Abstract
Background
Chlamydomonas reinhardtii is a unicellular green alga, which is a most commonly used model organism for basic research and biotechnological applications. Generation of transgenic strains, which usually requires selectable markers, is instrumental in such studies/applications. Compared to other organisms, the number of selectable markers is limited in this organism. Nourseothricin (NTC) N-acetyl transferase (NAT) has been reported as a selectable marker in a variety of organisms but not including C. reinhardtii. Thus, we investigated whether NAT was useful and effective for selection of transgenic strains in C. reinhardtii. The successful use of NAT would provide alterative choice for selectable markers in this organism and likely in other microalgae.
Results
C. reinhardtii was sensitive to NTC at concentrations as low as 5 µg/ml. There was no cross-resistance to nourseothricin in strains that had been transformed with hygromycin B and/or paromomycin resistance genes. A codon-optimized NAT from Streptomyces noursei was synthesized and assembled into different expression vectors followed by transformation into Chlamydomonas. Around 500 transformants could be obtained by using 50 ng DNA on selection with 10 µg/ml NTC. The transformants exhibited normal growth rate and were stable at least for 10 months on conditions even without selection. We successfully tested that NAT could be used as a selectable marker for ectopic expression of IFT54-HA in strains with paromomycin and hygromycin B resistance markers. We further showed that the selection rate for IFT54-HA positive clones was greatly increased by fusing IFT54-HA to NAT and processing with the FMDV 2A peptide.
Conclusions
This work represents the first demonstration of stable expression of NAT in the nuclear genome of C. reinhardtii and provides evidence that NAT can be used as an effective selectable marker for transgenic strains. It provides alterative choice for selectable markers in C. reinhardtii. NAT is compatible with paromomycin and hygromycin B resistance genes, which allows for multiple selections.
Similar content being viewed by others
Background
Chlamydomonas reinhardtii (C. reinhardtii), a unicellular green alga, is a widely used model organism for basic scientific research as well as biotechnological applications [1]. Generation of transgenic strains plays a critical role in our deeper understanding of molecular mechanisms involved in various cellular processes and genetic engineering for producing valuable products [2, 3].
Because of low efficiency of transformation, a selectable marker is usually needed for selection of transgenic strains. Currently, there are three types of selections used in nuclear transformation of C. reinhardtii: auxotrophy rescue, herbicide resistance and antibiotic resistance [1,2,3]. Auxotrophy rescue involves using parental strains with mutations thus limiting its application. For example, Nit1, a gene encoding nitrate reductase, can only be used in transformation of strains with defects in Nit1 [4]. Several herbicide resistance markers have been reported [5,6,7]. The herbicides used include dichlorophenyl dimethyl urea (DCMU), norflurazon, oxyfluorfen, glyphosate and sulfadiazine. For reasons unknown, the herbicide resistance markers are rarely adopted in the community. It is likely due to high dose application of herbicide, poor transformation efficiency and/or other reasons. Six antibiotics have been used in C. reinhardtii for selection of transgenic strains transformed with corresponding selectable markers [1, 3]. The antibiotics used include paromomycin, zeocin, spectinomycin, hygromycin B, kanamycin and tetracycline. According to our understanding, only paromomycin, hygromycin B and zeocin resistance genes are commonly used as selectable markers [8,9,10]. Zeocin for selecting of Ble gene transformants is light sensitive and may induce genomic damages even in cells harboring the selection marker [11]. Compared to higher number of selectable markers in higher plant and mammalian cells [12, 13], the number of effective selectable markers is limited in C. reinhardtii. Therefore, availability of additional selectable markers in C. reinhardtii will enable complex experimental design, for example triple or more selection for transgenic strains.
Nourseothricin (NTC), a metabolite produced by Streptomyces noursei, belongs to streptothricin-class aminoglycoside antibiotics that inhibit protein synthesis [14]. NTC N-acetyl transferase (NAT) derived from S. noursei inactivates NTC by acetylating the beta-amino group of the beta-lysine residue [15]. NTC is highly soluble in water (1 g/ml) and stable for 2 years even in solution. NAT has been used as a selectable marker in a variety of organisms including bacteria, fungi, plant and mammalian cells (https://www.jenabioscience.com/images/741d0cd7d0/NTC-Flyer.pdf). However, NAT has been used in diatoms but not in other microalgae including C. reinhardtii [16].
In this report, we have shown that NAT is an effective selectable marker for nuclear transformation of C. reinhardtii. NTC, as low as 5 µg/ml, effectively kills or suppresses the growth of C. reinhardtii wild type cells as well as strains harboring paromomycin and/or hygromycin B resistant genes. Codon-optimized NAT from S. noursei is expressible in C. reinhardtii and confers cell resistance to NTC. We further show that NAT can be used as a selectable marker for transgenic strains even in strains harboring paromomycin and/or hygromycin B resistant genes. Furthermore, by fusing of a target gene to NAT and processing with the FMDV 2A peptides, the selection efficiency for targeted transgenic transformants is dramatically increased.
Results
Wild type C. reinhardtii strain is sensitive to nourseothricin
To explore the possibility to use NAT gene as a selectable marker for nuclear transformation, we first tested the sensitivities of C. reinhardtii to NTC. The selection concentrations for other organisms range from 20–400 µg/ml (https://www.jenabioscience.com/images/741d0cd7d0/NTC-Flyer.pdf). C. reinhardtii cells were placed on agar plates supplemented with different concentrations of NTC and grown for 4 days. The cells were sensitive to NTC at concentrations even as low as 2.5 µg/ml. At concentrations of 5 µg/ml and above, no viable cells were observed microscopically (Fig. 1) and even after 14 days (data not shown). Thus, we conclude that C. reinhardtii is sensitive to NTC, which paves the way for using NAT as a selectable maker.
NTC is compatible with paromomycin and hygromycin B selections
For studies involved with transgenic strains, multiple selections are usually required. For example, rescue of insertional mutants generated by using antibiotic resistance genes requires another antibiotic selectable maker. Commonly used antibiotics for selection of transgenic strains of C. reinhardtii include paromomycin and hygromycin B [1]. We wondered whether NTC is compatible with these two antibiotics for selection in C. reinhardtii. NTC, paromomycin and hygromycin B all inhibit protein synthesis, but their working mechanisms are different. Paromomycin inhibits initiation of translation or earlier steps of elongation while hygromycin B potently inhibits elongation [17]. NTC inhibits protein synthesis with miscoding activity [14]. It has been shown in mammalian cells and fungus that NTC is compatible with selections with hygromycin B [18]. Whether it is compatible with selection with paromomycin and/or hygromycin B in Chlamydomonas cells is not known.
Previously, our lab has generated strains with resistance to paromomycin, hygromycin B or both, which provided the needed resource for testing NTC compatibility with paromomycin and/or hygromycin B for selection. Wild type strain 21gr and strains with paromomycin, hygromycin B or both were grown on agar plate supplemented the antibiotics as indicated. As shown in Fig. 2, application of NTC killed wild type cells as well as cells with paromomycin, hygromycin B and both. Thus, there is no cross-resistance to NTC in strains that had been transformed with hygromycin B and/or paromomycin resistance genes, indicating that NTC can be used for multiple selection with paromomycin and/or hygromycin B.
No cross-resistance to NTC in strains with paromomycin and/or hygromycin B resistance. Wild type (WT) cells and transgenic strains with resistance to paromomycin, hygromycin B or both were grown for 4 days on TAP agar plates without antibiotics (a) or supplemented with paromomycin (b), hygromycin B (c) or NTC (d). paro, paromomycin resistant strains; hygro, hygromycin B resistant strains; paro/hygro, paromomycin and hygromycin B double resistance strains. Data shown are representative of three experiments
Expression of codon optimized NAT gene confers resistance to NTC of C. reinhardtii
The Chlamydomonas genome has a higher GC content and its gene expression exhibits bias of codon usage. Divergence of these properties in a foreign gene may adversely affect gene expression in Chlamydomonas likely due to changes in chromatin structure and abnormal low abundance of some tRNAs, respectively [19,20,21,22]. Though the GC content of NAT is similar to that of Chlamydomonas, some rarely used codons in Chlamydomonas are present in NAT. To test whether NAT can be used a selectable marker, the NAT gene from S. noursei was synthesized with codon optimization (Fig. 3). The NAT gene encodes a protein of 190 aa. We generated a pHR-NAT-HA plasmid harboring codon optimized NAT (Fig. 4). The NAT ORF tagged with 3xHA was placed under the control of the HSP70a/RBCS2 promoter containing one copy of RBCS2 intron 1 and RBCS2 3′UTR (Fig. 4a). The construct was transformed by electroporation into wild type (WT) cells. The transformants were grown on agar plates with or without NTC (Fig. 4b). Transformants on agar plates without NTC grew like a lawn. In contrast, individual colonies were observed on agar plate with NTC, suggesting that these colonies are NTC resistant. The NTC resistance of these colonies may be caused by rare mutations or expressing NAT. To discern these possibilities, 7 colonies were randomly picked and subjected to analysis for NAT expression by immunoblotting. All the colonies showed expression of NAT (Fig. 4c). The protein mass of NAT was similar to calculated molecular weight. Thus, these data demonstrate that NAT can be successfully expressed in C. reinhardtii and its expression can confer transgenic strains with NTC resistance. The transformation efficiency was 528 cfu per 50 ng plasmid DNA for three independent transformations.
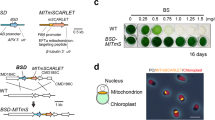
Expression of NAT confers cell resistance to NTC. a A diagram showing the construct for expressing NAT in C. reinhardtii. b WT cells were transformed with plasmid pHR-NAT-HA and grown on TAP agar plates supplemented with or without NTC (10 µg/ml). c Expression of NAT-HA was examined by immunoblotting with anti-HA antibody. α-tubulin was used as a loading control. d The extent of NTC resistance is correlated with the levels of NAT expression. Cells of wild type and strains with higher (#1 and #6) or lower (#2) expression levels of NAT were grown on agar plates supplemented with different concentrations of NTC as indicated
To examine the sensitivity of the transgenic strains with different levels of NAT expression, cells of strains with higher expression (strains #1 and #6) and lower expression (strain #2) were grown on agar plates supplemented with various concentrations of NTC (Fig. 4d). Consistent with results shown above, wild type cells were killed at 10 µg/ml NTC while the transformants grew normally. However, at higher concentrations of NTC, the transformants showed different extent of growth. For strain #2, which had lower expression of NAT, strong growth inhibition was observed at 50 µg/ml of NTC. In contrast, for strains #1 and #6, which had relatively higher expression of NAT, strong inhibition was observed at 100 µg/ml of NTC. These data further demonstrate that the tolerance to NTC is conferred by the expression of NAT and reveal that the extent of tolerance is correlated with the levels of NAT expression.
NAT can be used as a selectable marker for transgenic strains
Given that NAT can be expressed and confer NTC resistance in C. reinhardtii, we decided to examine it as a selectable marker for transgenic strains. ift54 is a mutant defective in IFT54, which was generated by insertional mutagenesis with paromomycin resistance gene AphVIII [23]. Loss of IFT54 blocks cilia formation (Fig. 5a). To prove our hypothesis, we generated a plasmid carrying NAT resistant gene as well as protein expression cassette of IFT54 (Fig. 5b). The plasmid was transformed into ift54 and the transformants were selected on agar plates with 10 µg/ml NTC. 12 out of 192 (6.5%) colonies grown on NTC selection agar plates expressed IFT54-HA as examined by immunoblotting (Fig. 5c and data not shown). And all strains expressing IFT54-HA rescued the aflagellar phenotype of ift54 (Fig. 5a). These data demonstrate that NAT can be used as a selectable marker even in strains with paromomycin resistance.
NAT as a selectable marker that is compatible with paromomycin and/or hygromycin B resistant genes. a Differential interference contrast (DIC) images showing wild type cells, ift54 mutant and rescued cells. Please note that ift54 did not have flagella and flagellar formation was rescued in IFT54-HA expressing cells. Bar, 10 μm. b A diagram showing construct that harbors expression cassettes of IFT54-HA and NAT. c Expression of IFT54-HA in strains harboring paromomycin resistant gene by using NAT as a selectable marker. The construct listed above was transformed into ift54, which harbors paromomycin resistant gene. The transformants were selected on agar plates supplemented with 10 µg/ml NTC. Cell lysates from randomly picked transformants were subjected to immunoblotting using the indicated antibodies. CDPK3 was used as a loading control. *Denotes non-specific bands. Wild type (WT) and ift54 cells were used as control. d Expression of IFT54-HA in strains harboring paromomycin and hygromycin B resistant genes by using NAT as a selectable marker. wdr92::WDR92-YFP strains with both paromomycin and hygromycin B resistant genes were transformed. The expression of IFT54-HA from cells grown on selection plate was examined by immunoblotting. WT and wdr92::WDR92-YFP (paro/hygro) cells were used as control
Next, we examined whether NAT can be used as a selectable marker in strains with both paromomycin and hygromycin B resistance. The pPSAD-IFT54-HA(NAT+) was transformed into a strain with paromomycin and hygromycin B resistance. 7 out of 116 (6.03%) colonies grown on the NTC selection plates expressed IFT54-HA as examined by immunoblotting (Fig. 5d and data not shown). Taken together, we have shown that NAT is an efficient selectable marker, which is compatible with paromomycin and/or hygromycin B resistance genes.
Fusion of a target gene IFT54-HA to NAT and processing with the FMDV 2A peptide increases gene expression efficiency
It has been reported that the gene expression efficiency is much improved by fusion of a target gene to a selectable marker and processing with the FMDV 2A peptide [24]. Due to cleavage after gene translation at the 2A peptide sequence, the resulting protein is processed into two discrete proteins: a protein from the target gene and the selectable marker protein fused with short 2A peptide [25]. To demonstrate whether NAT can be used as such a selectable marker, we made a construct by fusing IFT54-HA to NAT and DNA sequence of FMDV 2A (Fig. 6a). The construct was transformed into ift54 and the transformants were selected on agar plates with 10 µg/ml NTC. 122 out of 204 (59.8%) colonies had flagella. Examination of a few transformants with flagella showed that they all expressed IFT54-HA (Fig. 6b). Thus, we predicted that all the transformants that had formed flagella should have expressed IFT54-HA. Compared to the construct that does not fuse IFT54-HA to NAT as shown in Fig. 5b, this construct led to a ninefold increase of selection efficiency (6.5% to 59.8%) for transgenic strains. However, by examination of the protein levels of IFT54-HA in transformants derived from these two constructs (Figs. 5c and 6b), we have not observed increased protein expression level by such a fusion construct as reported [24].
Fusion of IFT54-HA to NAT and FMDV 2A for cell transformation. a Schematic representation of the construct used to drive NAT-2A-IFT54-HA fusion protein expression. b Immunoblot analysis of IFT54-HA expression in transformants from the fusion construct mediated transformation. Cell lysates from WT, ift54 and the transformants were subjected to immunoblotting with IFT54 and CDPK3 antibodies. CDPK3 was used as a loading control. *Denotes a band in each lane that reacts non-specifically. Please note, the predicted fusion protein NAT-FMDV 2A–IFT54-HA was cleaved in the cell to generate IFT54-HA
Discussion
The ability to generate transgenic cells is crucial for genetic engineering widely used in basic research as well as in biotechnological applications. As a model organism, C. reinhardtii is widely used for exploration of basic cellular processes such as cilia biogenesis and photosynthesis and for producing commercially valuable products as a cell factory [1,2,3]. Although a few selectable markers have been developed in this organism, few of them have been widely used. An ideal selectable marker may possess the following properties: (1) high stability, aqueous solubility and low dosage of the selection reagents; (2) non-toxicity of the selection reagents in the presence of a selectable marker; (3) non-toxicity of the selectable markers; 4) high efficiency of transformation; (5) compatibility with other selectable makers and 6) no genotype requirement for the parental strains.
We have shown that NAT is an effective and stable selectable marker in C. reinhardtii that confers resistance to NTC. C. reinhardtii is very sensitive to NTC. No viable colonies were observed even in the presence of 5 µg/ml NTC though we have used 10 µg/ml for the selection. NTC is soluble in water and highly stable. The transformation efficiency of NAT is high. Around 500 cfu were routinely obtained by using 50 ng plasmid DNA for transformation. Expression of NAT was stable even in the absence of NTC. We have not observed any growth defects in NAT transgenic strains. As NTC is an antibiotic, it does not require strains with specific genotype. Thus, NAT provides an alternative choice for selectable markers in C. reinhardtii. Random insertion of foreign DNA into the genome of C. reinhardtii occurs during transformation and this property has been used to generate insertional mutants [26, 27]. Though we have not examined the patterns of integration of NAT into the genome of C. reinhardtii, NAT is expected to behave as other foreign DNA fragments. Thus, NAT may be used for generation of insertional mutants from which desired functional genes can be cloned.
We have tested using NAT as a selectable marker for transgenic expression of a target gene IFT54. We have used parental strains that had been previously transformed with paromomycin and/or hygromycin B resistant genes. Around 6.5% IFT54 transgenic strains were obtained from NTC resistant colonies, demonstrating that NAT can be used as a selectable marker. These data also indicate that NAT is compatible with hygromycin B and paromomycin resistant genes, which allow for multiple selections. We have developed a construct by fusing the target gene IFT54 to NAT and processing with FMDV 2A peptide. Compared to the non-fusion construct, the efficiency of expression of IFT54-HA has increased around ninefold. Thus, this fusion expression system can increase selection efficiency of transgenic strains. Because the NTC resistance is correlated with the expression levels of NAT, this system may also be used for obtaining strains with higher expression of target genes by selection at higher concentrations of NTC.
NAT as a selectable marker has been used in microalgae but so far only in marine diatoms including Chaetoceros gracilis, P. tricornutum and T. pseudonana [16, 28, 29]. Our demonstration that NAT can be used a selectable marker in a fresh water green alga, opening a promising prospect in using NAT in other microalgae, especially in those algae with fewer choices for selectable markers. For example, Dunaliella, a saline green alga, is a popular model organism for the study of adaptation of eukaryotic cells to high salt concentrations and some Dunaliella species are of economic value for producing beta-carotene [30]. However, Dunaliella is resistant to paromomycin, hygromycin B, spectinomycin and kanamycin [3]. Thus, NTC resistance needs to be tested in Dunaliella before the NAT/NTC selection system can be used.
Conclusions
We have developed a new stable selectable marker for selection of transgenic strains in C. reinhardtii that confers resistance to NTC, which provides an alternative choice for selectable markers. In addition, NAT is compatible with paromomycin and hygromycin B resistance genes, two most commonly used selectable markers in C. reinhardtii, which allows combination of multiple selectable markers in transgenic studies.
Methods
Strains and culture
Chlamydomonas reinhardtii wild type strain 21gr (CC-1690, mt+) was from the Chlamydomonas Resource Center. ift54 (a paromomycin resistant strain) [23], lf4::LF4-HA (a hygromycin B resistant strain) [31] and wdr92::WDR92-YFP (a paromomycin and hygromycin B double resistant strain) [32] were generated in our own lab. Unless otherwise stated, cells were grown at 23 °C in M liquid medium in a 14/10 light/dark cycle [33]. Cells used for transformation were grown at 23 °C in liquid TAP medium under continuous light [34].
Reagents
Paromomycin and hygromycin B were purchased from Merck Millipore, USA, while NTC was obtained from Jena biosciences, Germany. The antibiotics were solubilized in water and sterilized by filtration. The concentrations used for selection for paromomycin, hygromycin B and NTC were 10, 20 and 10 µg/ml, respectively.
Drug sensitivity assay
To determine the sensitivity of C. reinhardtii to NTC, 1 × 106 of wild type cells were spotted on 1.5% TAP agar plates supplemented with various concentrations of NTC (0, 2.5, 5, 10 and 40 µg/ml) and incubated for 4 days at 23 °C in a 14/10 light/dark cycle. To test whether strains with paromomycin and/or hygromycin B resistant genes are sensitive to NTC, 1 × 106 cells of wild type and strains with various resistant genes were grown on 1.5% TAP agar plates supplemented with different antibiotics as indicated in the text.
Construction of the transformation vectors
To generate a construct for expressing NAT, the coding region of NAT from S. noursei was codon optimized for C. reinhardtii and chemically synthesized (Genscript, China). Codon optimized NAT tagged with 3× HA tag at the 3′ end driven by HSP70a/RBCS2 and terminated by RBCS2 terminator was cloned into ZT4-blunt vector. The HSP70a/RBCS2 promoter and RBCS2 terminator were cloned from pCB740 [35]. The 3× HA tag was cloned from pKL-3XHA [36]. The final construct was termed pHR-NAT-HA. To enable expression of IFT54-HA in C. reinhardtii with NAT as a selectable marker, the expression cassette for hygromycin B in the vector pPSAD-IFT54-HA-Hyg was replaced with the NAT expression cassette in pHR-NAT-HA with 3xHA being removed [23]. The resulting construct was termed pPSAD-IFT54-HA(NAT+). To generate construct pHR-NAT-2A-IFT54-HA for fusion of IFT54-HA to NAT and processing by FMDV 2A peptide, the sequence for Ble and GFP-tubulin in the vector pBR25-sfGFP-TUA were replaced by NAT and IFT54-HA, respectively [37]. All the constructs were verified by sequencing.
Electroporation transformation
Transformation of Chlamydomonas was performed by electroporation using BTX ECM630 (Harvard Apparatus Inc, USA) following a previously published protocol [38]. For each transformation, 5 × 107 cells were mixed with 50 ng plasmid DNA linearized by AclI. After electroporation, the transformation mixture was diluted in 10 ml TAP + 50 mM sorbitol and kept away from light for 8 h. Transformants were selected on agar plates supplemented with 10 µg/ml NTC.
SDS-PAGE and immunoblotting
SDS-PAGE and immunoblotting analysis were performed as described previously [39]. Briefly, cells were lysed with Buffer A (50 mM Tris–HCl pH 7.5, 10 mM MgCl2, 1 mM EDTA, and 1 mM DTT) containing protease inhibitor cocktail (Roche, Switzerland) and boiled for 10 min in 1× SDS loading buffer. The proteins were separated in 10% SDS-PAGE, transferred to polyvinylidene difluoride (PVDF) membranes (Merck Millipore, USA) and probed with the indicated antibodies. The primary antibodies used include the following: rat anti-HA (Roche, Switzerland), 1:3000; mouse anti-α-tubulin (Sigma-Aldrich, USA), 1: 3000; rabbit anti-IFT54, 1:3000 [23] and rabbit anti-CDPK3, 1: 5000 [38].
Cell imaging
After cell fixation in 1% glutaraldehyde, DIC images were captured by Zeiss Axio Observer Z1 microscope (Carl Zeiss, Germany) equipped with a CCD camera (QuantEM512SC, Photometrics, USA). The images were processed in Photoshop and/or Illustrator (Adobe, USA).
Availability of data and materials
All data generated or analyzed during this study are included in this published article. Experimental materials generated during the current study are available from the corresponding author on reasonable request.
Abbreviations
- NAT:
-
nourseothricin (NTC) N-acetyl transferase
- NTC:
-
nourseothricin
- FMDV 2A:
-
foot-and-mouth-disease-virus 2A
- HR:
-
hSP70a/RBCS2
- IFT54:
-
intraflagellar transport 54
- CDPK3:
-
calcium dependent kinase 3
- DIC:
-
differential interference contrast
References
Salome PA, Merchant SS. A series of fortunate events: introducing Chlamydomonas as a reference organism. Plant Cell. 2019;31:1682–707.
Doron L, Segal N, Shapira M. Transgene expression in microalgae-from tools to applications. Front Plant Sci. 2016;7:505.
Vazquez-Villegas P, Torres-Acosta MA, Garcia-Echauri SA, Aguilar-Yanez JM, Rito-Palomares M, Ruiz-Ruiz F. Genetic manipulation of microalgae for the production of bioproducts. Front Biosci. 2018;10:254–75.
Kindle KL. High-frequency nuclear transformation of Chlamydomonas reinhardtii. Proc Natl Acad Sci USA. 1990;87:1228–322.
Brueggeman AJ, Kuehler D, Weeks DP. Evaluation of three herbicide resistance genes for use in genetic transformations and for potential crop protection in algae production. Plant Biotechnol J. 2014;12:894–902.
Tabatabaei I, Dal Bosco C, Bednarska M, Ruf S, Meurer J, Bock R. A highly efficient sulfadiazine selection system for the generation of transgenic plants and algae. Plant Biotechnol J. 2019;17:638–49.
Erickson JM, Rahire M, Bennoun P, Delepelaire P, Diner B, Rochaix JD. Herbicide resistance in Chlamydomonas reinhardtii results from a mutation in the chloroplast gene for the 32-kilodalton protein of photosystem II. Proc Natl Acad Sci USA. 1984;81:3617–21.
Berthold P, Schmitt R, Mages W. An engineered Streptomyces hygroscopicus aph 7" gene mediates dominant resistance against hygromycin B in Chlamydomonas reinhardtii. Protist. 2002;153:401–12.
Sizova I, Fuhrmann M, Hegemann P. A streptomyces rimosus aphVIII gene coding for a new type phosphotransferase provides stable antibiotic resistance to Chlamydomonas reinhardtii. Gene. 2001;277:221–9.
Stevens DR, Rochaix JD, Purton S. The bacterial phleomycin resistance gene ble as a dominant selectable marker in Chlamydomonas. Mol Gen Genet. 1996;251:23–30.
van Peer AF, de Bekker C, Vinck A, Wosten HA, Lugones LG. Phleomycin increases transformation efficiency and promotes single integrations in Schizophyllum commune. Appl Environ Microbiol. 2009;75:1243–7.
Sundar IK, Sakthivel N. Advances in selectable marker genes for plant transformation. J Plant Physiol. 2008;165:1698–716.
Mortensen RM, Kingston RE. Selection of transfected mammalian cells. Curr Protoc Mol Biol. 2009. https://doi.org/10.1002/0471142727.
I Haupt R Hubener H Thrum 1978 Streptothricin F, an inhibitor of protein synthesis with miscoding activity. J Antibiot. 1978;31:1137–1142
Zahringer U, Voigt W, Seltmann G. Nourseothricin (streptothricin) inactivated by a plasmid pIE636 encoded acetyl transferase of Escherichia coli: location of the acetyl group. FEMS Microbiol Lett. 1993;110:331–4.
Zaslavskaia LA, Lippmeier JC, Kroth PG, Grossman AR, Apt KE. Transformation of the diatom Phaeodactylum tricornutum (Bacillariophyceae) with a variety of selectable marker and reporter genes. J Phycol. 2000;36:379–86.
Eustice DC, Wilhelm JM. Fidelity of the eukaryotic codon-anticodon interaction: interference by aminoglycoside antibiotics. Biochemistry. 1984;23:1462–7.
Kochupurakkal BS, Iglehart JD. Nourseothricin N-acetyl transferase: a positive selection marker for mammalian cells. PLoS ONE. 2013;8:e68509.
Barahimipour R, Strenkert D, Neupert J, Schroda M, Merchant SS, Bock R. Dissecting the contributions of GC content and codon usage to gene expression in the model alga Chlamydomonas reinhardtii. Plant J. 2015;84:704–17.
Shao N, Bock R. A codon-optimized luciferase from Gaussia princeps facilitates the in vivo monitoring of gene expression in the model alga Chlamydomonas reinhardtii. Curr Genet. 2008;53:381–8.
Fuhrmann M, Oertel W, Hegemann P. A synthetic gene coding for the green fluorescent protein (GFP) is a versatile reporter in Chlamydomonas reinhardtii. Plant J. 1999;19:353–61.
Kane JF. Effects of rare codon clusters on high-level expression of heterologous proteins in Escherichia coli. Curr Opin Biotechnol. 1995;6:494–500.
Zhu X, Liang Y, Gao F, Pan J. IFT54 regulates IFT20 stability but is not essential for tubulin transport during ciliogenesis. Cell Mol Life Sci. 2017;74:3425–37.
Rasala BA, Lee PA, Shen Z, Briggs SP, Mendez M, Mayfield SP. Robust expression and secretion of Xylanase1 in Chlamydomonas reinhardtii by fusion to a selection gene and processing with the FMDV 2A peptide. PLoS ONE. 2012;7:e43349.
Ryan MD, King AM, Thomas GP. Cleavage of foot-and-mouth disease virus polyprotein is mediated by residues located within a 19 amino acid sequence. J Gen Virol. 1991;72:2727–32.
Gonzalez-Ballester D, de Montaigu A, Galvan A, Fernandez E. Restriction enzyme site-directed amplification PCR: a tool to identify regions flanking a marker DNA. Anal Biochem. 2005;340:330–5.
Tam LW, Lefebvre PA. Cloning of flagellar genes in Chlamydomonas reinhardtii by DNA insertional mutagenesis. Genetics. 1993;135:375–84.
Ifuku K, Yan D, Miyahara M, Inoue-Kashino N, Yamamoto YY, Kashino Y. A stable and efficient nuclear transformation system for the diatom Chaetoceros gracilis. Photosynth Res. 2015;123:203–11.
Poulsen N, Chesley PM, Kröger N. molecular genetic manipulation of the diatom Thalassiosira pseudonana (Bacillariophyceae)1. J Phycol. 2006;42:1059–65.
Oren A. A Century of Dunaliella Research: 1905–2005. In: Gunde-Cimerman N, Oren A, Plemenitaš A, editors. Adaptation to life at high salt concentrations in archaea, bacteria, and eukarya. Dordrecht: Springer; 2005. p. 491–502.
Wang Y, Ren Y, Pan J. Regulation of flagellar assembly and length in Chlamydomonas by LF4, a MAPK-related kinase. FASEB J. 2019;33:6431–41.
Liu G, Wang L, Pan J. Chlamydomonas WDR92 in association with R2TP-like complex and multiple DNAAFs to regulate ciliary dynein preassembly. J Mol Cell Biol. 2019;11:770–80.
Sager R, Granick S. Nutritional studies with Chlamydomonas reinhardtii. Ann N Y Acad Sci. 1953;56:831–8.
Gorman DS, Levine RP. Cytochrome f and plastocyanin: their sequence in the photosynthetic electron transport chain of Chlamydomonas reinhardtii. Proc Natl Acad Sci USA. 1965;54:1665–9.
Schroda M, Blocker D, Beck CF. The HSP70A promoter as a tool for the improved expression of transgenes in Chlamydomonas. Plant J. 2000;21:121–31.
Lechtreck KF, Luro S, Awata J, Witman GB. HA-tagging of putative flagellar proteins in Chlamydomonas reinhardtii identifies a novel protein of intraflagellar transport complex B. Cell Motil Cytoskeleton. 2009;66:469–82.
Craft JM, Harris JA, Hyman S, Kner P, Lechtreck KF. Tubulin transport by IFT is upregulated during ciliary growth by a cilium-autonomous mechanism. J Cell Biol. 2015;208:223–37.
Liang Y, Pan J. Regulation of flagellar biogenesis by a calcium dependent protein kinase in Chlamydomonas reinhardtii. PLoS ONE. 2013;8:e69902.
Pan J, Snell WJ. Regulated targeting of a protein kinase into an intact flagellum An aurora/Ipl1p-like protein kinase translocates from the cell body into the flagella during gamete activation in Chlamydomonas. J Biol Chem. 2000;275:24106–14.
Acknowledgements
We would like to thank Dr. Karl F. Lechtreck (University of Georgia) for providing plasmids pBR25-sfGFP-TUA.
Funding
This work was supported by the National Key R&D Program of China (2018YFA0902500) and the National Natural Science Foundation of China (31671387) (to J.P.)
Author information
Authors and Affiliations
Contributions
XJY and JLP performed the experiments. XJY and JMP analyzed the data. JMP provided reagents and supervised the project. JMP and XJY wrote the paper. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated.
About this article
Cite this article
Yang, X., Peng, J. & Pan, J. Nourseothricin N-acetyl transferase (NAT), a new selectable marker for nuclear gene expression in Chlamydomonas. Plant Methods 15, 140 (2019). https://doi.org/10.1186/s13007-019-0526-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s13007-019-0526-5