Abstract
Background
Cryptosporidium is one of the most prevalent parasites infecting both birds and mammals. To examine the prevalence of Cryptosporidium species and evaluate the public health significance of domestic chickens in Guangdong Province, southern China, we analyzed 1001 fecal samples from 43 intensive broiler chicken farms across six distinct geographical regions.
Methods
Individual DNA samples were subjected to nested PCR-based amplification and sequencing of the small subunit of the nuclear ribosomal RNA gene (SSU rRNA). Analysis of the 60 kDa glycoprotein gene (gp60) was performed to characterize the subtypes of C. meleagridis.
Results
The overall prevalence of Cryptosporidium was 13.2% (95% CI 11.1–15.3) (24 of 43 farms), with C. meleagridis (7.8%), C. baileyi (4.8%) and mixed infections (0.6%). Using the gp60 gene, three subtype families, IIIb, IIIe and IIIg, were identified, including six subtypes: one novel (IIIgA25G3R1a) and five previously reported (IIIbA23G1R1c, IIIbA24G1R1, IIIbA21G1R1a, IIIeA17G2R1 and IIIeA26G2R1). Within these subtypes, five known subtypes were genetically identical to those identified in humans.
Conclusions
This is the first report of C. meleagridis in chickens from Guangdong. The frequent occurrence of C. meleagridis in domestic chickens and the common C. meleagridis subtypes identified in both humans and chickens is of public health significance. Our study indicates that broiler chickens represent a potential zoonotic risk for the transmission of Cryptosporidium in this region.
Graphical Abstract

Similar content being viewed by others
Background
Cryptosporidium is a protozoan parasite that infects a wide range of vertebrate hosts, including humans and birds [1]. In birds, Cryptosporidium was first found in the order Galliformes, and since then has been reported in more than 30 avian species worldwide [2].
Currently, only five valid species, namely C. meleagridis, C. baileyi, C. galli, C. avium and C. proventriculi, and at least 15 genotypes including avian genotypes I–II, IV and VI–IX, goose genotypes I–V, black duck genotype, Eurasian woodcock genotype and C. xiaoi–like genotype have been documented in a wide range of birds worldwide [3,4,5,6,7,8,9,10,11]. In addition, mammal-specific Cryptosporidium species including C. hominis, C. parvum, C. andersoni, C. muris and C. canis are rarely detected in birds [12,13,14,15,16], partly because birds ingest oocysts from contaminated food or water and shed oocysts mechanically. Cryptosporidium baileyi infection usually occurs in the respiratory system, causing high morbidity and mortality, and C. meleagridis infects the gut and is associated with intestinal clinical signs (enteritis and diarrhea), whereas C. galli and C. proventriculi infect the proventriculus, manifesting symptoms associated with anorexia, weight loss and chronic vomiting [6]. Cryptosporidium avium primarily infects the bursa fabricii, but has been described with no pathogenicity [7, 17].
The identification and characterization of species, genotypes and subtypes of Cryptosporidium by molecular methods is crucial for the tracing of contamination sources and the assessment of public health importance. Small subunit (SSU) rRNA and the 60-kDa glycoprotein (gp60) genes are commonly used for determining Cryptosporidium species/genotypes and subtyping, respectively [18,19,20]. Currently, C. meleagridis is considered the third most common species infecting humans and the only species that infects both birds and mammals [19]. In China, C. meleagridis has been recorded previously in diarrheic children and chickens in Wuhan, Hubei Province; importantly, some gp60 C. meleagridis subtypes characterized from diarrheic children were shared by chickens in the same location [8, 21]. Recently, avian-specific Cryptosporidium species C. baileyi pulmonary infection was found in an immunocompetent woman with a benign tumor in Poland [22]. This information highlights the epidemiological importance of avian hosts as significant reservoirs for human cryptosporidiosis, although the extent of cross-species transmission of these zoonotic species remains unclear [4].
Guangdong Province, southern China, is particularly rich in domestic poultry producers. The interaction between humans and domestic poultry poses the potential for zoonotic transmission. However, there are no reports on Cryptosporidium isolated from commercial broiler chickens in this region to date, only a study in domestic pigeons [23]. The aim of the present study was to estimate the occurrence and genetic diversity of Cryptosporidium species and C. meleagridis subtypes, and the public health significance of chickens in intensive farms in Guangdong Province.
Methods
Sample collection
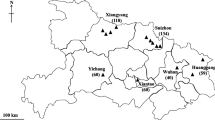
Fresh pooled fecal samples from the floor were randomly collected from broiler chickens (Qing Yuan-ma chickens) from 43 medium-sized to large intensive farms (with 1000–25,000 chickens per farm on average) across six distinct geographical regions (Qingyuan, Maoming, Huizhou, Meizhou, Yangjiang and Shanwei) in Guangdong Province (23°13′S, 113°26′W), China, from June 2020 to March 2021 (Fig. 1; Table 1). Each sample contained 4–5 single fecal deposit droppings from different areas inside the poultry house that were pooled into a single sample. Samples of different consistency and color were chosen on the ground within 1–2 m2 to avoid sampling error. Five to 10 samples were collected per farm from broiler flocks comprising 50–100 chickens. A total of 1001 fecal samples of broiler chickens, comprising animals aged < 30 days (n = 169), 31–60 days (n = 440), 61–90 days (n = 348) and > 90 days (n = 44), were collected. All samples were collected from apparently healthy flocks. Care was taken to avoid sampling fecal material that had been in contact with the ground. The pooled fecal samples (approximately 50 g) were collected into clean plastic bags, kept in ice boxes and marked with the region, number and date. Samples were then transported immediately to the laboratory and stored at 4 °C. Samples were examined within 24 h after collection.
DNA extraction and polymerase chain reaction (PCR) amplification
For genomic DNA extraction, approximately 200 mg of fecal samples was suspended in 100 ml of distilled water and centrifuged at 3000×g for 10 min. The process was repeated at least three times until the supernatant was clear. Genomic DNA samples were extracted from individual treated materials using the E.Z.N.A.® Stool DNA Kit (Omega Bio-tek Inc., Norcross, GA, USA) in accordance with the manufacturer's instructions and then frozen at −20 °C prior to PCR analysis.
Individual DNA samples were subjected to nested PCR-based amplification and sequencing of the small subunit of the nuclear ribosomal RNA gene (SSU rRNA, ~ 830 base pairs [bp]) [24]. Subtypes of C. meleagridis and mixed-species infections were determined by amplification of the 60 kDa glycoprotein gene (gp60; ~ 900 bp) from positive C. meleagridis samples and positive C. baileyi samples, respectively [20]. PCR was conducted in a 50-μl reaction mixture containing 1× PCR buffer (Takara Shuzo Co., Ltd., Otsu, Japan), 3.0 mM of MgCl2, 0.2 mM of each deoxynucleotide triphosphate, 50 pM of each primer, 1 unit of rTaq DNA polymerase (Takara Shuzo Co., Ltd.), 2 μl of DNA sample and 1 μl of bovine serum albumin. Known test-positive (cattle DNA) and test-negative (distilled water) controls were included with each PCR reaction. The amplification products were separated by electrophoresis in 1.5% agarose gels, stained with ethidium bromide and visualized on an ultraviolet (UV) transilluminator.
Nucleotide sequencing and analysis
All secondary PCR amplicons were sequenced using an ABI PRISM™ 3730xl DNA Analyzer (Applied Biosystems, Foster City, CA, USA) in both directions. Sequences were aligned by the Clustal version 2.1 program and adjusted manually by BioEdit 7.04 software. The adjusted sequences were submitted to a BLAST [Basic Local Alignment Search Tool] search to initially define the species and to further confirm the high similarity with other known sequences of Cryptosporidium spp. in the GenBank database.
Phylogenetic analysis
Phylogenetic analysis was performed using Bayesian inference (BI) and Markov chain Monte Carlo (MCMC) methods in MrBayes version 3.2.6. The best-fit nucleotide substitution model was generalized time-reversible model (GTR + G) determined by ModelTest version 3.7. Based on 1,000,000 generations with four simultaneous tree-building chains, posterior probability values were estimated with trees being saved every 1000th generation. A 50% majority rule consensus tree for each analysis was constructed based on the final 75% of trees generated. To ensure convergence and insensitivity to priors, analyses were run three times by BI. Posterior probabilities of > 0.95 are indicated at all major nodes.
Statistical analysis
Statistical analysis was performed by chi-square tests, and differences were considered significant when P < 0.01 was obtained using SAS version 9.1 (SAS Institute Inc., Cary, NC, USA). Odds ratios (ORs) and 95% confidence intervals (95% CIs) were calculated by SPSS version 22.0 software (IBM Corp., Armonk, NY, USA).
Results
Prevalence of Cryptosporidium
Of 1001 broiler chicken DNA samples, 132 samples tested positive by PCR amplification of the SSU rRNA gene, equating to an overall prevalence of Cryptosporidium of 13.2% (95% CI 11.1–15.3%). The PCR-positive chickens were detected on 24 of 43 farms from six geographical regions examined, with prevalence ranging from 4.0% to 62.5% (Additional file 1: Table S1). There was no significant statistical difference in geographical provenance or prevalence among regions (χ2 = 14.209, df = 5, P = 0.014) (Table 1).
Species and age distribution
Cryptosporidium species and genotypes were identified through sequencing of SSU amplicons (n = 132). This analysis revealed C. baileyi in 48 (4.8%) and C. meleagridis in 78 (7.8%) of the 132 SSU rRNA -positive samples. Six mixed-species infections were also detected. Seven distinct SSU rRNA sequences were deposited under GenBank accession numbers OK560460–OK560466. Cryptosporidium was detected in all age groups (Table 2), and chickens aged 61–90 days (17.8%) showed a significantly higher infection rate than chickens < 30 days (8.3%), 31–60 days (12.0%) and > 90 days (6.8%) of age (χ2 = 12.123, df = 3, P = 0.007). Cryptosporidium baileyi was detected in chickens of all age groups (Table 2), and 61–90-day-old chickens tended to have a higher infection rate (8.6%) than other age groups (χ2 = 20.600, df = 3, P = 0.000). Cryptosporidium meleagridis was only detected in chickens ≤ 90 days of age, and no statistically significant differences were observed among three age groups (χ2 = 0.092, df = 2, P = 0.955).
Subtypes of C. meleagridis
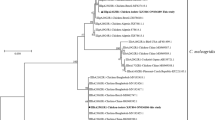
Among 78 C. meleagridis-positive specimens, 64 yielded gp60 PCR products of the expected size. Alignment of gp60 nucleotide sequences obtained here and references downloaded from the GenBank database revealed the presence of six subtypes, including five known (IIIbA23G1R1c, IIIbA24G1R1, IIIbA21G1R1a, IIIeA17G2R1 and IIIeA26G2R1) and one previously unreported IIIg25G3R1 variant (IIIbA25G3R1a) (GenBank: OK 562,693–562,699). As expected, the phylogenetic tree revealed three distinct clusters, representing three subtype families (IIIb, IIIe and IIIg) (Fig. 2). The most common subtype family, IIIb, was identified in 51 samples, including known subtypes IIIbA23G1R1c (n = 24), IIIbA24G1R1 (n = 20) and IIIbA21G1R1a (n = 7). The second most common subtype family, IIIe, was identified in 11 chicken isolates and comprised three distinct subtypes, two known IIIeA17G2R1 subtypes (n = 8) and a known IIIeA26G2R1 subtype (n = 3). Finally, for subtype family IIIg, IIIgA25G3R1a was identified in two chicken isolates.
Phylogenetic relationship of the nuclear 60-kDa glycoprotein gene (pgp60) of Cryptosporidium meleagridis in chickens by Bayesian inference (BI). Posterior probabilities of > 0.95 are indicated at all major nodes. Subtypes tested in this study are labeled after the specimen numbers. The scale bar represents the number of substitutions per site
Discussion
To our knowledge, this is the first report on the presence and prevalence of Cryptosporidium in intensively farmed chickens in Guangdong Province, although previous studies have been reported in Hubei, Zhejiang, Henan and Anhui in China [3, 5, 8, 25]. In our study, the overall prevalence of Cryptosporidium in chickens (13.2%; 132/1001) was comparable to previous numbers reported for domestic chickens in Brazil (12.6%) [10], China (10.2%) [8] and Syria (9.9%) [26], and higher than Iran (0.5%) [27], Tunisia (4.5%) [28], Jordan (4.8%) [29] and Germany (5.7%) [15], but lower than Brazil (25.6%) [30] and Algeria (34.4%) [4]. Differences in hygiene, management practices, sample origin and detection methods may contribute to these differences in the prevalence of Cryptosporidium in poultry flocks.
In addition, Cryptosporidium infection in broiler chickens appeared to be age-related. However, unlike the age-related infection pattern whereby the infection rate decreases with increasing age of infected animals in ruminants [31], the highest infection rate of 17.8% was detected in 61–90-day-old broiler chickens, compared with the other age groups (P < 0.01) (Table 2). In a previous study in China, chickens ≤ 4 months had the highest infection rate [8], which was partially in agreement with our results. It is worth noting that most broiler chickens are sold by the age of 61–90 days; therefore, oocysts may be disseminated during the process of transfer and new infection may result. These broiler chickens should be considered as important reservoirs of Cryptosporidium, although the age-related association should be verified by further research.
Cryptosporidium meleagridis and C. baileyi were confirmed by molecular characterization of the SSU rRNA gene, which was consistent with previous studies reported in farmed and wild birds in China, including chickens, domestic pigeons, quails, ducks, ostriches and white Java sparrows [3, 5, 10, 15, 23, 28, 32, 33]. Cryptosporidium baileyi, originally isolated from commercial broiler chickens [34], has a broad range of avian hosts and is considered the predominant avian Cryptosporidium species. In China, C. baileyi has been reported in a wide variety of birds, including chickens, quails, ostriches, Pekin ducks, domestic pigeons and geese, as well as some pet birds [3, 5, 8, 23, 33, 35]. Recently, C. baileyi was also found in an immunocompetent woman with a benign tumor in Poland [22]. Evidence has shown that C. baileyi causes respiratory disease and production loss in chickens, causing reduced weight gain in broilers and decreased egg production in laying chickens [36]. One study showed that C. baileyi is one cause of Newcastle disease and/or avian influenza vaccination failure in poultry farms [37].
Cryptosporidium meleagridis has been detected in various avian hosts, including chickens, turkeys, cockatiels, pigeons and quails, as well as some pet birds [3, 8, 23, 33, 38,39,40]. Cryptosporidium meleagridis has also been frequently detected in humans worldwide, especially in immunocompromised individuals such as neonates and patients with HIV/AIDS [2]. In China, C. meleagridis has been detected in diarrheic children in Wuhan [21], HIV-positive patients in Henan [41] and pediatric patients in Shanghai [42]. Cryptosporidium meleagridis is an emerging human pathogen and constitutes the third most common human-pathogenic Cryptosporidium species after C. hominis and C. parvum [43,44,45]. Moreover, molecular studies have revealed that identical C. meleagridis subtypes were shared between humans and birds in the same location in Sweden, Peru and China [46], suggesting cross-species transmission of C. meleagridis between birds and humans. Chickens may act as a source of infection and a mechanical vector by shedding C. meleagridis oocysts into the environment.
Surprisingly, among 132 test-positive chicken fecal samples, C. baileyi was detected in the minority (40%) of samples, while C. meleagridis was detected in the majority (60%) of samples. The prevalence of C. meleagridis in the present study is significantly higher than that of the typical avian species C. baileyi [2, 3, 5, 8, 10]. The predominance of C. meleagridis among the chicken fecal samples in the present study is consistent with the results of a previous study in poultry in Brazil [47]. Because of the lack of related epidemiological data on cryptosporidiosis in mammals/humans in the investigated areas, the source of infection of domesticated chickens with C. meleagridis remains to be elucidated. Whether chickens acquire the infection by contamination of water, feed and/or litter in poultry houses with oocysts from human origin requires further investigation.
To date, at least nine subtype families (IIIa to IIIi) of C. meleagridis have been identified by nucleotide sequence analysis of the gp60 gene [4, 48, 49]. In the present study, three subtype families of C. meleagridis, IIIb, IIIe and IIIg, were detected. Subtypes IIIbA21G1R1a, IIIbA24G1R1, IIIbA23G1R1c, IIIeA17G2R1 and IIIeA26G2R1 have previously been reported sporadically in birds [8, 47], but predominantly in humans, especially those with a travel history to Asia [20, 21, 41, 50]. For example, subtype IIIbA23G1R1c, the predominant subtype found here, had previously been isolated from a Swedish patient with a history of travel to Malaysia, while other variants of this subtype, IIIbA23G1R1a and IIIbA23G1R1b, were reported in patients who had traveled to other developing countries (Indonesia or Thailand) prior to infection [20, 50]. Similarly, subtype IIIbA24G1R1, IIIbA21G1R1a and IIIeA17G2R1 infections had previously been linked to travel to Asian countries (China, Thailand or Vietnam). These reports indicate that foreign travel is a significant risk factor for infection with C. meleagridis. Meanwhile, subtype IIIeA26G2R1 identified in chickens in this study was also previously identified in HIV-positive patients in China [41]. This accumulated information suggests the cross-transmission of cryptosporidiosis between chickens and humans in this region. Therefore, to better prevent human cryptosporidiosis, specific management measures are needed on poultry farms, including adherence to an appropriate feeding model as well as strict hygiene and waste management procedures.
Conclusions
This is the first large-scale molecular study on the occurrence and genetic identity of Cryptosporidium in farm-raised chickens in Guangdong Province, China. Two species, C. meleagridis and C. baileyi, were identified. Five of the six subtypes of C. meleagridis detected in this study matched those identified in humans. The dominance of C. meleagridis infection among chickens and the detection of zoonotic subtypes IIIbA21G1R1a and IIIbA24G1R1 are indicative of cross-transmission of cryptosporidiosis between chickens and humans. Domestic chickens are of public health significance as potential reservoirs of zoonotic Cryptosporidium. Further epidemiological investigations are needed to confirm the source of infection of domesticated chickens with C. meleagridis.
Availability of data and materials
The data supporting the conclusions of this article are included within the article and its additional file. Seven distinct SSU rRNA sequences and subtypes sequences of C. meleagridis were deposited under GenBank accession numbers OK560460–OK560466 and OK562693–OK562699, respectively.
Abbreviations
- SSU rRNA:
-
Small subunit of nuclear ribosomal RNA
- gp60:
-
60 KDa glycoprotein
- C. meleagridis :
-
Cryptosporidium meleagridis
- C. baileyi :
-
Cryptosporidium baileyi
References
Fayer R. Taxonomy and species delimitation in Cryptosporidium. Exp Parasitol. 2010;124:90–7.
Ryan UM. Cryptosporidium in birds, fish and amphibians. Exp Parasitol. 2010;124:113–20.
Wang R, Jian F, Sun Y, Hu Q, Zhu J, Wang F, et al. Large-scale survey of Cryptosporidium spp. in chickens and Pekin ducks (Anas platyrhynchos) in Henan, China: prevalence and molecular characterization. Avian Pathol. 2010;39:447–51.
Baroudi D, Khelef D, Goucem R, Adjou KT, Adamu H, Zhang H, et al. Common occurrence of zoonotic pathogen Cryptosporidium meleagridis in broiler chickens and turkeys in Algeria. Vet Parasitol. 2013;196:334–40.
Wang L, Xue X, Li J, Zhou Q, Yu Y, Du A. Cryptosporidiosis in broiler chickens in Zhejiang Province, China: molecular characterization of oocysts detected in fecal samples. Parasite. 2014;21:36.
Nakamura AA, Meireles MV. Cryptosporidium infections in birds-a review. Rev Bras Parasitol Vet. 2015;24:253–67.
Holubová N, Sak B, Horčičková M, Hlásková L, Květoňová D, Menchaca S, et al. Cryptosporidium avium n. sp. (Apicomplexa: Cryptosporidiidae) in birds. Parasitol Res. 2016;115:2243–51.
Liao C, Wang T, Koehler AV, Fan Y, Hu M, Gasser RB. Molecular investigation of Cryptosporidium in farmed chickens in Hubei Province, China, identifies “zoonotic” subtypes of C. meleagridis. Parasit Vectors. 2018;11:484.
Holubová N, Zikmundová V, Limpouchová Z, Sak B, Konečný R, Hlásková L, et al. Cryptosporidium proventriculi sp. n. (Apicomplexa: Cryptosporidiidae) in Psittaciformes birds. Eur J Protistol. 2019;69:70–87.
Santana BN, Kurahara B, Nakamura AA, da Silva Camargo V, Ferrari ED, da Silva GS, et al. Detection and characterization of Cryptosporidium species and genotypes in three chicken production systems in Brazil using different molecular diagnosis protocols. Prev Vet Med. 2018;151:73–8.
Liu X, Zhu H, Meng W, Dong H, Han Q, An Z, et al. Occurrence of a Cryptosporidium xiaoi-like genotype in peafowl (Pavo cristatus) in China. Parasitol Res. 2019;118:3555–9.
Ryan UM, Xiao L, Read C, Sulaiman IM, Monis P, Lal AA, et al. A redescription of Cryptosporidium galli Pavlasek, 1999 (Apicomplexa: Cryptosporidiidae) from birds. J Parasitol. 2003;89:809–13.
Zhou L, Kassa H, Tischler ML, Xiao L. Host-adapted Cryptosporidium spp. in Canada geese (Branta canadensis). Appl Environ Microbiol. 2004;70:4211–5.
Ng J, Pavlasek I, Ryan U. Identification of novel Cryptosporidium genotypes from avian hosts. Appl Environ Microbiol. 2006;72:7548–53.
Helmy YA, Krücken J, Abdelwhab EM, Samson-Himmelstjerna G, Hafez HM. Molecular diagnosis and characterization of Cryptosporidium spp. in turkeys and chickens in Germany reveals evidence for previously undetected parasite species. PLoS ONE. 2017;12:e0177150.
Ferrari ED, Nakamura AA, Nardi ARM, Santana BN, da Silva CV, Nagata WB, et al. Cryptosporidium spp. in caged exotic psittacines from Brazil: evaluation of diagnostic methods and molecular characterization. Exp Parasitol. 2018;184:109–14.
Cui Z, Song D, Qi M, Zhang S, Wang R, Jian F, et al. Revisiting the infectivity and pathogenicity of Cryptosporidium avium provides new information on parasitic sites within the host. Parasit Vectors. 2018;11:514.
Ryan U, Fayer R, Xiao L. Cryptosporidium species in humans and animals: current understanding and research needs. Parasitology. 2014;141:1667–85.
Xiao L. Molecular epidemiology of cryptosporidiosis: an update. Exp Parasitol. 2010;124:80–9.
Stensvold CR, Beser J, Axén C, Lebbad M. High applicability of a novel method for gp60-based subtyping of Cryptosporidium meleagridis. J Clin Microbiol. 2014;52:2311–9.
Wang T, Fan Yi, Koehler AV, Ma G, Li T, Hu M, et al. First survey of Cryptosporidium, Giardia and Enterocytozoon in diarrhoeic children from Wuhan, China. Infect Genet Evol. 2017;51:127–31.
Kopacz Ż, Kváč M, Piesiak P, Szydłowicz M, Hendrich AB, Sak B, et al. Cryptosporidium baileyi pulmonary infection in immunocompetent woman with Benign Neoplasm. Emerg Infect Dis. 2020;26:1958–61.
Li J, Lin X, Zhang L, Qi N, Liao S, Lv M, et al. Molecular characterization of Cryptosporidium spp. in domestic pigeons (Columba livia domestica) in Guangdong Province, Southern China. Parasitol Res. 2015;114:2237–41.
Xiao L, Singh A, Limor J, Graczyk TK, Gradus S, Lal A. Molecular characterization of Cryptosporidium oocysts in samples of raw surface water and wastewater. Appl Environ Microbiol. 2004;67:1097–101.
Gong Z, Kan Z, Huang J, Fang Z, Liu X, Gu Y, et al. Molecular prevalence and characterization of Cryptosporidium in domestic free-range poultry in Anhui Province. China Parasitol Res. 2021;120:3519–27.
Kassouha M, Soukkarieh C, Alkhaled A. First genotyping of Cryptosporidium spp. in pre-weaned calves, broiler chickens and children in Syria by PCR-RFLP analysis. Vet Parasitol. 2016;225:86–90.
Hamidinejat H, Jalali MH, Jafari RA, Nourmohammadi K. Molecular determination and genotyping of Cryptosporidium spp. in fecal and respiratory samples of industrial poultry in Iran. Asian Pac J Trop Med. 2014;7:517–20.
Soltane R, Guyot K, Dei-Cas E, Ayadi A. Prevalence of Cryptosporidium spp. (Eucoccidiorida: Cryptosporiidae) in seven species of farm animals in Tunisia. Parasite. 2007;14:335–8.
Hijjawi N, Mukbel R, Yang R, Ryan U. Genetic characterization of Cryptosporidium in animal and human isolates from Jordan. Vet Parasitol. 2016;228:116–20.
Ewald MPC, Martins FDC, Caldart ET, Vieira FEG, Yamamura MH, Sasse JP, et al. The first study of molecular prevalence and species characterization of Cryptosporidium in free-range chicken (Gallus gallus domesticus) from Brazil. Rev Bras Parasitol Vet. 2017;26:472–8.
Santín M, Trout JM, Xiao L, Zhou L, Greiner E, Fayer R. Prevalence and age-related variation of Cryptosporidium species and genotypes in dairy calves. Vet Parasitol. 2004;122:103–17.
Amer S, Wang C, He H. First detection of Cryptosporidium baileyi in Ruddy Shelduck (Tadorna ferruginea) in China. J Vet Med Sci. 2010;72:935–8.
Wang R, Wang F, Zhao J, Qi M, Ning C, Zhang L, et al. Cryptosporidium spp. in quails (Coturnix coturnix japonica) in Henan, China: molecular characterization and public health significance. Vet Parasitol. 2012;187:534–7.
Current W, Upton SJ, Haynes TB. The life cycle of Cryptosporidium baileyi n. sp. (Apicomplexa, Cryptosporidiidae) infecting chickens. J Protozool. 1986;33:289–96.
Zhao W, Zhou H, Ma T, Cao J, Lu G, Shen Y. PCR-based detection of Cryptosporidium spp. and Enterocytozoon bieneusi in farm-raised and free-ranging geese (Anser anser f. domestica) from Hainan Province of China: natural infection rate and the species or genotype distribution. Front Cell Infect Microbiol. 2019;9:416.
Goodwin MA, Brown J, Resurreccion RS, Smith JA. Respiratory coccidiosis (Cryptosporidium baileyi) among northern Georgia broilers in one company. Avian Dis. 1996;40:572–5.
Eladl AH, Hamed HR, Khalil MR. Consequence of Cryptosporidiosis on the immune response of vaccinated broiler chickens against Newcastle disease and/or avian influenza. Vet Res Commun. 2014;38:237–47.
Laatamna AE, Holubova N, Sak B, Kvac M. Cryptosporidium meleagridis and C. baileyi (Apicomplexa) in domestic and wild birds in Algeria. Folia Parasitol. 2017;64:018.
Abe N, Iseki M. Identification of Cryptosporidium isolates from cockatiels by direct sequencing of the PCR-amplified small subunit ribosomal RNA gene. Parasitol Res. 2004;92:523–6.
Qi M, Wang R, Ning C, Li X, Zhang L, Jian F, et al. Cryptosporidium spp. in pet birds: genetic diversity and potential public health significance. Exp Parasitol. 2011;128:336–40.
Wang L, Zhang H, Zhao X, Zhang L, Zhang G, Guo M, et al. Zoonotic Cryptosporidium species and Enterocytozoon bieneusi genotypes in HIV-positive patients on antiretroviral therapy. J Clin Microbiol. 2013;51:557–63.
Feng Y, Wang L, Duan L, Gomez-Puerta LA, Zhang L, Zhao X, et al. Extended outbreak of cryptosporidiosis in a pediatric hospital, China. Emerg Infect Dis. 2012;18:312–4.
McLauchlin J, Amar C, Pedraza-Díaz S, Nichols GL. Molecular epidemiological analysis of Cryptosporidium spp. in the United Kingdom: results of genotyping Cryptosporidium spp. in 1,705 fecal samples from humans and 105 fecal samples from livestock animals. J Clin Microbiol. 2000;38:3984–90.
Matos O, Alves M, Xiao L, Cama V, Antunes F. Cryptosporidium felis and C. meleagridis in persons with HIV. Portugal Emerg Infect Dis. 2004;10:2256–7.
Cama VA, Bern C, Roberts J, Cabrera L, Sterling CR, Ortega Y, et al. Cryptosporidium species and subtypes and clinical manifestations in children, Peru. Emerg Infect Dis. 2008;14:1567–74.
Silverlås C, Mattsson JG, Insulander M, Lebbad M. Zoonotic transmission of Cryptosporidium meleagridis on an organic Swedish farm. Int J Parasitol. 2012;42:963–7.
da Cunha MJR, Cury MC, Santín M. Molecular characterization of Cryptosporidium spp. in poultry from Brazil. Res Vet Sci. 2018;118:331–5.
Feng Y, Lal AA, Li N, Xiao L. Subtypes of Cryptosporidium spp. in mice and other small mammals. Exp Parasitol. 2011;127:238–42.
Abal-Fabeiro JL, Maside X, Llovo J, Bartolomé C. Aetiology and epidemiology of human cryptosporidiosis cases in Galicia (NW Spain), 2000–2008. Epidemiol Infect. 2015;143:3022–35.
Lebbad M, Winiecka-Krusnell J, Stensvold CR, Beser J. High diversity of Cryptosporidium species and subtypes identified in cryptosporidiosis acquired in Sweden and abroad. Pathogens. 2021;10:523.
Acknowledgements
We are grateful to the chicken farmers for collecting fecal samples. Thanks to Liwen Bianji, Edanz Group China (www.liwenbianji.cn/ac), for editing the English text of a draft of this manuscript.
Funding
Project support was provided in part by the State Key Laboratory of Veterinary Etiological Biology, Lanzhou Veterinary Research Institute, Chinese Academy of Agricultural Sciences (Grant No. SKLVEB2019KFKT016). Key Realm R&D Program of Guangdong Province (2020B0202080004, 2020B0202090004), NSFC grants (31872460), NSF grant of Guangdong Province (2021A1515010521, 2021A1515012401, 2019A1515010913), Science and Technology Project of Heyuan (2021008), Science and Technology Project of Guangzhou (202102080459), Special Fund for Scientific Innovation Strategy-Construction of High Level Academy of Agriculture Science (202110TD, 202122TD, R2020PY-JC001, R2019YJ-YB3010, R2020PY-JG013, R2020QD-048), and Guangdong Provincial Special Fund for Modern Agriculture Industry Technology Innovation teams (2021KJ119).
Author information
Authors and Affiliations
Contributions
XJ, MS and MQ planned the study. XL, LX, NQ and ML collected samples. SL, JL and MH undertook the laboratory and analytical work. XL and HC wrote the manuscript, with active inputs from JZ and JH. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Additional file 1: Table S1
. Samples from 43 intensive chicken farms across six distinct geographical regions (Qingyuan, Maoming, Huizhou, Meizhou, Yangjiang and Shanwei) in Guangdong Province.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated in a credit line to the data.
About this article
Cite this article
Lin, X., Xin, L., Qi, M. et al. Dominance of the zoonotic pathogen Cryptosporidium meleagridis in broiler chickens in Guangdong, China, reveals evidence of cross-transmission. Parasites Vectors 15, 188 (2022). https://doi.org/10.1186/s13071-022-05267-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s13071-022-05267-x