligible for allogeneic transplantation. Results of the GOELAMS AML 2006 IR study. Blood. 2011;118:79.
Pigneux A, Bene MC, Salmi LR, et al. Improved survival by adding Lomustine to conventional chemotherapy for elderly patients with AML without unfavorable cytogenetics: results of the LAM-SA 2007 FILO trial. J Clin Oncol. 2018;36:3203–10.
Handschuh L, Kazmierczak M, Milewski MC, et al. Gene expression profiling of acute myeloid leukemia samples from adult patients with AML-M1 and -M2 through boutique microarrays, real-time PCR and droplet digital PCR. Int J Oncol. 2018;52:656–78.
Koubi M, Poplineau M, Vernerey J, et al. Regulation of the positive transcriptional effect of PLZF through a non-canonical EZH2 activity. Nucleic Acids Res. 2018;46:3339–50.
Acknowledgments
The authors thank the FACS core facility at CRCM for support with flow cytometry, the oncogenomics platform at IPC for sharing patient mutation data, the Biopathology Department at IPC for support with MGG coloration, and the TrGET platform for cell culture advices. The authors are grateful to Jihane Pakradouni Demestre for helping with collecting patient data.
Availability of the data and materials
The datasets generated and analyzed during the current study are available on reasonable request. In addition, the Microarray data are accessible under the accession number E-MTAB-6997.
Funding
This study was supported by the Institut National de la Santé et de la Recherche Médicale, the Centre National de la Recherche Scientifique, Inserm-Transfert to E.D. and J.V., Institut Thématique Multi-Organisme-cancer (P036560) to E.D. and L.NG., l’Institut National du Cancer (20141PLBIO06–1 to E.D. and M.Popl.), SIRIC grant INCa-DGOS-Inserm 6038 to N.V., E.D. and C.Che) and Groupement des Entreprises Françaises dans la Lutte contre le Cancer (GEFLUC) to E.D. S.G. was supported by the Fondation pour la Recherche Médicale (FFDM20160640830).
Author information
Authors and Affiliations
Contributions
SG and LN designed and performed the research and analyzed the data; PF, BC, DB, and FB provided and analyzed microarray data; CC and JV performed the research and analyzed the data; MP and NP contributed to experimental data. SA and MP performed and analyzed proteomic data. CR characterized and provided patient samples. DB contributed to the design and analysis of the study. CC and NV provided clinical data, and reviewed the manuscript. ED supervised the study, contributed to the design, and analysis of the study. SG and ED wrote the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
Informed consent was provided by all patients according to the Declaration of Helsinki and subjected to ethical institutional review board approval.
Consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Additional file 1.
Table S1. Clinical and molecular characteristics in the GOELAMS Cohort (n = 46) according to H3K27me3 HIST1 status. Table S2. Multivariate analyses in the validation CN-AML cohort (n = 46). Table S3. Univariate and Multivariate Analyses for 3-HIST1-mRNA signature in TCGA and Metzeler cohorts. Table S4. Clinical characteristics of NPM1mut patients selected for transcriptomic analysis.
Additional file 2.
Figure S1. Related to Fig. 2: Heatmap of the H3K27me3 level in a new cohort of 78 NPM1mut CN-AML patients. Figure S2. Related to Fig. 2: Analysis of H3K27me3 HIST1 status in CD34low and CD34high sorted blasts. Figure S3 Related to Fig. 4. Representative Integrative Genomics Viewer (IGV) tracks of H3K27me3 signal obtained from ChIP-chip data published in Tiberi et al., 2015 Figure S4. Related to Fig. 5: Histone protein extraction in NPM1mut patients. Figure S5. Related to Fig. 5:Total protein abundance of each histone types determined by IBAQ label-free quantification method. Figure S6 related to Fig. 6. Effect of H1-3 KD on histone H1 subtype mRNA and protein expression. Figure S7 related to Fig. 6. Effect of H1d KD on CD11b expression in shRNA1 (clones KD#2 and KD#3) and in shRNA2 conditions. Figure S7 related to Figure 6. Effect of H1d KD on CD11b expression in shRNA1 (clones KD#2 and KD#3) and in shRNA2 conditions
Additional file 3.
Supplemental Methods
Additional file 4.
Table S5. GES 81 g, LAM HIST1 16 ‘high (1)’ vs. 11 ‘low (0)’
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated.
About this article
Cite this article
Garciaz, S., N’guyen Dasi, L., Finetti, P. et al. Epigenetic down-regulation of the HIST1 locus predicts better prognosis in acute myeloid leukemia with NPM1 mutation. Clin Epigenet 11, 141 (2019). https://doi.org/10.1186/s13148-019-0738-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s13148-019-0738-6